Abstract
Memory T cells are critical for the immune response to recurring infections. Their instantaneous reactivity to pathogens is empowered by the persistent expression of cytokine-encoding mRNAs. How the translation of proteins from pre-formed cytokine-encoding mRNAs is prevented in the absence of infection has remained unclear. Here we found that protein production in memory T cells was blocked via a 3′ untranslated region (3′ UTR)-mediated process. Germline deletion of AU-rich elements (AREs) in the Ifng-3′ UTR led to chronic cytokine production in memory T cells. This aberrant protein production did not result from increased expression and/or half-life of the mRNA. Instead, AREs blocked the recruitment of cytokine-encoding mRNA to ribosomes; this block depended on the ARE-binding protein ZFP36L2. Thus, AREs mediate repression of translation in mouse and human memory T cells by preventing undesirable protein production from pre-formed cytokine-encoding mRNAs in the absence of infection.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Masopust, D. & Schenkel, J. M. The integration of T cell migration, differentiation and function. Nat. Rev. Immunol. 13, 309–320 (2013).
Sheridan, B. S. & Lefrançois, L. Regional and mucosal memory T cells. Nat. Immunol. 12, 485–491 (2011).
Harty, J. T., Tvinnereim, A. R. & White, D. W. CD8+ T cell effector mechanisms in resistance to infection. Annu. Rev. Immunol. 18, 275–308 (2000).
Veiga-Fernandes, H., Walter, U., Bourgeois, C., McLean, A. & Rocha, B. Response of naïve and memory CD8+ T cells to antigen stimulation in vivo. Nat. Immunol. 1, 47–53 (2000).
London, C. A., Lodge, M. P. & Abbas, A. K. Functional responses and costimulator dependence of memory CD4+ T cells. J. Immunol. 164, 265–272 (2000).
Whitmire, J. K., Eam, B. & Whitton, J. L. Tentative T cells: memory cells are quick to respond, but slow to divide. PLoS Pathog. 4, e1000041 (2008).
Guidotti, L. G. & Chisari, F. V. Noncytolytic control of viral infections by the innate and adaptive immune response. Annu. Rev. Immunol. 19, 65–91 (2001).
Soudja, S. M., Ruiz, A. L., Marie, J. C. & Lauvau, G. Inflammatory monocytes activate memory CD8+ T and innate NK lymphocytes independent of cognate antigen during microbial pathogen invasion. Immunity 37, 549–562 (2012).
Weng, N. P., Araki, Y. & Subedi, K. The molecular basis of the memory T cell response: differential gene expression and its epigenetic regulation. Nat. Rev. Immunol. 12, 306–315 (2012).
Wherry, E. J. et al. Molecular signature of CD8+ T cell exhaustion during chronic viral infection. Immunity 27, 670–684 (2007).
Philip, M. et al. Chromatin states define tumour-specific T cell dysfunction and reprogramming. Nature 545, 452–456 (2017).
Kaech, S. M., Hemby, S., Kersh, E. & Ahmed, R. Molecular and functional profiling of memory CD8 T cell differentiation. Cell 111, 837–851 (2002).
Swanson, B. J., Murakami, M., Mitchell, T. C., Kappler, J. & Marrack, P. RANTES production by memory phenotype T cells is controlled by a posttranscriptional, TCR-dependent process. Immunity 17, 605–615 (2002).
Willinger, T., Freeman, T., Hasegawa, H., McMichael, A. J. & Callan, M. F. Molecular signatures distinguish human central memory from effector memory CD8 T cell subsets. J. Immunol. 175, 5895–5903 (2005).
Salerno, F., Paolini, N. A., Stark, R., von Lindern, M. & Wolkers, M. C. Distinct PKC-mediated posttranscriptional events set cytokine production kinetics in CD8+ T cells. Proc. Natl. Acad. Sci. USA 114, 9677–9682 (2017).
Hodge, D. L. et al. IFN-gamma AU-rich element removal promotes chronic IFN-gamma expression and autoimmunity in mice. J. Autoimmun. 53, 33–45 (2014).
Kontoyiannis, D., Pasparakis, M., Pizarro, T. T., Cominelli, F. & Kollias, G. Impaired on/off regulation of TNF biosynthesis in mice lacking TNF AU-rich elements: implications for joint and gut-associated immunopathologies. Immunity 10, 387–398 (1999).
