Abstract
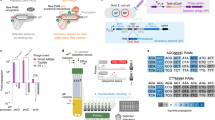
Compact Cas9 nucleases hold great promise for therapeutic applications. Although several compact Cas9 nucleases have been developed, many genomic loci still could not be edited due to a lack of protospacer adjacent motifs (PAMs). We previously developed a compact SlugCas9 recognizing an NNGG PAM. Here we demonstrate that SlugCas9 displays comparable activity to SpCas9. We developed a simple phage-assisted evolution to engineer SlugCas9 for unique PAM requirements. Interestingly, we generated a SlugCas9 variant (SlugCas9-NNG) that could recognize an NNG PAM, expanding the targeting scope. We further developed a SlugCas9-NNG-based adenine base editor and demonstrated that it could be delivered by a single adeno-associated virus to disrupt PCSK9 splice donor and splice acceptor. These genome editors greatly enhance our ability for in vivo genome editing.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All HTS data have been uploaded to the NCBI Sequence Read Archive database under accession code PRJNA985249. There are no restrictions on data availability. Source data are provided with this paper.
Change history
05 January 2024
A Correction to this paper has been published: https://doi.org/10.1038/s41589-024-01544-1
References
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013).
Mali, P. et al. RNA-guided human genome engineering via Cas9. Science 339, 823–826 (2013).
Xie, Y. et al. An episomal vector-based CRISPR/Cas9 system for highly efficient gene knockout in human pluripotent stem cells. Sci. Rep. 7, 2320 (2017).
Qi, T. et al. Base editing mediated generation of point mutations into human pluripotent stem cells for modeling disease. Front. Cell Dev. Biol. 8, 590581 (2020).
Wang, B. et al. krCRISPR: an easy and efficient strategy for generating conditional knockout of essential genes in cells. J. Biol. Eng. 13, 35 (2019).
Wang, S. et al. Identification of SaCas9 orthologs containing a conserved serine residue that determines simple NNGG PAM recognition. PLoS Biol. 20, e3001897 (2022).
Jinek, M. et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816–821 (2012).
Mojica, F. J. M., Diez-Villasenor, C., Garcia-Martinez, J. & Almendros, C. Short motif sequences determine the targets of the prokaryotic CRISPR defence system. Microbiology 155, 733–740 (2009).
Gaudelli, N. M. et al. Programmable base editing of A*T to G*C in genomic DNA without DNA cleavage. Nature 551, 464–471 (2017).
Komor, A. C., Kim, Y. B., Packer, M. S., Zuris, J. A. & Liu, D. R. Programmable editing of a target base in genomic DNA without double-stranded DNA cleavage. Nature 533, 420–424 (2016).
Anzalone, A. V. et al. Search-and-replace genome editing without double-strand breaks or donor DNA. Nature 576, 149–157 (2019).
Paquet, D. et al. Efficient introduction of specific homozygous and heterozygous mutations using CRISPR/Cas9. Nature 533, 125–129 (2016).
Agudelo, D. et al. Versatile and robust genome editing with Streptococcus thermophilus CRISPR1–Cas9. Genome Res. 30, 107–117 (2020).
Chatterjee, P., Jakimo, N. & Jacobson, J. M. Minimal PAM specificity of a highly similar SpCas9 ortholog. Sci. Adv. 4, eaau0766 (2018).
Chatterjee, P. et al. An engineered ScCas9 with broad PAM range and high specificity and activity. Nat. Biotechnol. 38, 1154–1158 (2020).
Kleinstiver, B. P. et al. Engineered CRISPR–Cas9 nucleases with altered PAM specificities. Nature 523, 481–485 (2015).
Hu, J. H. et al. Evolved Cas9 variants with broad PAM compatibility and high DNA specificity. Nature 556, 57–63 (2018).
Miller, S. M. et al. Continuous evolution of SpCas9 variants compatible with non-G PAMs. Nat. Biotechnol. 38, 471–481 (2020).
Walton, R. T., Christie, K. A., Whittaker, M. N. & Kleinstiver, B. P. Unconstrained genome targeting with near-PAMless engineered CRISPR–Cas9 variants. Science 368, 290–296 (2020).
