Abstract
The adaptor CARD9 functions downstream of C-type lectin receptors (CLRs) for the sensing of microbial infection, which leads to responses by the TH1 and TH17 subsets of helper T cells. The single-nucleotide polymorphism rs4077515 at CARD9 in the human genome, which results in the substitution S12N (CARD9S12N), is associated with several autoimmune diseases. However, the function of CARD9S12N has remained unknown. Here we generated CARD9S12N knock-in mice and found that CARD9S12N facilitated the induction of type 2 immune responses after engagement of CLRs. Mechanistically, CARD9S12N mediated CLR-induced activation of the non-canonical transcription factor NF-κB subunit RelB, which initiated production of the cytokine IL-5 in alveolar macrophages for the recruitment of eosinophils to drive TH2 cell–mediated allergic responses. We identified the homozygous CARD9 mutation encoding S12N in patients with allergic bronchopulmonary aspergillosis and revealed activation of RelB and production of IL-5 in peripheral blood mononuclear cells from these patients. Our study provides genetic and functional evidence demonstrating that CARD9S12N can turn alveolar macrophages into IL-5-producing cells and facilitates TH2 cell–mediated pathologic responses.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Gross, O. et al. Card9 controls a non-TLR signalling pathway for innate anti-fungal immunity. Nature 442, 651–656 (2006).
Jia, X. M. et al. CARD9 mediates Dectin-1-induced ERK activation by linking Ras-GRF1 to H-Ras for antifungal immunity. J. Exp. Med. 211, 2307–2321 (2014).
Glocker, E. O. et al. A homozygous CARD9 mutation in a family with susceptibility to fungal infections. N. Engl. J. Med. 361, 1727–1735 (2009).
Lanternier, F. et al. Deep dermatophytosis and inherited CARD9 deficiency. N. Engl. J. Med. 369, 1704–1714 (2013).
Rosentul, D. C. et al. Genetic variation in the dectin-1/CARD9 recognition pathway and susceptibility to candidemia. J. Infect. Dis. 204, 1138–1145 (2011).
Rosentul, D. C. et al. Gene polymorphisms in pattern recognition receptors and susceptibility to idiopathic recurrent vulvovaginal candidiasis. Front. Microbiol. 5, 483 (2014).
Venselaar, H., Te Beek, T. A., Kuipers, R. K., Hekkelman, M. L. & Vriend, G. Protein structure analysis of mutations causing inheritable diseases. An e-Science approach with life scientist friendly interfaces. BMC Bioinformatics 11, 548 (2010).
Bertin, J. et al. CARD9 is a novel caspase recruitment domain-containing protein that interacts with BCL10/CLAP and activates NF-κB. J. Biol. Chem. 275, 41082–41086 (2000).
Ramensky, V., Bork, P. & Sunyaev, S. Human non-synonymous SNPs: server and survey. Nucleic Acids Res. 30, 3894–3900 (2002).
Franke, A. et al. Genome-wide meta-analysis increases to 71 the number of confirmed Crohn’s disease susceptibility loci. Nat. Genet. 42, 1118–1125 (2010).
McGovern, D. P. et al. Genome-wide association identifies multiple ulcerative colitis susceptibility loci. Nat. Genet. 42, 332–337 (2010).
Janse, M. et al. Three ulcerative colitis susceptibility loci are associated with primary sclerosing cholangitis and indicate a role for IL2, REL, and CARD9. Hepatology 53, 1977–1985 (2011).
Burghardt, K. M. et al. A CARD9 polymorphism is associated with decreased likelihood of persistent conjugated hyperbilirubinemia in intestinal failure. PLoS One 9, e85915 (2014).
Rivas, M. A. et al. Deep resequencing of GWAS loci identifies independent rare variants associated with inflammatory bowel disease. Nat. Genet. 43, 1066–1073 (2011).
Ader, F. et al. Invasive pulmonary aspergillosis in chronic obstructive pulmonary disease: an emerging fungal pathogen. Clin. Microbiol. Infect. 11, 427–429 (2005).
Rivera, A. et al. Dectin-1 diversifies Aspergillus fumigatus-specific T cell responses by inhibiting T helper type 1 CD4 T cell differentiation. J. Exp. Med. 208, 369–381 (2011).
