Abstract
To tackle the complexity of cross-reactive and pathogen-specific T cell responses against related Salmonella serovars, we used mass cytometry, unbiased single-cell cloning, live fluorescence barcoding, and T cell–receptor sequencing to reconstruct the Salmonella-specific repertoire of circulating effector CD4+ T cells, isolated from volunteers challenged with Salmonella enterica serovar Typhi (S. Typhi) or Salmonella Paratyphi A (S. Paratyphi). We describe the expansion of cross-reactive responses against distantly related Salmonella serovars and of clonotypes recognizing immunodominant antigens uniquely expressed by S. Typhi or S. Paratyphi A. In addition, single–amino acid variations in two immunodominant proteins, CdtB and PhoN, lead to the accumulation of T cells that do not cross-react against the different serovars, thus demonstrating how minor sequence variations in a complex microorganism shape the pathogen-specific T cell repertoire. Our results identify immune-dominant, serovar-specific, and cross-reactive T cell antigens, which should aid in the design of T cell–vaccination strategies against Salmonella.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Change history
12 March 2019
In the version of this article initially published, the first affiliation lacked ‘MRC’; the correct name of the institution is ‘MRC Weatherall Institute of Molecular Medicine’. Two designations (SP110Y and ST110H) were incorrect in the legend to Fig. 6f,h,i. The correct text is as follows: for panel f, “...loaded with either the CdtB(105–125)SP110Y (DRB4*SP110Y) or the CdtB(105–125)ST110H (DRB4*ST110H) peptide variants...”; for panel h, “...decorated by the DRB4*SP110Y tetramer (lower-right quadrant), the DRB4*ST110H (upper-left quadrant)...”; and for panel i, “...stained ex vivo with DRB4*SP110Y, DRB4*ST110H...”. In Fig. 8e, the final six residues (LTEAFF) of the sequence in the far right column of the third row of the table were missing; the correct sequence is ‘CASSYRRTPPLTEAFF’. In the legend to Fig. 8d, a designation (HLyE) was incorrect; the correct text is as follows: “(HlyE?).” Portions of the Acknowledgements section were incorrect; the correct text is as follows: “This work was supported by the UK Medical Research Council (MRC) (MR/K021222/1) (G.N., M.A.G., A.S., V.C., A.J.P.),...the Oxford Biomedical Research Centre (A.J.P., V.C.),...and core funding from the Singapore Immunology Network (SIgN) (E.W.N.) and the SIgN immunomonitoring platform (E.W.N.).” Finally, a parenthetical element was phrased incorrectly in the final paragraph of the Methods subsection “T cell cloning and live fluorescence barcoding”; the correct phrasing is as follows: “...(which in all cases included HlyE, CdtB, Ty21a, Quailes, NVGH308, and LT2 strains and in volunteers T5 and T6 included PhoN)...”. Also, in Figs. 3c and 4a, the right outlines of the plots were not visible; in the legend to Fig. 3, panel letter ‘f’ was not bold; and in Fig. 8f, ‘ND’ should be aligned directly beneath DRB4 in the key and ‘ND’ should be removed from the diagram at right, and the legend should be revised accordingly as follows: “...colors indicate the HLA class II restriction (gray indicates clones for which restriction was not determined (ND)). Clonotypes are grouped on the basis of pathogen selectivity (continuous line), protein specificity (dashed line) and epitope specificity; for ten HlyE-specific clones (pixilated squares), the epitope specificity was not determined...”. The errors have been corrected in the HTML and PDF versions of the article.
References
Crump, J. A., Luby, S. P. & Mintz, E. D. The global burden of typhoid fever. Bull. World Health Organ. 82, 346–353 (2004).
Crump, J. A. & Mintz, E. D. Global trends in typhoid and paratyphoid fever. Clin. Infect. Dis. 50, 241–246 (2010).
Mogasale, V. et al. Burden of typhoid fever in low-income and middle-income countries: a systematic, literature-based update with risk-factor adjustment. Lancet Glob. Health 2, e570–e580 (2014).
Dougan, G. & Baker, S. Salmonella enterica serovar Typhi and the pathogenesis of typhoid fever. Annu. Rev. Microbiol. 68, 317–336 (2014).
Gal-Mor, O., Boyle, E. C. & Grassl, G. A. Same species, different diseases: how and why typhoidal and non-typhoidal Salmonella enterica serovars differ. Front. Microbiol. 5, 391 (2014).
