Abstract
Legionella pneumophila (Lp) is a flagellated, intracellular bacterium that can cause Legionnaires’ disease (LD). Lp activates multiple innate immune receptors, and TOLLIP dampens MyD88-dependent signaling and may influence susceptibility to LD. We evaluated the effect of TOLLIP on innate immunity, pneumonia severity, and LD susceptibility in mouse lungs and human populations. To accomplish this, we evaluated the effect of TOLLIP on lung-specific Lp control and immune response and associated a common functional TOLLIP variant with Lp-induced innate immune responses and LD susceptibility in humans. After aerosol Lp infection, Tollip−/− mice demonstrated significantly fewer bacterial colony-forming unit and increased cytokine responses from BAL fluid. Tollip−/− macrophages also suppressed intracellular Lp replication in a flagellin-independent manner. The presence of a previously characterized, functionally active SNP associated with decreased TOLLIP mRNA transcript in monocytes was associated with increased TNF and IL-6 secretion after Lp stimulation of PBMC ex vivo. This genotype was separately associated with decreased LD susceptibility (309 controls, 88 cases, p = 0.008, OR 0.36, 95% CI 0.16–0.76) in a candidate gene association study. These results suggest that TOLLIP decreases lung-specific TLR responses to increase LD susceptibility in human populations. Better understanding of TOLLIP may lead to novel immunomodulatory therapies.
Similar content being viewed by others
Introduction
An outbreak of severe pneumonia at a Philadelphia American Legion convention in 1976 led to the identification of Legionella spp. as a cause of pneumonia.1 Legionella pneumonia ranges from a mild illness to a severe multifocal pneumonia termed Legionnaires’ disease (LD) with significant mortality.2 In mammals, Lp infects alveolar macrophages after inhalation into the deep airways.3 Lp is unique from other pulmonary pathogens; it contains flagellin, displays LPS that avoids detection by TLR4, and secretes multiple immunoregulatory proteins into the cytosol of infected macrophages to impair host immune responses.4,5,6 To survive intracellularly, Lp alters the phagolysosome within the macrophage, creating the Legionella-containing vacuole (LCV).6 Lp recruits proteins to the LCV that inhibit macrophage proinflammatory responses and permit its intracellular survival.7,8 IL-1β and TNF produced by infected and bystander macrophages induce antibacterial defense by activating and recruiting macrophages to the infected site9,10. Macrophages recognize Lp via multiple pathogen-recognition receptors, including TLR2, TLR5, NLRC4, and NAIP5 in mice.11,12,13,14,15 Understanding the factors that influence signaling activity of these pathways may provide insight into unique mechanisms of host resistance and susceptibility to infection.
Toll-Interacting Protein (TOLLIP) regulates many innate immune processes that influence Lp immunity, including TLR2 and IL-1R, autophagy, and endosomal transport.16,17,18,19 In humans, common genetic variation in the TOLLIP gene is associated with susceptibility to multiple intracellular infections.20,21,22 How TOLLIP influences infectious disease host defense within the lung is not well understood. We hypothesized that TOLLIP, acting as a ubiquitin-binding autophagy receptor, negatively regulates the innate immune response after Lp infection, which results in increased susceptibility to LD.
Host genetic variation influences innate immune responses and susceptibility to many infections.23 Functional polymorphisms in TLR5 and STING are associated with altered cytokine responses and susceptibility to LD, but assessing genetic associations are challenging given the episodic nature of pneumonia outbreaks.24,25 In prior studies, we identified a functionally active SNP at rs5743854 that is associated with decreased TOLLIP mRNA expression in monocytes and increased TNF after TLR2 stimulation.20,26 In this paper, we characterize effect of this variant on Lp-specific macrophage immune responses in a cohort of healthy volunteers and evaluate the genetic association of this variant on LD susceptibility in a second well-characterized cohort from the Netherlands. Overall, using a mouse model, cellular studies, human cellular immunology, and a case–control population study of highly exposed individuals, we herein evaluate the role of TOLLIP on Lp pathogenesis and critical host immune mechanisms of control.
Methods
Human Subjects
Approval for human study protocols was obtained from the human subjects’ review boards at the University of Amsterdam Medical Center and the University of Washington Medical Center. All participants gave written informed consent. Genomic DNA was purified form peripheral blood leukocytes from 10 ml of blood. Enrollment of the cases and controls from a LD outbreak at a flower show in the Netherlands was described previously.25,27 Of the 188 cases identified in the original investigation of the flower show outbreak, 141 consented for the study. Eighteen individuals died and no DNA was available for genotyping. Eighty-eight cases were available with both DNA and epidemiologic data for TOLLIP genotyping. Control subjects (N = 309) were drawn from the exhibitioners who worked at the same flower show and were at high risk for exposure to Legionella pneumophila (Lp). Studies in Seattle included 62 healthy volunteers who donated peripheral blood for immunogenetic analysis. The ethnic composition of these subjects was 73% Caucasian (45/62) and 27% East Asian (17/62).
Genotyping
Genomic DNA was isolated from whole blood and genotyping of selected SNPs was performed using a Fluidigm Biomark 96 × 96 chip (Fluidigm, Inc.). Cluster plots were visually inspected to ensure accurate genotyping calls. SNPs were manually assessed for data quality and only high-quality calls were accepted. Eighty-eight cases and 309 controls had high-quality genotyping data available for analysis. Genotypes were assessed for Hardy–Weinberg equilibrium (HWE) with a Chi-square test comparing observed and expected frequencies in the control population. No SNPs violated HWE (P < 0.001).