Greenberger, P. A. Allergic bronchopulmonary aspergillosis. J. Allergy Clin. Immunol. 110, 685–692 (2002).
Kauffman, H. F. Immunopathogenesis of allergic bronchopulmonary aspergillosis and airway remodeling. Front. Biosci. 8, e190–e196 (2003).
Knutsen, A. P. et al. Increased sensitivity to IL-4 in cystic fibrosis patients with allergic bronchopulmonary aspergillosis. Allergy 59, 81–87 (2004).
Gavino, C. et al. CARD9 deficiency and spontaneous central nervous system candidiasis: complete clinical remission with GM-CSF therapy. Clin. Infect. Dis. 59, 81–84 (2014).
Steele, C. et al. The β-glucan receptor dectin-1 recognizes specific morphologies of Aspergillus fumigatus. PLoS Pathog. 1, e42 (2005).
Loures, F. V. et al. Recognition of Aspergillus fumigatus hyphae by human plasmacytoid dendritic cells is mediated by dectin-2 and results in formation of extracellular traps. PLoS Pathog. 11, e1004643 (2015).
Fei, M. et al. TNF-α from inflammatory dendritic cells (DCs) regulates lung IL-17A/IL-5 levels and neutrophilia versus eosinophilia during persistent fungal infection. Proc. Natl. Acad. Sci. USA 108, 5360–5365 (2011).
Voehringer, D., Shinkai, K. & Locksley, R. M. Type 2 immunity reflects orchestrated recruitment of cells committed to IL-4 production. Immunity 20, 267–277 (2004).
Gessner, A., Mohrs, K. & Mohrs, M. Mast cells, basophils, and eosinophils acquire constitutive IL-4 and IL-13 transcripts during lineage differentiation that are sufficient for rapid cytokine production. J. Immunol. 174, 1063–1072 (2005).
Hogan, M. B., Piktel, D. & Landreth, K. S. IL-5 production by bone marrow stromal cells: implications for eosinophilia associated with asthma. J. Allergy Clin. Immunol. 106, 329–336 (2000).
Toussaint, M. et al. Host DNA released by NETosis promotes rhinovirus-induced type-2 allergic asthma exacerbation. Nat. Med. 23, 681–691 (2017).
Rivera, A. et al. Innate immune activation and CD4+ T cell priming during respiratory fungal infection. Immunity 25, 665–675 (2006).
Karta, M. R., Broide, D. H. & Doherty, T. A. Insights into group 2 Innate lymphoid cells in human airway disease. Curr. Allergy Asthma Rep. 16, 8 (2016).
Jhingran, A. et al. Tracing conidial fate and measuring host cell antifungal activity using a reporter of microbial viability in the lung. Cell Rep. 2, 1762–1773 (2012).
Hara, H. & Saito, T. CARD9 versus CARMA1 in innate and adaptive immunity. Trends Immunol. 30, 234–242 (2009).
Hailfinger, S. et al. Malt1-dependent RelB cleavage promotes canonical NF-κB activation in lymphocytes and lymphoma cell lines. Proc. Natl. Acad. Sci. USA 108, 14596–14601 (2011).
Chauhan, B., Knutsen, Ap, Hutcheson, P. S., Slavin, R. G. & Bellone, C. J. T cell subsets, epitope mapping, and HLA-restriction in patients with allergic bronchopulmonary aspergillosis. J. Clin. Invest. 97, 2324–2331 (1996).
Chauhan, B. et al. The association of HLA-DR alleles and T cell activation with allergic bronchopulmonary aspergillosis. J. Immunol. 159, 4072–4076 (1997).
Risma, K. A. et al. V75R576 IL-4 receptor alpha is associated with allergic asthma and enhanced IL-4 receptor function. J. Immunol. 169, 1604–1610 (2002).
Brouard, J. et al. Influence of interleukin-10 on Aspergillus fumigatus infection in patients with cystic fibrosis. J. Infect. Dis. 191, 1988–1991 (2005).
Saxena, S., Madan, T., Shah, A., Muralidhar, K. & Sarma, P. U. Association of polymorphisms in the collagen region of SP-A2 with increased levels of total IgE antibodies and eosinophilia in patients with allergic bronchopulmonary aspergillosis. J. Allergy Clin. Immunol. 111, 1001–1007 (2003).