McClelland, M. et al. Comparison of genome degradation in Paratyphi A and Typhi, human-restricted serovars of Salmonella enterica that cause typhoid. Nat. Genet. 36, 1268–1274 (2004).
Holt, K. E. et al. Pseudogene accumulation in the evolutionary histories of Salmonella enterica serovars Paratyphi A and Typhi. BMC Genomics 10, 36 (2009).
Coward, C. et al. The effects of vaccination and immunity on bacterial infection dynamics in vivo. PLoS Pathog. 10, e1004359 (2014).
Griffin, A. J. & McSorley, S. J. Development of protective immunity to Salmonella, a mucosal pathogen with a systemic agenda. Mucosal Immunol. 4, 371–382 (2011).
Lee, S. J., Dunmire, S. & McSorley, S. J. MHC class-I-restricted CD8 T cells play a protective role during primary Salmonella infection. Immunol. Lett. 148, 138–143 (2012).
Dunstan, S. J. et al. Variation at HLA-DRB1 is associated with resistance to enteric fever. Nat. Genet. 46, 1333–1336 (2014).
Waddington, C. S. et al. An outpatient, ambulant-design, controlled human infection model using escalating doses of Salmonella Typhi challenge delivered in sodium bicarbonate solution. Clin. Infect. Dis. 58, 1230–1240 (2014).
McCullagh, D. et al. Understanding paratyphoid infection: study protocol for the development of a human model of Salmonella enterica serovar Paratyphi A challenge in healthy adult volunteers. BMJ Open 5, e007481 (2015).
Blohmke, C. J. et al. Interferon-driven alterations of the host’s amino acid metabolism in the pathogenesis of typhoid fever. J. Exp. Med. 213, 1061–1077 (2016).
Levine, J. H. et al. Data-driven phenotypic dissection of aml reveals progenitor-like cells that correlate with prognosis. Cell 162, 184–197 (2015).
Krutzik, P. O. & Nolan, G. P. Fluorescent cell barcoding in flow cytometry allows high-throughput drug screening and signaling profiling. Nat. Methods 3, 361–368 (2006).
Haghjoo, E. & Galán, J. E. Salmonella typhi encodes a functional cytolethal distending toxin that is delivered into host cells by a bacterial-internalization pathway. Proc. Natl. Acad. Sci. USA 101, 4614–4619 (2004).
Song, J., Gao, X. & Galán, J. E. Structure and function of the Salmonella Typhi chimaeric A(2)B(5) typhoid toxin. Nature 499, 350–354 (2013).
Liang, L. et al. Immune profiling with a Salmonella Typhi antigen microarray identifies new diagnostic biomarkers of human typhoid. Sci. Rep. 3, 1043 (2013).
Rodriguez-Rivera, L. D., Bowen, B. M., den Bakker, H. C., Duhamel, G. E. & Wiedmann, M. Characterization of the cytolethal distending toxin (typhoid toxin) in non-typhoidal Salmonella serovars. Gut Pathog. 7, 19 (2015).
von Rhein, C. et al. ClyA cytolysin from Salmonella: distribution within the genus, regulation of expression by SlyA, and pore-forming characteristics. Int. J. Med. Microbiol. 299, 21–35 (2009).
Oscarsson, J. et al. Characterization of a pore-forming cytotoxin expressed by Salmonella enterica serovars typhi and paratyphi A. Infect. Immun. 70, 5759–5769 (2002).
Barat, S. et al. Immunity to intracellular Salmonella depends on surface-associated antigens. PLoS Pathog. 8, e1002966 (2012).
Stubbington, M. J. T. et al. T cell fate and clonality inference from single-cell transcriptomes. Nat. Methods 13, 329–332 (2016).
Lindestam Arlehamn, C. S. et al. Memory T cells in latent Mycobacterium tuberculosis infection are directed against three antigenic islands and largely contained in a CXCR3+CCR6+ Th1 subset. PLoS Pathog. 9, e1003130 (2013).
Bhuiyan, S. et al. Cellular and cytokine responses to Salmonella enterica serotype Typhi proteins in patients with typhoid fever in Bangladesh. Am. J. Trop. Med. Hyg. 90, 1024–1030 (2014).
Reynolds, C. J. et al. The serodominant secreted effector protein of Salmonella, SseB, is a strong CD4 antigen containing an immunodominant epitope presented by diverse HLA class II alleles. Immunology 143, 438–446 (2014).