Turner, M. & Hodson, D. Regulation of lymphocyte development and function by RNA-binding proteins. Curr. Opin. Immunol. 24, 160–165 (2012).
Kafasla, P., Skliris, A. & Kontoyiannis, D. L. Post-transcriptional coordination of immunological responses by RNA-binding proteins. Nat. Immunol. 15, 492–502 (2014).
Salerno, F. & Wolkers, M. C. T-cells require post-transcriptional regulation for accurate immune responses. Biochem. Soc. Trans. 43, 1201–1207 (2015).
Bronevetsky, Y. et al. T cell activation induces proteasomal degradation of Argonaute and rapid remodeling of the microRNA repertoire. J. Exp. Med. 210, 417–432 (2013).
Wu, H. et al. miRNA profiling of naïve, effector and memory CD8 T cells. PLoS One 2, e1020 (2007).
Garneau, N. L., Wilusz, J. & Wilusz, C. J. The highways and byways of mRNA decay. Nat. Rev. Mol. Cell Biol. 8, 113–126 (2007).
Grammatikakis, I., Abdelmohsen, K. & Gorospe, M. Posttranslational control of HuR function. Wiley Interdiscip. Rev. RNA 8, e1372 (2017).
Jeltsch, K. M. et al. Cleavage of roquin and regnase-1 by the paracaspase MALT1 releases their cooperatively repressed targets to promote TH17 differentiation. Nat. Immunol. 15, 1079–1089 (2014).
Vlasova-St Louis, I. & Bohjanen, P. R. Post-transcriptional regulation of cytokine signaling by AU-rich and GU-rich elements. J. Interferon Cytokine Res. 34, 233–241 (2014).
Beisang, D. & Bohjanen, P. R. Perspectives on the ARE as it turns 25 years old. Wiley Interdiscip. Rev. RNA 3, 719–731 (2012).
Schoenberg, D. R. & Maquat, L. E. Regulation of cytoplasmic mRNA decay. Nat. Rev. Genet. 13, 246–259 (2012).
Zehn, D., Lee, S. Y. & Bevan, M. J. Complete but curtailed T-cell response to very low-affinity antigen. Nature 458, 211–214 (2009).
Penix, L., Weaver, W. M., Pang, Y., Young, H. A. & Wilson, C. B. Two essential regulatory elements in the human interferon gamma promoter confer activation specific expression in T cells. J. Exp. Med. 178, 1483–1496 (1993).
Hamilton, S. E., Wolkers, M. C., Schoenberger, S. P. & Jameson, S. C. The generation of protective memory-like CD8+ T cells during homeostatic proliferation requires CD4+ T cells. Nat. Immunol. 7, 475–481 (2006).
Leppek, K. & Stoecklin, G. An optimized streptavidin-binding RNA aptamer for purification of ribonucleoprotein complexes identifies novel ARE-binding proteins. Nucleic Acids Res. 42, e13 (2014).
Brooks, S. A. & Blackshear, P. J. Tristetraprolin (TTP): interactions with mRNA and proteins, and current thoughts on mechanisms of action. Biochim. Biophys. Acta 1829, 666–679 (2013).
Galloway, A. et al. RNA-binding proteins ZFP36L1 and ZFP36L2 promote cell quiescence. Science 352, 453–459 (2016).
Ogilvie, R. L. et al. Tristetraprolin mediates interferon-gamma mRNA decay. J. Biol. Chem. 284, 11216–11223 (2009).
Taylor, G. A. et al. A pathogenetic role for TNF alpha in the syndrome of cachexia, arthritis, and autoimmunity resulting from tristetraprolin (TTP) deficiency. Immunity 4, 445–454 (1996).
Hodson, D. J. et al. Deletion of the RNA-binding proteins ZFP36L1 and ZFP36L2 leads to perturbed thymic development and T lymphoblastic leukemia. Nat. Immunol. 11, 717–724 (2010).
Lindstein, T., June, C. H., Ledbetter, J. A., Stella, G. & Thompson, C. B. Regulation of lymphokine messenger RNA stability by a surface-mediated T cell activation pathway. Science 244, 339–343 (1989).
Ma, F. et al. The microRNA miR-29 controls innate and adaptive immune responses to intracellular bacterial infection by targeting interferon-γ. Nat. Immunol. 12, 861–869 (2011).
Steiner, D. F. et al. MicroRNA-29 regulates T-box transcription factors and interferon-γ production in helper T cells. Immunity 35, 169–181 (2011).