Miller, P. W. et al. Cystic fibrosis transmembrane conductance regulator (CFTR) gene mutations in allergic bronchopulmonary aspergillosis. Am. J. Hum. Genet. 59, 45–51 (1996).
Faccioli, L. H. et al. IL-5 drives eosinophils from bone marrow to blood and tissues in a guinea-pig model of visceral larva migrans syndrome. Mediators Inflamm. 5, 24–31 (1996).
Dubucquoi, S. et al. Interleukin 5 synthesis by eosinophils: association with granules and immunoglobulin-dependent secretion. J. Exp. Med. 179, 703–708 (1994).
Plaut, M. et al. Mast cell lines produce lymphokines in response to cross-linkage of FcεRI or to calcium ionophores. Nature 339, 64–67 (1989).
Takeda, K. et al. Development of eosinophilic airway inflammation and airway hyperresponsiveness in mast cell-deficient mice. J. Exp. Med. 186, 449–454 (1997).
Wilson, S. J., Shute, J. K., Holgate, S. T., Howarth, P. H. & Bradding, P. Localization of interleukin (IL)-4 but not IL-5 to human mast cell secretory granules by immunoelectron microscopy. Clin. Exp. Allergy 30, 493–500 (2000).
Hamelmann, E. et al. Allergen-specific IgE and IL-5 are essential for the development of airway hyperresponsiveness. Am. J. Respir. Cell Mol. Biol. 16, 674–682 (1997).
Robinson, D. S. Mepolizumab for severe eosinophilic asthma. Expert Rev. Respir. Med. 7, 13–17 (2013).
Walsh, G. M. Profile of reslizumab in eosinophilic disease and its potential in the treatment of poorly controlled eosinophilic asthma. Biologics 7, 7–11 (2013).
Ghazi, A., Trikha, A. & Calhoun, W. J. Benralizumab–a humanized mAb to IL-5Rα with enhanced antibody-dependent cell-mediated cytotoxicity–a novel approach for the treatment of asthma. Expert Opin. Biol. Ther. 12, 113–118 (2012).
Stranick, K. S. et al. Identification of transcription factor binding sites important in the regulation of the human interleukin-5 gene. J. Biol. Chem. 272, 16453–16465 (1997).
Mori, A. et al. p38 mitogen-activated protein kinase regulates human T cell IL-5 synthesis. J. Immunol. 163, 4763–4771 (1999).
Bailey, E. et al. FLT3/D835Y mutation knock-in mice display less aggressive disease compared with FLT3/internal tandem duplication (ITD) mice. Proc. Natl. Acad. Sci. USA 110, 21113–21118 (2013).
Yang, H. et al. One-step generation of mice carrying reporter and conditional alleles by CRISPR/Cas-mediated genome engineering. Cell 154, 1370–1379 (2013).
Hsu, Y. M. S. et al. The adaptor protein CARD9 is required for innate immune responses to intracellular pathogens. Nat. Immunol. 8, 198–205 (2007).
Gersuk, G. M., Underhill, D. M., Zhu, L. & Marr, K. A. Dectin-1 and TLRs permit macrophages to distinguish between different Aspergillus fumigatus cellular states. J. Immunol. 176, 3717–3724 (2006).
Zhao, X. et al. JNK1 negatively controls antifungal innate immunity by suppressing CD23 expression. Nat. Med. 23, 337–346 (2017).
Shizuru, J. A., Taylor-Edwards, C., Banks, B. A., Gregory, A. K. & Fathman, C. G. Immunotherapy of the nonobese diabetic mouse: treatment with an antibody to T-helper lymphocytes. Science 240, 659–662 (1988).
Veillette, A., Bookman, M. A., Horak, E. M. & Bolen, J. B. The CD4 and CD8 T cell surface antigens are associated with the internal membrane tyrosine-protein kinase p56lck. Cell 55, 301–308 (1988).
Sauer, K. A., Scholtes, P., Karwot, R. & Finotto, S. Isolation of CD4+ T cells from murine lungs: a method to analyze ongoing immune responses in the lung. Nat. Protoc. 1, 2870–2875 (2006).