Sheikh, A. et al. Interferon-γ and proliferation responses to Salmonella enterica serotype Typhi proteins in patients with S. Typhi bacteremia in Dhaka, Bangladesh. PLoS Negl. Trop. Dis. 5, e1193 (2011).
Wahid, R., Fresnay, S., Levine, M. M. & Sztein, M. B. Cross-reactive multifunctional CD4+ T cell responses against Salmonella enterica serovars Typhi, Paratyphi A and Paratyphi B in humans following immunization with live oral typhoid vaccine Ty21a. Clin. Immunol. 173, 87–95 (2016).
Fuentes, J. A., Villagra, N., Castillo-Ruiz, M. & Mora, G. C. The Salmonella Typhi hlyE gene plays a role in invasion of cultured epithelial cells and its functional transfer to S. Typhimurium promotes deep organ infection in mice. Res. Microbiol. 159, 279–287 (2008).
Galán, J. E. Typhoid toxin provides a window into typhoid fever and the biology of Salmonella Typhi. Proc. Natl. Acad. Sci. USA 113, 6338–6344 (2016).
Groisman, E. A. The pleiotropic two-component regulatory system PhoP-PhoQ. J. Bacteriol. 183, 1835–1842 (2001).
Dobinson, H. C. et al. Evaluation of the clinical and microbiological response to Salmonella paratyphi a infection in the first paratyphoid human challenge model. Clin. Infect. Dis. 64, 1066–1073 (2017).
Wong, M. T. et al. Mapping the diversity of follicular helper T cells in human blood and tonsils using high-dimensional mass cytometry analysis. Cell Reports 11, 1822–1833 (2015).
Finck, R. et al. Normalization of mass cytometry data with bead standards. Cytometry A 83, 483–494 (2013).
Chen, H. et al. Cytofkit: a Bioconductor package for an integrated mass cytometry data analysis pipeline. PLOS Comput. Biol. 12, e1005112 (2016).
Kotecha, N., Krutzik, P. O. & Irish, J. M. Web-based analysis and publication of flow cytometry experiments. Curr. Protoc. Cytom. 53, 10.17.1–10.17.24 (2010).
Traunecker, A., Oliveri, F. & Karjalainen, K. Myeloma based expression system for production of large mammalian proteins. Trends Biotechnol. 9, 109–113 (1991).
Olsen, L. R. et al. BlockLogo: visualization of peptide and sequence motif conservation. J. Immunol. Methods 400-401, 37–44 (2013).
Bolotin, D. A. et al. MiXCR: software for comprehensive adaptive immunity profiling. Nat. Methods 12, 380–381 (2015).
Shugay, M. et al. Towards error-free profiling of immune repertoires. Nat. Methods 11, 653–655 (2014).
Shugay, M. et al. VDJtools: unifying post-analysis of T cell receptor repertoires. PLOS Comput. Biol. 11, e1004503 (2015).
Acknowledgements
This work was supported by the UK Medical Research Council (MRC) (MR/K021222/1) (G.N., M.A.G., A.S., V.C., A.J.P.), Cancer Research UK (CRUK) (C399/A2291) (V.C.), the NIHR Clinical Research Network Thames Valley (A.J.P.), the Oxford Biomedical Research Centre (A.J.P., V.C.), the Bill & Melinda Gates Foundation (OPP1084259) (A.J.P.), the European Vaccine Initiative (PIM) (A.J.P.), and core funding from the Singapore Immunology Network (SIgN) (E.W.N.) and the SIgN immunomonitoring platform (E.W.N.). G.D. is supported by The Wellcome Trust. We thank C. Waugh and the WIMM FACS facility for assistance with cell sorting, and T. Rostron for assistance with next-generation sequencing. The DRB4 tetramers were provided by B. Kwok and I.-T. Chow, from the Tetramer Core Laboratory at the Benaroya Research Institute at Virginia Mason.
Author information
Authors and Affiliations
Contributions
G.N. conceived and designed the study, processed the samples, performed and supervised the experiments, analyzed the data, and wrote the manuscript. P.K. processed the samples, performed experiments, and analyzed the data. L.H., P.d.H., L.P.-L., L.S., and M.S. assisted with experiments. A.A. designed and performed bacteria–macrophage coculture experiments. M.R. and D.M.S.P. optimized and performed TCR-repertoire experiments and analysis. K.W.W.T., E.B., M.T.W., and E.W.N. developed the metal-conjugated antibody panels, coordinated and performed the mass cytometry experiments, and provided assistance with the analysis of mass cytometry experiments. A.J.P. conceived the development of the human-challenge model and directed the clinical trial. H.D. and M.M.G. designed and directed the clinical trial. D.C., C.J., and C.J.B. coordinated the clinical trial and sampling. C.J., H.T.-B., and H.B.J. coordinated sampling and processed samples. L.R.O. designed peptide pools containing overlapping peptides. S.B., G.D., D.B., and D.P. provided reagents. G.N., M.A.G., A.S., V.C., and A.J.P. coordinated the study. V.C. conceived and designed the study, edited the manuscript, and supervised performing of experiments. All authors discussed the results and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Characterization of Ki67+ cells accumulating during enteric fever.