Genetic analysis
We assessed the genetic association between the TOLLIP rs5743854 G/G genotype and LD with a recessive genetic model using Stata 13 (Stata Corp, College Station, TX) and the user-written package “genass.”28
Reagents
Roswell Park Memorial Institute (RPMI) 1640 and Dulbecco's Modified Eagle Medium (DMEM) was purchased from Invitrogen (Carlsbad, CA.). Ultrapure LPS was purified from Salmonella minnesota R595 (List Biological Laboratories). Flagellin C (FliC) was purified from Salmonella typhimurium strain TH4778, which is flijB-/fliC+.29 Flagellin A (FlaA) was purified from L. pneumophila Corby strain by the same method with bacteria grown as described below on BCYE agar plates. Bone marrow was harvested from mice and grown in DMEM supplemented with 10% heat-inactivated fetal calf serum (FCS; Atlas Bio, Fort Collins, CO) and 20% LCL-conditioned medium. Bone marrow-derived macrophages (BMDM) were used after 4–10 days of culture. Bronchoalveolar fluid was obtained by intratracheal lavage with cold phosphate-buffered saline; PBS (10 ml). Cells were then plated in RPMI 1640 supplemented with 10% heat-inactivated FCS. After a 4 h adherence step, the cells were washed six times with PBS before stimulation immediately or the next day.
Microbiology
Lp was cultured in BCYE medium was prepared with α-ketoglutarate.30 Bacteria were inoculated from a frozen stock onto BCYE agar for 4 days at 35°C. Bacterial cultures used for cellular stimulation or replication assays were placed in RPMI culture medium supplemented with 10% FCS and thymidine. Aerosol infection experiments with WT Lp used Lp Philadelphia-01 strain (ATCC 33152). Macrophage experiments with mutant Lp used the Lp serogroup 1 JR32-derived (rpsLK88R and hsdR) strain lacking either flaA (ΔflaA Lp) or dot/icm locus (ΔdotA Lp).31,32
Knockout mouse generation and screening
Embryos from B6.Cg-Tolliptm1Kbns/Cnrm (Tollip−/−) mice were obtained from the European Mutant Mouse Archive (www.infrafrontier.eu), which were generated by Didierlaurent et al.33 Mice were backcrossed eight times on C57BL/6J background and were confirmed to be > 95% C57BL/6J genetically by screening 150 SNP ancestry informative markers (Jax Genome Scanning Services, Jax, Inc). Genotyping was performed using DNA primers for neomycin (Forward sequence: AGGATCTCCTGTCATCTCACCTTGCTCCTG; Reverse sequence AAGAACTCGTCAAGAAGGCGATAGAAGGCG) and the first exon of TOLLIP (Forward sequence: AGCTACTGGGAGGCCATACA; Reverse sequence: CGTGTACGGGAGACCCATTT). TOLLIP protein expression was confirmed in both knockout and control mice by western blot (Supplemental Fig. 1). TOLLIP antibodies for western blotting were polyclonal (#4748, Cell Signaling). All wild-type control mice were age-matched littermates of Tollip−/− mice to ensure a common genetic background.
Model of pneumonic legionellosis
The University of Washington Institutional Animal Care and Use Committee approved all animal protocols. Buffered charcoal yeast extract (BCYE) medium was prepared with alpha-ketoglutarate as described elsewhere.25 Bacteria were inoculated from a frozen stock onto BCYE agar for 4 days at 35 °C, harvested by rinsing plates with PBS, pelleted by centrifugation, and resuspended in PBS to a concentration of 1010 cfu/ml (estimated by optical density at 540 nm and confirmed by quantitative culture).25 The mice were exposed to aerosolized bacteria in a whole animal exposure chamber (Biaera Technologies, Hagerstown, MD) with aerosols generated by a mini-Heart Hi-Flo jet nebulizers (Westmed, Tucson, AZ).34 Bacterial deposition in the lungs was determined from quantitative culture of homogenized lung tissue harvested from mice killed with pentobarbital and exsanguinated immediately after aerosol exposure. At subsequent time points, mice were killed and exsanguinated for tissue harvests. The trachea was cannulated and the right lung was lavaged with 0.85% saline/0.6mml/L ethylenediaminetetraacetic acid to determine cell counts and differentials. The left lung was homogenized in PBS and serially diluted in Mueller–Hinton broth for quantitative culture on BCYE agar. The remaining lung homogenate was mixed 1:1 with lysis buffer containing 2 × protease inhibitor mixture, incubated for 30 min on ice, clarified by centrifugation at 2500 rpm, and then the supernatant was saved at − 80 °C.
Histology
To prepare organs for histology, the lung was inflated to 15 cm pressure with 4% paraformaldehyde, fixed in the same solution, embedded in paraffin, and then 4 µm sections were generated. Sections stained with hematoxylin and eosin were examined by a pathologist blinded to mouse genotype.
Cytokine analysis
Multiplex cytokine analysis was performed as follows: samples of WT and Tollip−/− mouse bronchoalveolar lavage fluid were thawed on ice and centrifuged at 10,000 g for 5–10 min to remove particulates. In total, 50 µL of each sample was added in triplicate to a 96-well plate. Samples were then added to 50 µL of magnetic Simplex beads and incubated with shaking for 1 h at room temperature. Beads were washed 3 × and stained with detection antibody cocktail (IFNγ; IL-12p70; IL-13; IL-1β; IL-2; IL-4; IL-5; IL-6; TNF; GM-CSF; IL-18; IL-10; IL-17A; IL-22; IL-23; IL-27; IL-9; GROα; IP-10; MCP-1; MCP-3; MIP-1α; MIP-1β; MIP-2; RANTES; Eotaxin; ThermoFisher Scientific) and incubated with shaking for 30 min at room temperature. Beads were washed 3 × and incubated with streptavidin-HRP with shaking for 30 min at RT. Beads with sample cytokines and detection antibody were resuspended in 120 µL of Reading Buffer and cytokine data was acquired immediately on Luminex 100/200. Analytes were captured by Ab-coated, fluorochrome-embedded microspheres and detected by biotin–streptavidin–phycoerythrin using reagents purchased from R&D or Luminex. For single cytokine analysis, concentrations were determined with enzyme-linked immunosorbent assay (ELISA; R&D Systems) according to the manufacturer’s instructions.
Western blotting
Cell extracts were prepared from BMDM by lysis with 100 µl Cell Extraction Buffer (Life Technologies) along with protease inhibitors (Thermo Scientific). Lysates were stored at − 20 °C. Western blotting of lysates was performed on 4–15% polyacrylamide gels and transferred onto PVDF membrane. Antibodies included anti-TOLLIP antibody (Pierce PA5-30747) at 1:1000 dilution, anti-GAPDH (Sigma, catalog A2066) at 1:2000 dilution, and anti-rabbit IgG-HRP secondary antibody (Sigma, catalog A0545) at 1:2000 dilution.