Nishimasu, H. et al. Engineered CRISPR–Cas9 nuclease with expanded targeting space. Science 361, 1259–1262 (2018).
Edraki, A. et al. A compact, high-accuracy Cas9 with a dinucleotide PAM for in vivo genome editing. Mol. Cell 73, 714–726 e714 (2019).
Ran, F. A. et al. In vivo genome editing using Staphylococcus aureus Cas9. Nature 520, 186–191 (2015).
Kim, E. et al. In vivo genome editing with a small Cas9 orthologue derived from Campylobacter jejuni. Nat. Commun. 8, 14500 (2017).
Gao, N. et al. Characterization of Brevibacillus laterosporus Cas9 (BlatCas9) for mammalian genome editing. Front. Cell Dev. Biol. 8, 583164 (2020).
Hu, Z. et al. Discovery and engineering of small SlugCas9 with broad targeting range and high specificity and activity. Nucleic Acids Res. 49, 4008–4019 (2021).
Wang, S. et al. Compact SchCas9 recognizes the simple NNGR PAM. Adv. Sci. 9, e2104789 (2022).
Wei, J. et al. Closely related type II-C Cas9 orthologs recognize diverse PAMs. eLife https://doi.org/10.7554/eLife.77825 (2022).
Huang, T. P. et al. High-throughput continuous evolution of compact Cas9 variants targeting single-nucleotide-pyrimidine PAMs. Nat. Biotechnol. https://doi.org/10.1038/s41587-022-01410-2 (2022).
Esvelt, K. M., Carlson, J. C. & Liu, D. R. A system for the continuous directed evolution of biomolecules. Nature 472, 499–503 (2011).
Richter, M. F. et al. Phage-assisted evolution of an adenine base editor with improved Cas domain compatibility and activity. Nat. Biotechnol. 38, 883–891 (2020).
Badran, A. H. & Liu, D. R. Development of potent in vivo mutagenesis plasmids with broad mutational spectra. Nat. Commun. 6, 8425 (2015).
Nishimasu, H. et al. Crystal structure of Staphylococcus aureus Cas9. Cell 162, 1113–1126 (2015).
Zettler, J., Schutz, V. & Mootz, H. D. The naturally split Npu DnaE intein exhibits an extraordinarily high rate in the protein trans-splicing reaction. FEBS Lett. 583, 909–914 (2009).
Miller, S. M., Wang, T. & Liu, D. R. Phage-assisted continuous and non-continuous evolution. Nat. Protoc. 15, 4101–4127 (2020).
Hu, Z. et al. A compact Cas9 ortholog from Staphylococcus auricularis (SauriCas9) expands the DNA targeting scope. PLoS Biol. 18, e3000686 (2020).
Leenay, R. T. et al. Identifying and visualizing functional PAM diversity across CRISPR–Cas systems. Mol. Cell 62, 137–147 (2016).
Hu, Z. et al. A highly sensitive GFP activation assay for detection of DNA cleavage in cells. Front. Cell Dev. Biol. 9, 771248 (2021).
Walton, R. T., Hsu, J. Y., Joung, J. K. & Kleinstiver, B. P. Scalable characterization of the PAM requirements of CRISPR–Cas enzymes using HT-PAMDA. Nat. Protoc. 16, 1511–1547 (2021).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Tsai, S. Q. et al. GUIDE-seq enables genome-wide profiling of off-target cleavage by CRISPR–Cas nucleases. Nat. Biotechnol. 33, 187–197 (2015).
Slaymaker, I. M. et al. Rationally engineered Cas9 nucleases with improved specificity. Science 351, 84–88 (2016).
NM_000335.5(SCN5A):c.1363A>G (p.Thr455Ala). National Center for Biotechnology Information (2016); https://www.ncbi.nlm.nih.gov/clinvar/variation/201550/#id_first
Huang, T. P., Newby, G. A. & Liu, D. R. Precision genome editing using cytosine and adenine base editors in mammalian cells. Nat. Protoc. 16, 1089–1128 (2021).