(a)Gating strategy to identify CD4+ and CD4+Ki67+ cells in Mass Cytometry experiments. Frequency of Cluster 19 cells within Ki67+ (b) and total CD4+ T cells (c) at different time points after infection (n = 6). (b) Friedman Test p = 0.0147, Dunn’s Multiple Comparison Test D0 vs D4, D28, D90 p = ns, D0 vs EF p = 0.027. (c)Friedman Test p = 0.0159, Dunn’s Multiple Comparison Test D0 vs D4, D28, D90 p = ns, D0 vs EF p = 0.0247. (d) Total Ki67+ events (grey) and Cluster 19 events (red) across all samples were concatenated and expression of 20 markers against CD38 in the distinct populations is shown.
Supplementary Figure 2 CCR7–CD38+CD4+ cells are enriched in Ki67+ accumulating during enteric fever.
(a) Expression of CCR7 and CD38 in the 20 distinct clusters identified by PhenoGraph analysis. (b) Proportion of PhenoGraph clusters within Ki67+ CD38+CCR7- cells at the peak of Ki67+ cell accumulation. (c) Frequency of Ki67+ CD38+CCR7- cells (Friedman Test p = 0.0130, Dunn’s Multiple Comparison Test D0 vs D4, D28, D90p = ns, D0 vs EF p = 0.0139) and of KI67 + cells non comprised within CD38+CCR7- cells over time in challenged volunteers (n = 6) (Friedman Test p = 0.23, Dunn’s Multiple Comparison Test D0 vs D4, EF, D28, D90 p = ns). Box extends from the 25th to the 75th percentile and whiskers from the minimum to the maximum value; line indicates median.
Supplementary Figure 3 Accumulation of CD4+CD38+CCR7– in whole blood of volunteers with enteric fever
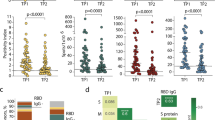
(a) Identification of CD4+CD38+CCR7- cells during enteric fever by Multicolour Flow Cytometry in whole blood from a representative volunteer challenged with S. Typhi, and one challenged with S. Paratyphi A. (b) Accumulation of CD38+CCR7- cells after diagnosis in volunteers challenged with S. Typhi and Paratyphi A who developed (Diagnosed) but not in those who did not develop enteric fever (Not Diagnosed). In challenged volunteers not diagnosed blood was collected 14 days after challenge (D14). Friedman Multi comparison Test p = 0.006 (S. Typhi), p < 0.0001 (S. Paratyphi). Dunn’s Multiple Comparison Test ED vs D0, p = 0.0065 (Typhi, n = 9); p = 0.0002 (S. Paratyphi A, n = 9). Box extends from the 25th to the 75th percentile and whiskers from the minimum to the maximum value; line indicates median.
Supplementary Figure 4 Gating strategy for live fluorescence barcoding of T cell clones
Shown is an example with four live fluorescent dyes. Cell were labelled with 9 combinations of three distinct dilution of Violet and Far red dyes. Cells with each of the nine cluster were labelled with 6 combinations of 4 distinct dilutions of CFSE and and 3 of CMTMR orange dyes. Dilutions used for adjacent clusters across the 640-670 axis were labelled with alternate dilutions of CFSE and CMTMR Orange (as shown in the overlaid plots) in order to reduce spill-over in case of suboptimal separation between clusters across the channel-dye combination with the lowest stain index.
Supplementary Figure 5 Increased frequency of HlyE- and CdtB- specific T cells in volunteers diagnosed with enteric fever
(a) Frequency of HlyE, CdtB and Flu specific T cell responses 28 days after challenge measured by IFN-g ELISPOT in volunteers challenged with S. Typhi or S. Paratyphi A who developed (Diagnosed, D, n = 12) or did not develop (Not Diagnosed, ND, n = 9)) enteric fever compared to uninfected controls (HC. n = 10). Freshly isolated PBMC were rested overnight and incubated with 5μg/ml of HlyE or CdtB peptide pools, or with inactivated seasonal influenza vaccine (Flu). Mann Witney two tailed t-test. * = p < 0.05; ** = p < 0.005. Box extends from the 25th to the 75th percentile and whiskers from the minimum to the maximum value; line indicates median.