Statistics
For in vivo colony-forming unit (CFU) count analysis, comparisons between groups were made by the Mann–Whitney test without presupposition of normality of the data. For mouse experiments, groups of five mice were compared with one another unless otherwise indicated. Cytokine measurements and cell counts were compared using a two-sided Students’ t test. A value of p < 0.05 was considered a statistically significant result. Statistics were calculated using Prism version 7.1 (GraphPad, Inc.).
Results
Aerosolized lung infection of mice with Lp
To understand the role of TOLLIP in Lp pathogenesis, we re-derived C57Bl/6 mice lacking the Tollip gene from embryos and confirmed lack of TOLLIP expression by Western blot (Supplemental Fig. 1). We infected groups of five WT and five Tollip−/− mice with Lp Philadelphia-1 strain and measured lung CFU at 0, 24, 72, and 144 h after infection. Tollip−/− mice demonstrated significantly fewer CFU in the lung compared with WT littermates at 72 and 144 h after infection (Fig. 1a, mean WT 4.3 × 105 CFU, mean Tollip−/− 8.5 × 104 CFU at 72 h, p = 0.014). We also examined the recruitment of neutrophils and mononuclear cells to the lungs after infection in these mice. We collected bronchoalveolar lavage (BAL) fluid and measured cell counts at 24, 72, and 144 h after infection. We did not detect significant differences in cell counts in Tollip−/− mice at 24 h after infection, but by 72 h after infection significantly fewer polymorphonuclear were present in BAL samples harvested from Tollip−/− lungs. (Fig. 1b, WT mean 2.2 × 106 cells/ml, mean Tollip−/− 1.3 × 106 cells/ml, p = 0.046). Similarly, monocyte counts were increased in WT mice compared with Tollip−/− mice after 72 h of infection (Fig. 1c, mean WT 9.4 × 105 cells/ml, mean Tollip−/− 6.4 × 105 cells/ml, p = 0.018). After 72 h, Tollip−/− mice demonstrated reduced airspace filling and leukocyte infiltration (Fig. 1d) compared with controls (Fig. 1e) by histopathology observations. In summary, Tollip−/− mice demonstrated accelerated bacterial clearance and decreased pulmonary inflammation after inhalation of Lp compared with WT controls.
TOLLIP-deficient mice demonstrate enhanced clearance of L. pneumophila (Lp) after aerosol infection. Mice were infected with approximately 106 CFU WT Lp Philadelphia-1 strain and at 24, 72, and 144 h post infection, lungs were harvested. a Bacterial counts were measured by CFU on BCYE agar plates. b Total polymorphonuclear cell counts from bronchoalveolar fluid. c Total mononuclear cells from bronchoalveolar fluid samples. d Hematoxylin and eosin staining of representative sections of d WT and e Tollip−/− mouse lung 72 h after Lp aerosol infection. *p < 0.05, Student’s t test, two sided. Error bars represent ± SEM. N = 5 mice in each group at each time point. Data are representative of three independent experiments
In vivo cytokine responses after Lp infection
Given TOLLIP’s known role regulating innate immune responses, we next evaluated whether TOLLIP influenced lung cytokine concentrations after Lp infection. 24 h after aerosol infection with 106 CFU Lp, we killed mice (n = 5 per group) and measured cytokine responses from BAL fluid and serum. Tollip−/− mice demonstrated significantly increased levels of multiple cytokines critical for Lp control in BAL fluid, including IL-1β (Fig. 2a; p = 0.001), TNF (p = 0.025), IL-12p40 (p = 0.001), and IL-18 (p = 0.04) by ELISA. We also evaluated a panel of 26 cytokines from BAL fluid in WT and Tollip−/− mice after Lp infection by multiplex array. We found that Tollip−/− mice produce significantly more IFNγ, IL-12p70, IL-18, IL-6, TNF, IL-22, IL-17A, Eotaxin, IL-27, and IP-10 (p < 0.05 with false discovery rate <1%, Fig. 2b). We detected significantly decreased MIP-2 in Tollip−/− mice, but no differences in GM-CSF, IL-23, IL-9, αKC, MCP-1, MCP-3, MIP-1α, and MIP-1β. IL-13, IL-2, IL-4, IL-5, and IL-10 cytokine concentrations were below the limit of detection (data not shown). We did not detect statistically significant differences between WT and Tollip−/− cytokine concentrations in serum (Fig. 2c). In summary, Tollip−/− mice demonstrated broadly increased cytokine responses from BAL fluid but not in serum after Lp infection.
Cytokine responses from WT and Tollip−/− mice after L. pneumophila (Lp) aerosol infection. Mice were infected with 106 CFU WT Lp Philadelphia-1 strain and after 24 h, cytokine concentrations were measured from bronchoalveolar (BAL) fluid and serum. a BAL cytokine concentrations measured by ELISA from WT and Tollip−/− mice. *p < 0.05, Student’s two-sided t test. b BAL cytokines, measured by Luminex cytokine assay 24 h after Lp aerosol infection. *p < 0.05, **p < 0.01, ***p < 0.001, at 1% false discovery rate. Error bars represent±SEM. c Serum cytokine concentrations measured by ELISA. *p < 0.05, Student’s two-sided t test. Error bars represent±SEM. Data are representative of three independent experiments with N = 5 in each group
Pattern-recognition receptor (PRR) responses and TOLLIP
Macrophages are essential reservoirs for Lp in vivo.35 TOLLIP downregulates PRR signaling in macrophages, however, and which specific PRR signaling complexes explain the improved control of Lp infection is not known. We hypothesized that multiple PRRs that are necessary for Lp control, including TLR2, TLR4, NLRC4, and NLRP3, are influenced by TOLLIP in macrophages. To evaluate the role of TOLLIP on TLR signaling, we stimulated WT and Tollip−/− peritoneal macrophages (PEM) with PAM3 (TLR2/1 agonist) or LPS (TLR4 agonist) for 24 h, then measured TNF and IL-10 cytokine concentrations from cell culture supernatants by ELISA. Tollip−/− macrophages secreted increased concentrations of TNF after PAM3 (Fig. 3a, p = 0.002) or LPS stimulation (Fig. 3a, p = 0.01) when compared with WT macrophages. Conversely, Tollip−/− PEM secreted significantly less IL-10 than WT PEM after 24 h' stimulation with either PAM3 (p = 0.019) or LPS (p = 0.012, Fig. 3b).