Zhang, H. et al. Adenine base editing in vivo with a single adeno-associated virus vector. GEN Biotechnol. 1, 285–299 (2022).
Davis, J. R. et al. Efficient in vivo base editing via single adeno-associated viruses with size-optimized genomes encoding compact adenine base editors. Nat. Biomed. Eng. 6, 1272–1283 (2022).
Cohen, J. C., Boerwinkle, E., Mosley, T. H. Jr. & Hobbs, H. H. Sequence variations in PCSK9, low LDL, and protection against coronary heart disease. N. Engl. J. Med. 354, 1264–1272 (2006).
Chen, P. J. et al. Enhanced prime editing systems by manipulating cellular determinants of editing outcomes. Cell 184, 5635–5652 e5629 (2021).
Acknowledgements
This work was supported by grants from the National Key Research and Development Program of China (2021YFC2701103 to Yongming W., 2021YFA0910602 to Yongming W.); the National Natural Science Foundation of China (82370254 to Yongming W., 82070258 to Yongming W.); the Open Research Fund of State Key Laboratory of Genetic Engineering, Fudan University (No. SKLGE-2104 to Yongming W.); the Science and Technology Research Program of Shanghai (19DZ2282100 to Yongming W., 23ZR1426000 to Yongming W.); the Scientific Research Foundation of Peking University Shenzhen Hospital (KYQD202100X to W.V.Z.); Shenzhen Science and Technology Program (JCYJ20220818102818039 to W.V.Z.); BYD Charity Foundation (PUSH-BYD JCYJ202201 to W.V.Z., PUSH-BYD JCYJ202204 to W.V.Z.); Shenzhen Biomedical Industry Major Public Service Platform and Core Technology Research Special Support Plan (XMHT20220104048 to W.V.Z.); National Natural Science Foundation of China (Nos. 82222007 to J.T., 82170281 to J.T.); Henan Province Medical Science and Technology Key Joint Project (SBGJ202101012 to J.T.); Funding for Scientific Research and Innovation Team of The First Affiliated Hospital of Zhengzhou University (QNCXTD2023001 to J.T.).
Author information
Authors and Affiliations
Contributions
T.Q. and Yongming W. designed the research. T.Q. and J.L. performed evolution. T.Q., Yao W., S.G. and Y.T. conducted human cell experiments. T.Q. and Y.Y. analyzed the data. Q.H. employed AlphaFold to predict the structures of SlugCas9 and SlugCas9-NNG. T.Q., Yongming W., and Yao W. wrote the manuscript. Yongming W., J.T. and W.V.Z. guided the project.
Corresponding authors
Ethics declarations
Competing interests
Yongming W. and T.Q. have submitted a patent application (2023106924996) on the basis the results reported in this study. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Chemical Biology thanks Pranam Chatterjee, Benjamin W. Thuronyi and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 SlugCas9 exhibits comparable editing activity to SpCas9.
a, Schematic of the Cas9 expression constructs. Both SpCas9 and SlugCas9 constructs have the same backbone. b, Expression levels of SpCas9 and SlugCas9 measured by RT-qPCR. Two-tailed, paired Student’s t-test was used to determine the statistical significance between the two groups. Data are mean + s.d. of n = 3 independent experiments. c, Schematic of sgRNA design. d, Indel frequency of SpCas9 and SlugCas9 at 23 endogenous target sites in HEK293T cells. Cas9 and sgRNA expression plasmids were transfected into HEK293T cells, followed by puromycin selection. Data are mean + s.d. of n = 3 independent experiments. e, Summary of the indel frequency of SpCas9 and SlugCas9. Two-tailed, paired Student’s t-test was used to determine the statistical significance between the two groups. The mean and s.d. of all individual values of n = 3 independent biological replicates are plotted. Statistical source data are available (Source Data Extended Data Fig. 1).
Extended Data Fig. 2 Split of SlugABE.
a, Sanger sequencing of the target sites on the T7RNAP gene-edited by SlugABE or split SlugABE. Dotted boxes indicate the target adenines (shown in red). b, Black arrows indicate the positions of the split sites on SlugCas9. Amino acid positions are shown above.