Zhao, X. Q. et al. C-type lectin receptor dectin-3 mediates trehalose 6,6′-dimycolate (TDM)-induced Mincle expression through CARD9/Bcl10/MALT1-dependent nuclear factor (NF)-κB activation. J. Biol. Chem. 289, 30052–30062 (2014).
Acknowledgements
This work was supported by the National Natural Science Foundation of China (31622023 and 81571611 to X.M.J, 81630058 and 91542107 to X.L.), Outstanding academic leader program of Shanghai health and Family Planning Commission (2017BR024 to X.M.J.),Shanghai laboratory animal research fund (16140902600 to X.M.J), Shuguang Program of Shanghai Education Development Foundation and Shanghai Municipal Education Commission (17SG24 to X.M.J.),a start-up fund from the Tsinghua University-Peking University Joint Center for Life Sciences (with grants from Tsinghua University to X.L.) and Shanghai Leading Talent Program (2016036 to J.F.X).
Author information
Authors and Affiliations
Contributions
X.X., G.Z., J.-L.D., W.R., J.-H.G., Q.-Z.L., J.-X.L. and C.-X.C. performed experiments; J.-F.X. and H.-W.L. collected samples from patients; L.-Q.C., D.-D.Y., M.-C.W. and X.Z. contributed critical reagents; P.S. performed statistical analysis; X.X., J.-F.X.., X-M.J. and X.L. designed the study; and X.X, X-M. J., and X.L. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated Supplementary Information
Supplementary Figure 1 Characterization of CARD9S12N knock-in mice.
(a) Raw data of sequencing CARD9 gene amplified from the genome of F0 and F1 offsprings. (b) Raw data of sequencing CARD9 gene amplified from the genome of WT, heterozygous (Het) and homozygous CARD9S12N KI mice. (c) CARD9 protein levels in BMDMs from WT and KI mice. (d) ELISA results for TNF, IL-6 and IL-12p40 in the supernatant of BMDMs from WT, Het and KI mice, which were stimulated with Af SC (MOI = 5) for 6 hours. (e) CFU assay in lungs of WT and KI mice at day 2 and 6 after single exposure to Af conidia (1×107). *P < 0.05, **P < 0.01 and ***P < 0.001, by one-way ANOVA and post hoc Tukey test (d) and two-tailed unpaired t-test (e). Data shown are representative of three independent experiments.
Supplementary Figure 2 Characterization of type-2 responses in wide-type and CARD9S12N KI mice in chronic asthma models.
(a) Strategy of developing chronic murine asthma model, in which WT and KI mice were sensitized with 5×106 Af conidia, challenged with 1×106 Af conidia for eight times, and sacrificed for sequent assay at day 18. (b–e) Eosinophil (SiglecF+, b and c) and Neutrophil (Ly6G+, d and e) counts in lungs of the above disposed mice. (f) ELISA results for lung IL-4 and IL-5 and serum IgE in the above disposed mice. *P < 0.05, **P < 0.01 and ***P < 0.001, by two-tailed unpaired t-test (c, e-f). Data shown are representative of three independent experiments.
Supplementary Figure 3 Determining T cell responses in wide-type and CARD9S12N KI mice in response to Af challenging.
(a) Gating strategy for flow assay of Th2 (CD3+CD4+ICOS+ST2+) cell recruitment in the lungs of WT and CARD9S12N KI mice at day 6 after single intratracheal exposure to Af conidia (1×107). (b) Gating strategy for flow assay of T-bet and GATA3 proteins in CD3+CD4+T cells from the lungs of the above disposed mice. (c) Gating strategy for flow assay of IL-4-producing CD3+CD4+T cells in the mediastinal lymph node (MLN) from the above disposed mice. (d) Gating strategy for flow assay of IL-17A-producing and IFN-γ-producing CD4+T cells in the MLN of the above disposed mice.
Supplementary Figure 4 Depletion of CD4+ T cell in CARD9S12N KI mice.
(a and b) T cell counts in lungs of KI mice, which were intravenously injected with 200 μg anti-CD4 or control IgG per mouse at 24 hours before challenge and assayed at day 6 after single intratracheal exposure to Af conidia (1×107). (c and d) Neutrophil (Ly6G+) and eosinophil (SiglecF+) counts in lungs of the above disposed mice. *P < 0.05, **P < 0.01 and ***P < 0.001. By one-way ANOVA and post hoc Tukey test. Data shown are representative of three independent experiments.