WT and Tollip−/− macrophage cytokine responses after TLR and NLR stimulation. Bone marrow-derived macrophages (BMDM) were isolated from WT and Tollip−/− mice and stimulated with media, PAM3 (250 ng/ml), a TLR2/1 agonist, or LPS (10 ng/ml), TLR4 agonist, for 24 hr, and concentrations of a TNF and b IL-10 were measured from cellular supernatants. c We stimulated mouse BMDM with LPS 100 pg/ml for 2 h, then stimulated overnight with c media control or the NLRP3 ligand nigericin (100 µg/ml), and d NLRC4 ligand Salmonella flagellin (FliC, 100 ng/ml) or bovine serum albumin (100 ng/ml), coincubated with Lipofectamine 2000 to permit cytosolic translocation of each protein. IL-1β concentrations from cellular supernatants were measured by ELISA. Error bars are means ± SEM; statistical analyses were performed by paired t tests using Prism 7.0e. *p < 0.05, Student’s two-sided t test. The experiment shown is representative of three independent experiments, each conducted in triplicate
We also evaluated the effect of TOLLIP deficiency on NLRP3- and NLRC4-induced cytokine responses. We stimulated WT and Tollip−/− BMDM with 100 pg/ml LPS, followed 2 h later by nigericin (NLRP3 ligand; 10 µg/mL). We found that BMDM isolated from Tollip−/− mice demonstrated significantly increased IL-1β release when compared with BMDM from WT mice (Fig. 3c, p = 0.026). Next, we compared IL-1β secretion from BMDM transfected with flagellin (NLRC4 ligand) via coincubation with cationic lipids as compared with bovine serum albumin-transfected BMDM controls. As with nigericin, we found that Tollip−/− BMDM secreted significantly more IL-1β after overnight incubation (Fig. 3d, p = 0.005). No BMDM, from either WT or Tollip−/− mice, produced detectable TNF or IL-6 after flagellin stimulation without transfection (50 ng/ml, data not shown), which is consistent with prior data demonstrating that murine BMDM do not express the flagellin receptor TLR5 on the cell surface.36 These data demonstrate that TOLLIP-deficient macrophages induce increased TNF and IL-1β after TLR and NLR stimulation, responses that may contribute to the in vivo phenotype observed in Fig. 1.
Cytokine responses to live Lp macrophage infection
Lp influences immune activation in macrophages, so we tested the hypothesis that cytokine responses in TOLLIP-deficient macrophages were increased after live Lp infection in vitro. We plated 105 BMDM from WT and Tollip−/− mice and infected them at MOI 10 with wild-type Lp, ΔflaA Lp, or ΔdotA Lp that lacks the Type IV secretion system, making them incapable of intracellular replication, and collected supernatants after 24 h’ infection. As compared with WT BMDM controls, BMDM lacking TOLLIP secreted significantly more TNF after infection with WT (Fig. 4a, p = 0.025), ΔflaA Lp (Fig. 4a, p = 0.004), and ΔdotA Lp (Fig. 4a, p = 0.018). Further, IL-1β was increased in Tollip−/− BMDM after WT Lp (Fig. 4b, p = 0.047) and ΔflaA Lp (Fig. 4b, p = 0.002) infection, but no IL-1β was detected after ΔDotA Lp incubation. IL-6 secretion was not significantly different between WT and Tollip−/− BMDM after infection with any of the Lp strains tested (Fig. 4c). These data show that TOLLIP deletion induces more TNF and IL-1β after infection with WT or ΔflaA Lp, suggesting that flagellin is not necessary for altered Lp cytokine responses by TOLLIP.
WT and Tollip−/− macrophage cytokine responses after L. pneumophila (Lp) Infection in vitro. Bone marrow-derived macrophages were isolated and infected with either negative control (mock), WT L. pneumophila (WT Lp), flagellin-deficient L. pneumophila (ΔflaA Lp), or dot-icm-locus-deficient L. pneumophila (ΔdotA Lp) on the JR32 background at MOI 10 overnight and a TNF, b IL-1β, and c IL-6 concentrations were measured from cellular supernatants. *p < 0.05, Student’s two-sided t test. Experiment shown is representative of three independent experiments, each performed in triplicate. Error bars represent ±SEM of mean
Intracellular replication in murine macrophages
Although TOLLIP influences the innate immune response to Lp, many of these responses may be induced by bystander macrophages. Further, Lp induces an immune signature by inhibiting host protein translation.37 We assessed overall control of Lp within macrophages by measuring Lp intracellular replication in macrophages. We infected WT and Tollip−/− BMDM with ΔflaA Lp (MOI 0.02), which is replication-competent in macrophages.38 Tollip−/− macrophages markedly suppressed replication ΔflaA Lp in comparison with WT BMDM (Fig. 5a, p < 0.001). These data demonstrate that Lp replication is suppressed within Tollip−/− macrophages.