Fleischer, T. C., Weaver, C. M., McAfee, K. J., Jennings, J. L. & Link, A. J. Systematic identification and functional screens of uncharacterized proteins associated with eukaryotic ribosomal complexes. Genes Dev. 20, 1294–1307 (2006).
Mackay, L. K. et al. Hobit and Blimp1 instruct a universal transcriptional program of tissue residency in lymphocytes. Science 352, 459–463 (2016).
Hudson, B. P., Martinez-Yamout, M. A., Dyson, H. J. & Wright, P. E. Recognition of the mRNA AU-rich element by the zinc finger domain of TIS11d. Nat. Struct. Mol. Biol. 11, 257–264 (2004).
Goldrath, A. W. et al. Cytokine requirements for acute and basal homeostatic proliferation of naive and memory CD8+ T cells. J. Exp. Med. 195, 1515–1522 (2002).
Hochweller, K. et al. Dendritic cells control T cell tonic signaling required for responsiveness to foreign antigen. Proc. Natl. Acad. Sci . 107, 5931–5936 (2010).
Swamy, M. et al. A cholesterol-based allostery model of T cell receptor phosphorylation. Immunity 44, 1091–1101 (2016).
Abdelsamed, H. A. et al. Human memory CD8 T cell effector potential is epigenetically preserved during in vivo homeostasis. J. Exp. Med. 214, 1593–1606 (2017).
Piecyk, M. et al. TIA-1 is a translational silencer that selectively regulates the expression of TNF-alpha. EMBO J. 19, 4154–4163 (2000).
Gruber, A. R., Fallmann, J., Kratochvill, F., Kovarik, P. & Hofacker, I. L. AREsite: a database for the comprehensive investigation of AU-rich elements. Nucleic Acids Res. 39, D66–D69 (2011).
Stetson, D. B. et al. Constitutive cytokine mRNAs mark natural killer (NK) and NK T cells poised for rapid effector function. J. Exp. Med. 198, 1069–1076 (2003).
Gessner, A., Mohrs, K. & Mohrs, M. Mast cells, basophils, and eosinophils acquire constitutive IL-4 and IL-13 transcripts during lineage differentiation that are sufficient for rapid cytokine production. J. Immunol. 174, 1063–1072 (2005).
Villarino, A. V. et al. Posttranscriptional silencing of effector cytokine mRNA underlies the anergic phenotype of self-reactive T cells. Immunity 34, 50–60 (2011).
Hombrink, P. et al. Programs for the persistence, vigilance and control of human CD8+ lung-resident memory T cells. Nat. Immunol. 17, 1467–1478 (2016).
Zhang, L. et al. ZFP36L2 is required for self-renewal of early burst-forming unit erythroid progenitors. Nature 499, 92–96 (2013).
Lee, P. P. et al. A critical role for Dnmt1 and DNA methylation in T cell development, function, and survival. Immunity 15, 763–774 (2001).
Brummelkamp, T. R., Bernards, R. & Agami, R. Stable suppression of tumorigenicity by virus-mediated RNA interference. Cancer Cell 2, 243–247 (2002).
Kessels, H. W., Wolkers, M. C., van den Boom, M. D., van der Valk, M. A. & Schumacher, T. N. Immunotherapy through TCR gene transfer. Nat. Immunol. 2, 957–961 (2001).
Salerno, F., Guislain, A., Cansever, D. & Wolkers, M. C. TLR-mediated innate production of IFN-γ by CD8+ T cells is independent of glycolysis. J. Immunol. 196, 3695–3705 (2016).
McCausland, M. M. et al. SAP regulation of follicular helper CD4 T cell development and humoral immunity is independent of SLAM and Fyn kinase. J. Immunol. 178, 817–828 (2007).
López de Silanes, I., Zhan, M., Lal, A., Yang, X. & Gorospe, M. Identification of a target RNA motif for RNA-binding protein HuR. Proc. Natl. Acad. Sci. USA 101, 2987–2992 (2004).
Rappsilber, J., Ishihama, Y. & Mann, M. Stop and go extraction tips for matrix-assisted laser desorption/ionization, nanoelectrospray, and LC/MS sample pretreatment in proteomics. Anal. Chem. 75, 663–670 (2003).
Kulak, N. A., Pichler, G., Paron, I., Nagaraj, N. & Mann, M. Minimal, encapsulated proteomic-sample processing applied to copy-number estimation in eukaryotic cells. Nat. Methods 11, 319–324 (2014).