Supplementary Figure 5 ILC2s decreased in WT and CARD9S12N KI mice after infection.
(a) Gating strategy for flow assay of ILC2 (CD90.2+Lin-RORγt-GATA-3+) counts in lungs from WT and KI mice at day 2 after single intratracheal exposure to Af conidia (1×107). (b and c) ILC2 counts in the lung of the above disposed mice. *P < 0.05; **P < 0.01 and ***P < 0.001. Data shown are representative of three independent experiments. By two-tailed unpaired t-test (c), in a-c, n = 6 mice per group.
Supplementary Figure 6 Determining cellular responses in wide-type and CARD9S12N KI mice in response to Af challenging.
(a) Gating strategy for flow assay of alveolar macrophage (AM, CD11c+SiglecF+), eosinophil (SiglecF+) and neutrophil (Ly6G+) counts in lungs of WT, Het and KI mice at the indicated day after single intratracheal exposure to Af conidia (1 × 107). (b, c) AM, eosinophil and neutrophil counts in lungs of the above disposed mice.
Supplementary Figure 7 Intracellular staining for IL-4 expression in wide-type and CARD9S12N KI mice.
(a) Gating strategy of flow assay for sorting alveolar macrophage (CD11c+ SiglecF+) in lungs of WT and CARD9S12N KI mice at day 2 after single exposure to Af conidia (1 × 107). (b) Gating strategy of flow assay for sorting DCs (CD11c+, Q1), alveolar macrophage (CD11c+SiglecF+, Q2), CD11c-SiglecF-cells (Q3) and eosinophil (SiglecF+, Q4) in lungs of the above disposed mice. (c) Quantitative real-time PCR results for IL-4 expression normalized to GAPDH in DCs and AMs sorted from the above disposed mice at day 2. Data are mean ± s.e.m. *P < 0.05, **P < 0.01 and ***P < 0.001, by two-tailed t-test. Data shown are representative of three independent experiments.
Supplementary Figure 8 IL-5-producing cells in ABPA patients.
(a) Gating strategy and (b) raw data of flow assay for IL-5-producing PBMCs from ABPA patients carrying wild-type or homozygous S12N mutation of CARD9, which were determined by flow cytometry.
Supplementary Figure 9 Nuclear translocation of RelB in PBMCs from ABPA patients.
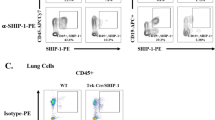
(a) Gating strategy and (b) raw data of RelB nuclear translocation in PBMCs (CD14+CD11b+ cells) from ABPA patients carrying wild-type or homozygous S12N mutation of CARD9, which were determined by Millipore-Amnis FlowSight.
Supplementary Figure 10 Nuclear translocation of p65 in PBMCs from ABPA patients.
(a) Gating strategy and (b) raw data of p65 nuclear translocation in PBMCs (CD14+CD11b+ cells) from ABPA patients carrying wild-type or homozygous S12N mutation of CARD9, which were determined by Millipore-Amnis FlowSight.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–10 and Supplementary Tables 1–3
Rights and permissions
About this article
Cite this article
Xu, X., Xu, JF., Zheng, G. et al. CARD9S12N facilitates the production of IL-5 by alveolar macrophages for the induction of type 2 immune responses. Nat Immunol 19, 547–560 (2018). https://doi.org/10.1038/s41590-018-0112-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-018-0112-4
This article is cited by
-
Eosinophils promote pulmonary matrix destruction and emphysema via Cathepsin L
Signal Transduction and Targeted Therapy (2023)
-
TRIM41 is required to innate antiviral response by polyubiquitinating BCL10 and recruiting NEMO
Signal Transduction and Targeted Therapy (2021)
-
Pathway paradigms revealed from the genetics of inflammatory bowel disease
Nature (2020)
-
Rapid generation and selection of Cas9-engineering TRP53 R172P mice that do not have off-target effects
BMC Biotechnology (2019)
-
CARD–BCL-10–MALT1 signalling in protective and pathological immunity
Nature Reviews Immunology (2019)