Intracellular L. pneumophila (Lp) replication and nonspecific autophagy in WT and Tollip−/− Macrophages. a Bone marrow-derived macrophages (BMDM) from WT and Tollip−/− mice were infected with flagellin-deficient L. pneumophila (JR32 strain) expressing the luminescent lux gene. Relative luminescence was measured over time. b Representative images depicting LC3 + organelles (green) and DAPI-stained nuclei (blue) from control (EV) or THP-1 cells with TOLLIP deleted via CRISPR-Cas9 gene editing (TOLLIP-KO) in RPMI supplemented with 10% FCS (ctrl) or EBSS (starve) for 4 h. c Tukey plot of the number of autophagosomes counted from 100 cells in EV and TOLLIP-KO cells in nutrient-enriched or starvation medium after 4 h. Box demonstrates median and interquartile range of values. *p < 0.05, Mann–Whitney U test, data are representative of three independent experiments. d Western blot of THP-1 cells comparing LC3-II expression with β-actin as a control in TOLLIP-KO and control cell lines under nutrient-rich and starvation conditions, with or without chloroquine. e Quantitative analysis of LC3-II expression, normalized to β-actin, in TOLLIP-deficient cells in control and TOLLIP-KO cells. Data are expressed as mean±SD of three separate experiments grouped together. *p < 0.05 by Student’s t test
Nonselective autophagy in macrophages
The mechanism by which TOLLIP influences Lp-specific cytokine responses and controls intracellular replication is not clear. TOLLIP is an autophagy receptor and influences clearance of lipids from atherosclerotic plaques and insoluble protein aggregates in brain tissue.18,39 Macrophages use autophagy for direct antimicrobial killing, but also to dampen immune responses.25 However, the role of TOLLIP on nonspecific autophagy in macrophages has not been established. We cultured THP-1 cells with TOLLIP deleted (TOLLIP-KO, Supplemental Fig. 2) using CRISPR-Cas9 gene editing, along with empty vector control (EV) cells, for 4 h in a nutrient rich (RPMI supplemented with 10% FCS) or starvation (EBSS) medium. Under typical conditions, starvation inhibits mTOR, a constitutive inhibitor of autophagic flux, leading to transient induction and organization of LC3-II, which forms the structural backbone of the autophagosome and attracts autophagy receptors to coordinate fusion with the lysosome.40 Starvation of EV cells for 4 h induced increased numbers of LC3+ autophagosomes (Fig. 5b, c, p < 0.001) compared with starvation of TOLLIP-KO cells. There were no significant differences in the number of autophagosomes present in EV and TOLLIP-KO cells in nutrient-rich conditions. Furthermore, EV cells, but not TOLLIP-KO cells incubated in starvation medium (EBSS) for 4 h demonstrated significantly more LC3-II protein expression than EV cells incubated with nutrient-rich media (RPMI; Fig. 5d, e, p < 0.05). Addition of chloroquine, a known inhibitor of lysosomal fusion and LC3 degradation, led to increased LC3-II levels in both EV and TOLLIP-KO cells (Fig. 5d, e). We did not detect differences in ubiquitin staining between EV and TOLLIP-KO cells (Supplemental Fig. 3). These data suggest that TOLLIP deficiency leads to reduced autophagic flux in monocytes and offers insight into the potential role of TOLLIP on specific autophagy functions.
TOLLIP SNP rs5743854 is associated with increased cytokine responses to live Lp
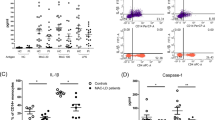
In prior studies, we characterized a functionally active promoter-region SNP in the TOLLIP gene region, rs5743854, that is associated with decreased TOLLIP expression in monocytes.41 We hypothesized that macrophages from individuals with this SNP would be associated with increased TNF responses after WT Lp and ΔflaA Lp infection. We isolated peripheral blood mononuclear cells from 62 healthy volunteers in the Seattle area and stimulated them with live WT and ΔflaA Lp (MOI 1) overnight, then measured IL-6 and TNF concentrations in cellular supernatants. Individuals with rs5743854 G/G genotype demonstrated increased TNF responses to WT Lp (Fig. 6a p = 0.013; C/C = 34, G/C = 24, G/G = 4) but not ΔflaA Lp (Fig. 6b, p = 0.82) when compared with those with either C/C or G/C genotype. Similarly, the same individuals demonstrated increased IL-6 responses to WT Lp (Fig. 6c, p = 0.0007) and ΔflaA Lp (Fig. 6d, p = 0.037). Thus, the presence of a functionally active common genetic variant that is associated with decreased TOLLIP mRNA expression in monocytes is also associated with increased TNF secretion after Lp infection.
rs5743854 G/G allele is associated with increased proinflammatory cytokine secretion after L. pneumophila (Lp) infection. Peripheral blood mononuclear cells were isolated from 62 healthy volunteers in Seattle and stimulated with media, WT Lp (MOI 1) or ΔflaA Lp (MOI 1) for 24 h. Secreted TNF and IL-6 levels were measured in supernatants via ELISA. TNF responses after a WT Lp or b ΔflaA Lp infection, stratified by rs5743854 genotype. Number of individuals tested per genotype−C/C = 34, G/C = 24, G/G = 4. IL-6 responses after c WT Lp or d ΔflaA Lp infection, stratified by rs5743854 genotype. Dots represent cytokine values for individual volunteers. Median value represented by red lines. *p < 0.05 by Mann–Whitney U test in a recessive genetic model (GG vs (CC + CG))
TOLLIP SNP rs5743854 is associated with decreased risk for LD
We further hypothesized that TOLLIP deficiency (genotype G/G) is associated with susceptibility to LD in humans.16,18 We performed a candidate gene case–control study on a cohort of individuals (N = 88 LD cases and 309 controls) were exposed to Lp during a flower show.26 The TOLLIP SNP rs5743854 G/G genotype was associated with decreased LD susceptibility in a recessive model (p = 0.032; OR 0.44, 95% confidence interval (CI) 0.22–0.96). After adjustment for smoking, age, gender, and comorbidities as covariates, the significance of this association increased (Table 1, p = 0.008, OR 0.36, 95% CI 0.16–0.76). We evaluated the this association in relation to other LD susceptibility SNPs in TLR525 and in STING.24 We conducted a logistic regression evaluating the association between rs5743854 G/G genotype and LD with presence of the TLR5 stop codon allele or the HAQ allele of STING as covariates. In the presence of inactivated TLR5, the association of rs5743854 G/G genotype remains robust (p = 0.009, 95% CI 0.16–0.77, but the presence of STING deficiency abolishes the association (p = 0.273, 95% CI 0.17–1.65). Overall, these data demonstrate that the presence of the TOLLIP deficiency SNP is associated with increased cytokine responses after Lp infection and decreased risk for LD after exposure in a well-characterized case–control study.