Supplementary Figure 6 CdtB-specific clones help macrophages limit bacterial spread
(a) Monocyte derived macrophages (MF) were differentiated for 5 days in the lower chamber of 24 well transwell plates. At day 5 a pool of 3 CdtB specific clones (T) was added to the upper well in the presence or absence of CdtB peptide pool (CdtB). After O.N. incubation, upper well was removed, macrophages were washed, infected with S. Typhimurium (LT2 strain) for 30 m, washed again and incubated with Gentamicin (30 mg/ml) to limit bacterial overgrowth. 2 h after infection cell culture supernatant was plated in agar plates and colonies counted after over night culture. Indicated is the percentage of bacterial growth compared to macrophages alone (n = 6, Friedman test (p = 0.0008) with Dunn’s multiple comparisons test (control vs T + CdtB p = 0.0169). Box extends from the 25th to the 75th percentile and whiskers from the minimum to the maximum value; line indicates median. (b) CFU after O.N. culture of macrophage supernatant in agar plate in one representative experiment.
Supplementary Figure 7 Increased frequency of CdtB, HlyE, and PhoN T cell responses after diagnosis with enteric fever
(a) Frequency of HlyE, CdtB, and Flu specific T cell responses at baseline and 28 days after challenge measured by IFN-g ELISPOT in frozen PBMC from volunteers challenged with S. Typhi or S. Paratyphi A who developed (n = 19) enteric fever (Wilcoxon matched-pairs signed rank test). (b) Frequency of HlyE, CdtB, PhoN and Flu specific T cell responses in HLADRB4 + and HLADRB4- volunteers 28 days after challenge (Mann Whitney test two-tailed p value). (c) Frequency of PhoN specific T cell responses at baseline and 28 days after challenge measured by IFN-g ELISPOT in frozen PBMC from volunteers challenged with S. Typhi or S. Paratyphi A who developed (n = 19) enteric fever and in HLADRB4+ and HLADRB4- individuals. ELISPOT analysis done on the same frozen samples as in panel a and b.
Supplementary Figure 8 Clonal expansion of Salmonella-specific effector T cells
(a) Rarefaction curve describing the estimated diversity of the CDR3β repertoire of CD38+CCR7- and CD38-CCR7- subsets in volunteers P1 and T4. Dots between continuous and dashed curves indicate the number of TCRs identified within each sample. Number of cells probed: P1 CD38+CCR7- = 5087 cells, CD38-CCR7- = 1.15e105 cells; T4 CD38+CCR7- = 9130 cells, CD38-CCR7- = 1.3e105 cells. (b) Pathogen selectivity, antigen specificity, restriction, CDR3α and CDR3β sequence of clones isolated from volunteer P1. Starred(*) sequences were identified also in the polyclonal analysis. (c) Pie Chart depicting the frequency of CDR3β sequences within the polyclonal repertoire of CD38+CCR7- cells (5087 cells probed) from T4, highlighted are the CDR3β sequences identified in the isolated clones. (d) Frequency of CDR3β within the polyclonal repertoire of CD38+CCR7- cells and CD38-CCR7- cells in volunteer P1 (1.15e105 cells probed). Red circles represent CDR3β sequences identified in the isolated clones.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8 and Supplementary Tables 1–8
Supplementary Note
Peptide pools
Rights and permissions
About this article
Cite this article
Napolitani, G., Kurupati, P., Teng, K.W.W. et al. Clonal analysis of Salmonella-specific effector T cells reveals serovar-specific and cross-reactive T cell responses. Nat Immunol 19, 742–754 (2018). https://doi.org/10.1038/s41590-018-0133-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-018-0133-z
This article is cited by
-
Resistance is futile? Mucosal immune mechanisms in the context of microbial ecology and evolution
Mucosal Immunology (2022)
-
An overview of immunoinformatics approaches and databases linking T cell receptor repertoires to their antigen specificity
Immunogenetics (2020)
-
Invasive Salmonella exploits divergent immune evasion strategies in infected and bystander dendritic cell subsets
Nature Communications (2018)