Cox, J. & Mann, M. MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat. Biotechnol. 26, 1367–1372 (2008).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Acknowledgements
We thank the animal caretakers of the NKI and the FACS facility of Sanquin Research and the Babraham Institute for excellent assistance. We thank D. Zehn (TU Munich) for the LM-OVA strain, J. Rohr (University of Freiburg) for Listeria cultures; H. Meijer for sharing the sucrose cushion protocol; G. Stoecklin (University of Heidelberg) for the 4xS1m aptamer construct; R. Arens (Leiden University), T. Schumacher (Netherlands Cancer Institute) and J. den Haan (Free University, Amsterdam) for providing mice; S. Libregts, B. van Steensel, K. Moore and B. Nicolet for technical help and advice; and D. Amsen, M. Nolte and R. van Lier for critical reading of the manuscript. M.T. and S.E.B. are supported by the Biotechnology and Biological Sciences Research Council. D.L.H. and H.A.Y. are funded through the intramural research program of the US NCI, NIH. The use of materials and reagents does not imply any endorsement of these products by the US government. This research was supported by the Dutch Science Foundation (VENI grant 916.76.127/VIDI grant 917.14.314, to M.C.W.).
Author information
Authors and Affiliations
Contributions
F.S. and M.C.W. designed experiments, F.S., S.E., M.v.d.B., F.P.J.v.A., A.G., W.Z., S.E.B. and M.C.W. performed experiments and analyzed the data, D.L.H. and H.A.Y. contributed the ARE-Del mice; S.E.B. and M.T. contributed the CD4cre-Zfp36l2flox/flox mice; M.v.L., M.T., H.A.Y. and J.P.M. provided intellectual input; M.C.W. directed the study; F.S. and M.C.W. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Ifng 3′-UTR-dependent regulation of GFP expression in T cells.
(a-b) Blood from LM-OVA infected mice (Fig. 1) was analyzed for GFP-MFI of GFPcontrol or GFP-Ifng3’UTR-expressing OTI cells. Data are represented as percentage of day 6 (n = 10 mice/group). (a) [One-way ANOVA with Dunnett’s multiple comparison to day 6 time point; *p < 0.05; **p < 0.005; ****p < 0.0001]. (b) [Unpaired Student t-test; **p < 0.005; ****p < 0.0001]. (c-h) T cells from (c-f) C57BL/6 J mice or (g-i) OTI mice were transduced with GFP-Ifng3’UTR or GFPcontrol. (c-f) Representative dot plots of GFP expression of unstimulated resting CD8+ T cells and CD4+ T cells (c,e), or reactivated for 4 h with PMA + ionomycin (d,f). Numbers in (c,e) indicate GFP-MFI. Graphs depict data compiled from 3 independently performed experiments (n = 4 mice/group). (c,e) [Unpaired Student t-test; **p < 0.005; ****p < 0.0001]. (d,f) [Paired Student t-test; *p < 0.05; **p < 0.005]. (g) Intracellular IFN-γ staining of resting (upper panel), or reactivated (OVA257-264 peptide; lower panel) GFP-Ifng3’UTR- or GFPcontrol-expressing OTI cells. Numbers indicate percentage of GFP+ and/or IFN-γ+ T cells. (h) Representative dot plot of GFPcontrol and GFP-Ifng 3’UTR expression levels under the minimal murine Ifng promoter in resting (left) and reactivated OTI cells (right). Numbers indicate GFP-MFI.
Supplementary Figure 2 All conserved AREs contribute to the Ifng 3′-UTR-mediated post-transcriptional regulation.
(a-b) GFP expression was measured in resting (gray histograms), and reactivated (black lines) OTI cells expressing wild-type GFP-Ifng 3’UTR or GFP-Ifng 3’UTR with indicated ARE mutants. Data are representative of three independently performed experiments. (c) Multiple sequence alignment of the Ifng 3’UTR of 9 representative species performed as described (Di Tommaso, P. et al. T-Coffee: a web server for the multiple sequence alignment of protein and RNA sequences using structural information and homology extension. Nucleic Acids Res. 39, W13–W17; 2011), combined with Boxshade analysis. For the alignment with the mRNA from gorilla, chimpanzee, green monkey, and Yangtze river dolphin, predicted sequences (Pubmed) have been used. ARE sequences are highlighted in yellow, underlined and numbered as in Fig. 2b. (d) Activated human T cells were transduced with wild-type human IFNG 3’UTR (WT), or with indicated ARE mutants. Graphs indicate GFP-MFI during resting phase (top) or GFP fold increase after reactivation with PMA + ionomycin (bottom). [One-way ANOVA with Dunnett’s multiple comparison to the control vector; n = 4 donors per group; *p < 0.005].