Discussion
Deletion of the TOLLIP gene in mice leads to faster clearance of Lp after aerosol infection and increased proinflammatory cytokine responses within the lungs of Lp-infected mice. Importantly, TOLLIP-deficient macrophages are resistant to Lp intracellular replication independently of flagellin, suggesting that TOLLIP acts independently of TLR5 and NLRC4 to influence immune responses. We link this finding to human immune responses and susceptibility to LD by demonstrating that a common, functionally active variant associated with TOLLIP deficiency (rs5743854 G/G genotype) is associated with increased cytokine responses to Lp and decreased risk for developing LD in a genetic case–control cohort from the Netherlands. Thus, we found that TOLLIP acts broadly to diminish innate immune responses and influence Lp pathogenesis.
We hypothesize that TOLLIP influences TLR2 signaling primarily to influence Lp disease. Lp is recognized by TLR2 (via Lp peptidoglycan-associated lipoprotein), TLR5, NLRP3, NLRC4, and AIM2. TNF is increased and IL-10 is decreased in TOLLIP-deficient macrophages after TLR2 stimulation, which both suppress intracellular Lp replication. Infection of Tollip−/− macrophages with ΔflaA Lp induces increased TNF and IL-1β and diminished intracellular replication, further supporting the critical effects of TOLLIP on TLR2 activity as sufficient for TOLLIP’s effect on Lp pathogenesis. Several questions remain, however. We demonstrate that TOLLIP acts upon TLR signaling and also influences IL-1β induction, even after infection with ΔflaA Lp. IL-1β is induced by the inflammasome, and inflammasome activation requires two signals: a priming signal, often MyD88-dependent, that upregulates expression of inflammasome components and pro-IL-1β, followed by an activation signal induced by ATP or nigericin for NLRP3, or cytosolic flagellin for NLRC4.42 TOLLIP-deficient macrophages demonstrate hyperactive induction of the first signal. TOLLIP may diminish the second signal of inflammasome activation as well. TOLLIP contains the CUE ubiquitin-binding domain, which has multiple binding partners and may thus influence signaling across multiple innate immune pathways. Other autophagy proteins, such as Atg16L, diminish inflammasome activation.43 Further, linear ubiquitin-binding accessory protein SHARPIN suppresses both TLR and inflammasome activation directly.44 Detailed study of TOLLIP’s activity after intracellular infection may provide insight into how ubiquitin signaling may influence innate immunity.
Lp avoids antimicrobial killing via multiple mechanisms, including by impairing autophagy.45 However, we found that TOLLIP-deficient macrophages demonstrated impaired starvation-induced autophagic flux, whereas simultaneously inhibiting Lp replication. To reconcile these seemingly contradictory data, we hypothesize that TOLLIP deficiency impairs the capacity for autophagy-induced immune inhibition. Depletion of critical autophagy proteins LC3 or Beclin leads to increased IL-1β production via two mechanisms: (1) decreased degradation of pro-IL-1β and (2) impaired degradation of inflammasome components.46 If TOLLIP similarly influences immune responses, then different autophagy receptors may impact specific functions. One of the challenges to using small molecule autophagy inhibitors in vivo are side effects from off target functions. Therefore, this insight may identify novel drug targets that influence autophagy innate immune responses with minimal non-immune activity.
Intriguingly, we identified that TLR2 signaling in the absence of TOLLIP induces increased TNF and proinflammatory cytokines but diminished anti-inflammatory IL-10. The mechanism of this discrepancy is unclear. First, TOLLIP may selectively suppresses proinflammatory cytokine production in the TLR signaling cascade via novel, non-IRAK-binding partners. Selective TRIF signaling, which occurs via nonclassical TLR4 activation induces Type I interferon activation followed by IL-10 induction.47 Alternately, induction of ER stress from infection in the setting TOLLIP deficiency may alter the cytokine balance in macrophages. ER stress occurs during the unfolded protein response (UPR), and a characteristic of ER stress is the selective induction of proinflammatory cytokines and inflammation.48 TOLLIP is a critical receptor for recycling insoluble protein aggregates, suggesting its importance in the UPR.18 Further understanding of how TOLLIP selectively influences proinflammatory cytokine induction will provide novel insights into immune regulation.
TOLLIP SNP rs5743854 was associated with increased cytokine responses to Lp and decreased susceptibility to LD. This finding is consistent with prior observations that the TOLLIP rs5743854 G allele was functionally active, associated with decreased TOLLIP mRNA expression in monocytes, and altered the innate and adaptive immune responses to mycobacteria.26 Currently, genetic variation in TOLLIP is also associated with susceptibility to tuberculosis, leprosy, leishmaniasis, and malaria. However, presence of the G/G allele is associated with increased risk for tuberculosis but decreased risk for LD and increased proinflammatory cytokine responses after infection with both pathogens. Given this well-characterized phenotype, we believe that the differences in susceptibility demonstrate the differential survival strategies of these two pathogens. Lp induces decreased inflammation to survive but M. tuberculosis requires inflammation in later stages of disease. Further comparative studies across diseases will improve our understanding of the role of inflammation in multiple infections.
Limitations of our study included the possible confounding of the genetic findings by heterogeneity of the population substructure.49 We performed logistic regression by self-identified ethnicity and comorbidity and report these results in this paper. Another possible limitation could be misclassification of cases and controls. However, study participants were followed for a year to determine case status. rs5743854 G/G genotype is a functionally active SNP associated with TOLLIP deficiency in monocytes26 and is associated with increased Lp-specific PBMC cytokine responses, suggesting that this data represent effects of TOLLIP deficiency. We used different Lp strains for in vivo infection experiments and macrophage assays—Philadelphia-1 and JR32 for ΔflaA and ΔdotA. These strains demonstrate some differences in the host response, apart from the presence of flagellin, including the presence of the lvh locus.50,51,52 We confirmed TOLLIP’s effect using specific ligands and live bacteria, and we controlled each infection with a comparison with WT littermates. Despite this, TOLLIP diminished TLR2-dependent activity and impaired the immune response to Lp in vivo and in human populations.
In this paper, we demonstrate that TOLLIP deficiency increases Lp clearance in macrophages and in vivo by influencing TLR2 activity. We also found that presence of a known TOLLIP deficiency genotype is associated with decreased susceptibility to LD. Further studies targeting TOLLIP as a target for therapeutics may lead to novel treatments for intracellular infections and inflammatory disorders.
References
Phin, N. et al. Epidemiology and clinical management of Legionnaires’ disease. Lancet Infect. Dis. 14, 1011–1021 (2014).
Dominguez, A. et al. Factors influencing the case-fatality rate of Legionnaires’ disease. Int J. Tuberc. Lung Dis. 13, 407–412 (2009).
Horwitz, M. A. Phagocytosis of the Legionnaires’ disease bacterium (Legionella pneumophila) occurs by a novel mechanism: engulfment within a pseudopod coil. Cell 36, 27–33 (1984).
Bornstein, N., Marmet, D., Dumaine, M. H., Surgot, M. & Fleurette, J. Detection of flagella in 278 Legionella strains by latex reagent sensitized with antiflagellum immunoglobulins. J. Clin. Microbiol. 29, 953–956 (1991).
Girard, R. et al. Lipopolysaccharides from Legionella and Rhizobium stimulate mouse bone marrow granulocytes via Toll-like receptor 2. J. Cell Sci. 116, 293–302 (2003).
Sherwood, R. K. & Roy, C. R. Autophagy evasion and endoplasmic reticulum saubversion: the Yin and Yang of legionella intracellularinfection. Annu. Rev. Microbiol. 70, 413–433 (2016).
Fernandez-Moreira, E., Helbig, J. H. & Swanson, M. S. Membrane vesicles shed by Legionella pneumophila inhibit fusion of phagosomes with lysosomes. Infect. Immun. 74, 3285–3295 (2006).
Hempstead, A. D. & Isberg, R. R. Inhibition of host cell translation elongation by Legionella pneumophila blocks the host cell unfolded protein response. Proc. Natl Acad. Sci. USA 112, E6790–E6797 (2015).
Ziltener, P., Reinheckel, T. & Oxenius, A. Neutrophil and alveolar macrophage-mediated innate immune control of Legionella pneumophila lung infection via TNF and ROS. PLoS Pathog. 12, e1005591 (2016).
Kawamoto, Y. et al. TNF-alpha inhibits the growth of Legionella pneumophila in airway epithelial cells by inducing apoptosis. J. Infect. Chemother. 23, 51–55 (2017).
Akamine, M. et al. Differential roles of Toll-like receptors 2 and 4 in in vitro responses of macrophages to Legionella pneumophila. Infect. Immun. 73, 352–361 (2005).
Molofsky, A. B. et al. Cytosolic recognition of flagellin by mouse macrophages restricts Legionella pneumophila infection. J. Exp. Med. 203, 1093–1104 (2006).
Zamboni, D. S. et al. The Birc1e cytosolic pattern-recognition receptor contributes to the detection and control of Legionella pneumophila infection. Nat. Immunol. 7, 318–325 (2006).
Hawn, T. R. et al. Altered inflammatory responses in TLR5-deficient mice infected with Legionella pneumophila. J. Immunol. 179, 6981–6987 (2007).
Zhao, Y. et al. The NLRC4 inflammasome receptors for bacterial flagellin and type III secretion apparatus. Nature 477, 596–600 (2011).
Burns, K. et al. Tollip, a new component of the IL-1RI pathway, links IRAK to the IL-1 receptor. Nat. Cell Biol. 2, 346–351 (2000).
Zhang, G. & Ghosh, S. Negative Regulation of Toll-like Receptor-mediated Signaling by Tollip. J. Biol. Chem. 277, 7059–7065 (2002).
Lu, K., Psakhye, I. & Jentsch, S. Autophagic clearance of polyQ proteins mediated by ubiquitin-Atg8 adaptors of the conserved CUET protein family. Cell 158, 549–563 (2014).
Jongsma, M. L. et al. An ER-associated pathway defines endosomal architecture for controlled cargo transport. Cell 166, 152–166 (2016).
Shah, J. A. et al. Human TOLLIP regulates TLR2 and TLR4 signaling and its polymorphisms are associated with susceptibility to tuberculosis. J. Immunol. 189, 1737–1746 (2012).
Araujo, F. J. et al. Polymorphisms in the TOLLIP gene influence susceptibility to cutaneous Leishmaniasis caused by Leishmania guyanensis in the Amazonas State of Brazil. PLoS Negl. Trop. Dis. 9, e0003875 (2015).
Shah, J. A. et al. Genetic variation in toll-interacting protein is associated with leprosy susceptibility and cutaneous expression of interleukin 1 receptor antagonist. J. Infect. Dis. 213, 1189–1197 (2016).
Casanova, J. L. & Abel, L. The genetic theory of infectious diseases: a brief history and selected illustrations. Annu Rev. Genomics. Hum. Genet. 14, 215–243 (2013).
Ruiz-Moreno, J. S. et al. The common HAQ STING variant impairs cGAS-dependent antibacterial responses and is associated with susceptibility to Legionnaires’ disease in humans. PLoS Pathog. 14, e1006829 (2018).
Hawn, T. R. et al. A common dominant TLR5 stop codon polymorphism abolishes flagellin signaling and is associated with susceptibility to legionnaires’ disease. J. Exp. Med. 198, 1563–1572 (2003).
Shah, J. A. et al. A functional TOLLIP variant is associated with BCG-specific immune responses and tuberculosis. Am. J. Respir. Crit. Care Med. 196, 502–511 (2017).
Den Boer, J. W. et al. A large outbreak of Legionnaires’ disease at a flower show, the Netherlands, 1999. Emerg. Infect. Dis. 8, 37–43 (2002).
Nahid, P. et al. Racial differences in tuberculosis infection in United States communities: the coronary artery risk development in young adults study. Clin. Infect. Dis. 53, 291–294 (2011).
Ibrahim, G. F., Fleet, G. H., Lyons, M. J. & Walker, R. A. Method for the isolation of highly purified Salmonella flagellins. J. Clin. Microbiol. 22, 1040–1044 (1985).