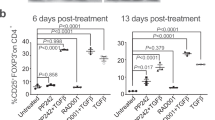
Supplementary Figure 3 Chronic IFN-γ production occurs in all TM subsets of IFN-γ-ARE-Del cells.
(a) Relative distribution of TEFF, TEM and TCM, and of SLEC/MPEC OTI cells 35 days post LM-OVA infection as defined in Fig. 3b (n = 8 mice/group). (b) Percentage (left panel) and MFI (right panel) of spontaneous IFN-γ production at day 48 post LM-OVA infection of spleen- and liver-residing wild-type and IFN-γ-ARE-Del TM cells. (c-d) IFN-γ MFI of spleen- (c) and IFN-γ MFI and percentage of IFN-γ producing T cells of liver- (d) residing wild-type (gray) and IFN-γ-ARE-Del (white) T cell subsets at day 35 post LM-OVA infection. (e) Ifng mRNA levels of sorted spleen-derived naive CD44lowCD62LhiCD8+ T cells and memory-like CD44 hiCD8+ T cells of 6–8 weeks old wild-type OTI mice. Results (mean ± SD) are pooled from 7 mice and five independently performed experiments. (f) Intracellular IFN-γ and TNF cytokine staining after 4h incubation with 1μg/ml brefeldin A. (g,h) IFN-γ production (g) and Ifng mRNA levels (h) of paired sorted and unstimulated CD44hiCD62Llow TEFF/EM and CD44hiCD62Lhi TCM CD8+ T cells. (a-h) [Unpaired Student t-test; ns = not significant; *p < 0.05; **p < 0.005; ***p < 0.0005].
Supplementary Figure 4 Identification of IFNG 3′ UTR ARE-binding proteins ZFP36L1 and ZFP36L2.
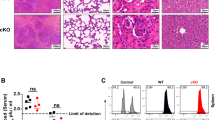
(a-c) Volcano plots represent RBPs quantified by mass spectrometry with a pull-down from human T cell lysates using 4xS1m mRNA aptamers expressing 189nt of the WT human IFNG 3’UTR (4xS1m-WT), the ARE mutant (4xS1m-MUT1-5) or the empty 4xS1m mRNA control. RBPs were eluted by on bead digestion using 50 mM Ammonium Bicarbonate solution (pH 8) containing 1 M urea and 5 mM DTT. Samples were further processed as described in ‘4xS1m RNA aptamer-protein pull-down’. Dotted lines indicate the cutoff of ± 1.75 fold change. (d) Heat map depicts Z-scored log2 LFQ values of proteins that were significantly enriched in the pull down shown in Fig. 4, and present in all three replicates of the second pull-down with fold change > 1.75. (e) ZFP36L1, ZFP36L2 and RhoGDI expression of naive CD44lowCD62LhiCD8+ T cells and memory-like CD44hiCD8+ T cells sorted from spleens of C57BL/6 J mice. CD44hi T cells were left untreated or stimulated for 2 h with PMA/ionomycin.
Supplementary Figure 5 Distribution of TM subsets in wild-type and Zfp36l2cKO mice, and mRNA expression levels in wild-type and IFN-γ-ARE-Del T cells.
(a) Relative distribution of CD8+ (left) and CD4+ (right) TEFF, TEM and TCM cells from 8-week-old Zfp36lcKO mice and wild-type littermates. (b-e) Tnf (b-c) and 18 S (d-e) mRNA decay in in vitro generated resting T cells (b,d), and in memory-like CD44hiCD8+ OTI cells (c,e) upon treatment with 1μg/ml ActD for indicated time points. Results ± SD are pooled from 4 independently performed experiments. (f-g) ActD treatment and analysis of Ifng mRNA decay of CD44hiCD8+ T cells or CD44hiCD4+ T cells from IFN-γ-ARE-Del (g) or Zfp36l2cKO (h) mice and their respective wild-type controls, as in Fig. 5g-h. (h) miR-29a, miR-29b, tbx21 and eomes mRNA expression levels of sorted CD44 hiCD8+ OTI cells analyzed by RT-PCR. Results ± SD are pooled from two independently performed experiments [Unpaired Student t-test; n = 4 mice].