Edelstein, P. H. [Biological diagnosis of Legionnaire’s disease]. Rev. Mal. Respir. 1, 11–14 (1984).
Rao, C., Benhabib, H. & Ensminger, A. W. Phylogenetic reconstruction of the Legionella pneumophila Philadelphia-1 laboratory strains through comparative genomics. PLoS One 8, e64129 (2013).
Zhu, W. et al. Sensing cytosolic RpsL by macrophages induces lysosomal cell death and termination of bacterial infection. PLoS Pathog 11, e1004704 (2015).
Didierlaurent, A. et al. Tollip regulates proinflammatory responses to interleukin-1 and lipopolysaccharide. Mol. Cell. Biol. 26, 735–742 (2006).
Walters, K. A. et al. Prior infection with Type A Francisella tularensis antagonizes the pulmonary transcriptional response to an aerosolized Toll-like receptor 4 agonist. BMC Genomics 16, 874 (2015).
Copenhaver, A. M. et al. Alveolar macrophages and neutrophils are the primary reservoirs for Legionella pneumophila and mediate cytosolic surveillance of type IV secretion. Infect. Immun. 82, 4325–4336 (2014).
Uematsu, S. et al. Detection of pathogenic intestinal bacteria by Toll-like receptor 5 on intestinal CD11c+ lamina propria cells. Nat. Immunol. 7, 868–874 (2006).
Fontana, M. F. et al. Secreted bacterial effectors that inhibit host protein synthesis are critical for induction of the innate immune response to virulent Legionella pneumophila. PLoS Pathog 7, e1001289 (2011).
Vinzing, M. et al. NAIP and Ipaf control Legionella pneumophila replication in human cells. J. Immunol. 180, 6808–6815 (2008).
Chen, K., Yuan, R., Zhang, Y., Geng, S. & Li, L. Tollip deficiency alters atherosclerosis and steatosis by disrupting lipophagy. J. Am. Heart Assoc. 6, e004078 (2017).
Deretic, V., Saitoh, T. & Akira, S. Autophagy in infection, inflammation and immunity. Nat. Rev. Immunol. 13, 722–737 (2013).
Deretic, V. et al. Immunologic manifestations of autophagy. J. Clin. Invest 125, 75–84 (2015).
Fontana, M. F. & Vance, R. E. Two signal models in innate immunity. Immunol. Rev. 243, 26–39 (2011).
Saitoh, T. et al. Loss of the autophagy protein Atg16L1 enhances endotoxin-induced IL-1beta production. Nature 456, 264–268 (2008).
Nastase, M. V. et al. An essential role for SHARPIN in the regulation of caspase 1 activity in sepsis. Am. J. Pathol. 186, 1206–1220 (2016).
Rolando, M., Escoll, P. & Buchrieser, C. Legionella pneumophila restrains autophagy by modulating the host’s sphingolipid metabolism. Autophagy 12, 1053–1054 (2016).
Nakahira, K. et al. Autophagy proteins regulate innate immune responses by inhibiting the release of mitochondrial DNA mediated by the NALP3 inflammasome. Nat. Immunol. 12, 222–230 (2011).
Sakaguchi, S. et al. Essential role of IRF-3 in lipopolysaccharide-induced interferon-beta gene expression and endotoxin shock. Biochem. Biophys. Res. Commun. 306, 860–866 (2003).
Zhang, K. & Kaufman, R. J. From endoplasmic-reticulum stress to the inflammatory response. Nature 454, 455–462 (2008).
Cordell, H. J. & Clayton, D. G. Genetic association studies. Lancet 366, 1121–1131 (2005).
Samrakandi, M. M., Cirillo, S. L., Ridenour, D. A., Bermudez, L. E. & Cirillo, J. D. Genetic and phenotypic differences between Legionella pneumophila strains. J. Clin. Microbiol. 40, 1352–1362 (2002).
Shames, S. R. et al. Multiple Legionella pneumophila effector virulence phenotypes revealed through high-throughput analysis of targeted mutant libraries. Proc. Natl Acad. Sci. USA 114, E10446–E10454 (2017).
Burstein, D. et al. Genomic analysis of 38 Legionella species identifies large and diverse effector repertoires. Nat. Genet. 48, 167–175 (2016).
Acknowledgements
We thank the families who participated in the study. We thank Kevin Hybiske, Meghan Zuck, and Stephen Hawn for valuable intellectual contributions and assay development. We also acknowledge the support of the Cell Analysis Flow Cytometry and Imaging Core in the Department of Immunology at the University of Washington. This work was supported by R01 AI136971, K08 AI102971, and VA R&D to JAS, R01 AI093646 to SJS, K24 AI137310 to TRH, K08 HL127075 to CFH. Lp strains were a gift of Klaus Heuner and Russell Vance.
Author information
Authors and Affiliations
Contributions
Study design: J.A.S., S.J.S. Manuscript preparation and editing: J.A.S., S.J.S., T.R.H. Data acquisition, analysis, and interpretation: J.A.S., R.E., B.L., S.V., J.D.S., M.B., C.F.H., S.J.S.. Human cohort recruitment and data analysis: J.A.S., J.M.P., A.V., T.R.H.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Shah, J.A., Emery, R., Lee, B. et al. TOLLIP deficiency is associated with increased resistance to Legionella pneumophila pneumonia. Mucosal Immunol 12, 1382–1390 (2019). https://doi.org/10.1038/s41385-019-0196-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41385-019-0196-7
This article is cited by
-
TOLLIP inhibits lipid accumulation and the integrated stress response in alveolar macrophages to control Mycobacterium tuberculosis infection
Nature Microbiology (2024)
-
Toll-interacting protein may affect doxorubicin resistance in hepatocellular carcinoma cell lines
Molecular Biology Reports (2023)
-
Tollip interaction with STAT3: a novel mechanism to regulate human airway epithelial responses to type 2 cytokines
Respiratory Research (2022)
-
Bedeutung der Genetik für die Pneumonie und COVID-19
Der Pneumologe (2021)
-
Toll-interacting protein impacts on inflammation, autophagy, and vacuole trafficking in human disease
Journal of Molecular Medicine (2021)