Supplementary Figure 6 Protein expression of T cells treated with ActD and CHX, and analysis of ribosome-bound mRNA.
(a) Percentage of live cells (defined as Near-IR-) and Geo-MFI of CD44 and CD8 expression of splenic CD44hi sorted from wild-type and IFN-γ-ARE-Del OTI cells, that were incubated for 4h with BFA alone (-), or with BFA together with 1μg/ml ActD or 10μg/ml CHX. (b) Total RNA distribution as determined by the RNAnano Chip assay of the cytosolic fraction of CD44hi T cells that were pre-treated or not with 20 mM EDTA, and that precipitated through a 20% sucrose cushion. (c,d) Tnf (c) and 18 S (d) mRNA levels analyzed before (input) and after (ribosomes) sucrose cushion of cytoplasmic RNA of CD44hiCD8+ wild-type and IFN-γ-ARE-Del OTI cells. Results were pooled from 3 independently performed experiments (mean ± SD). [Unpaired Student t-test; n = 3–6 mice; ns = not significant].
Supplementary Figure 7 ZFP36L2 expression and function of CD44hi memory-like T cells.
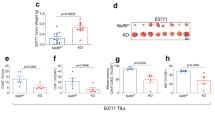
(a) Heat map represents Z-scored log2 LFQ values of proteins that were significantly upregulated or downregulated in the proteomics analysis depicted in Fig. 7a. Gray indicates no detection of peptides in the mass spectrometry analysis. Red stars depict proteins encoded by ARE-containing transcripts (Hierarchical clustering, k-means = 4). (b) Bar graphs display log2-normalized counts of transcripts encoding the proteins in panel a. Red indicates ARE-containing transcripts. Of note, Gm3839 and Mtnd4 gene expression was missing in the RNAseq data set (n.a.). (c) Zfp36l2 mRNA expression of CD44hiCD8+ or CD44hiCD4+ T cells from C57BL/6 J mice directly ex vivo, or upon 2 h PMA/ionomycin stimulation [Unpaired Student t-test; *p < 0.005; **p < 0.0005]. (d) Western blot analysis of ZFP36L2 and GAPDH expression in wild-type and in Zfp36l2cKO CD44hiCD8+ T cells that were stimulated for indicated time with PMA + ionomycin, or left untreated. Data are pooled (c) or representative of (d) 4 mice and 3 independently performed experiments.
Supplementary Figure 8 Flow cytometry gating strategies.
Representative gating strategies of spleen-derived OTI cells. T cell analysis from blood, liver, and bone marrow was performed in a similar manner. (a) Analysis of GFP+CD45.1+CD8+ OTI cells at day 35 post LM-OVA infection (as in Fig. 1). (b) Naive wild-type/CD45.1 and IFN-γ-ARE-Del/CD45.2 OTI cells were co-transferred into recipient mice. IFN-γ production at day 35 post LM-OVA infection (as in Fig. 3). (c) Expression levels of effector and memory markers of transferred OTI cells 48 days upon LM-OVA infection. (d) Example of sorting strategy of naive CD44lowCD62Lhi and memory-like CD44hi CD4+ or CD8+ T cells.
Supplementary information
Supplementary Figures
Supplementary Figures 1–8 and Supplementary Table 1
Rights and permissions
About this article
Cite this article
Salerno, F., Engels, S., van den Biggelaar, M. et al. Translational repression of pre-formed cytokine-encoding mRNA prevents chronic activation of memory T cells. Nat Immunol 19, 828–837 (2018). https://doi.org/10.1038/s41590-018-0155-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-018-0155-6
This article is cited by
-
Single-B cell analysis correlates high-lactate secretion with stress and increased apoptosis
Scientific Reports (2024)
-
An integrated proteome and transcriptome of B cell maturation defines poised activation states of transitional and mature B cells
Nature Communications (2023)
-
Post-transcriptional checkpoints in autoimmunity
Nature Reviews Rheumatology (2023)
-
RBP–RNA interactions in the control of autoimmunity and autoinflammation
Cell Research (2023)
-
Dynamic chromatin accessibility licenses STAT5- and STAT6-dependent innate-like function of TH9 cells to promote allergic inflammation
Nature Immunology (2023)