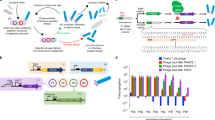
Extended Data Fig. 3 Phage propagation level detected by PCR amplification.
a, PCR amplification of the phage genome (1,615 bp). 1 µL of filtered supernatant containing phages from the phage propagation assay is used as a template. Forward and reverse primers are indicated by red arrows. b, PCR products are analyzed by agarose gel electrophoresis. The expected PCR band size (1,615 bp) is indicated on the right by the black arrow. Ctr, a titer of 106 phages, is input into 2 ml of medium containing bacteria (OD600 = 0.5) and immediately filtered without culture. The PAM for each site is shown. The experiment was independently repeated three times and yielded similar results (Source Data Extended Data Fig. 3).
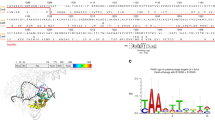
Extended Data Fig. 4 Dominant mutations obtained from the first-round evolution.
a, Mutations detected by Sanger sequencing. Mutations are marked in red. b, Influence of N984S on phage propagation. Data are mean + s.d. of n = 3 independent experiments. c, Influence of K1016I on phage propagation. Data are mean + s.d. of n = 3 independent experiments. Statistical source data are available (Source Data Extended Data Fig. 4).
Extended Data Fig. 5 A GFP-activation assay for PAM identification.
a, PAM logos of SlugCas9-N984S and SlugCas9-K1016I identified by the GFP-activation assay. b, PAM wheels of SlugCas9, SlugCas9-N984S, SlugCas9-K1016I, and SlugCas9-N984S/K1016I were identified by the GFP-activation assay.
Extended Data Fig. 6 Testing the activity of SlugCas9-N984S/K1016I using the GFP-activation reporter in HEK293T cells.
a, Schematic of the GFP-activation reporter. Dark lines indicate target sites; PAMs are indicated by red lines. b, Percentage of GFP-positive cells after genome editing by SlugCas9 or SlugCas9-N984S/K1016I. For the gating strategy, begin by drawing the FSC/SSC dual-parameter dot map, then encircle the target cell population and establish the gate R1. Subsequently, draft the GFP/SSC dual-parameter dot map. After applying Gate R1 to this map, set a gate to encircle GFP positive cells, R2, using the negative control as a reference. Two-tailed, paired Student’s t-test was used to determine the statistical significance between the two groups. A value of P < 0.05 was considered to be statistically significant (*P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001). Data are mean + s.d. of n = 3 independent experiments. Statistical source data are available (Source Data Extended Data Fig. 6).
Extended Data Fig. 7 Structure model of SlugCas9 and SlugCas9-NNG predicted by AlphaFold.
a, SlugCas9 structure predicted by AlphaFold. b, SlugCas9-NNG structure predicted by AlphaFold. c, The SlugCas9-NNG structure is superimposed on the SlugCas9 structure.
Extended Data Fig. 8 Adenine base editing with SlugABE-NNG in human iPS cells.
A to G conversion frequencies at 10 target sites in iPS cells. The PAM for each site is shown below. Data are mean + s.d. of n = 3 independent experiments. Statistical source data are available (Source Data Extended Data Fig. 8).
Extended Data Fig. 9 Base editing with SlugCBE-NNG in human cells.
a, Schematic of the SlugCBE-NNG. b, Examples of C to T conversion at target sites in HEK293T cells. The PAM for each site is shown below. Data are mean + s.d. of n = 3 independent experiments. c, Editing window generated from 40 genomic target sites. The mean and s.d. of all individual values of n = 3 independent biological replicates are plotted. Statistical source data are available (Source Data Extended Data Fig. 9).
Extended Data Fig. 10 SlugABE-NNG can be delivered via a single AAV for genome editing.
a, Schematic of the single-AAV SlugABE-NNG. EFS represents EF1A core promoter; bGH represents bovine growth hormone polyadenylation signal; sgRNA expression cassette includes hU6 promoter, sgRNA, and seven ‘T’ terminator. b, Sequences of the five target sites. The target adenines are marked in red, and the PAM sequences are highlighted in bold.
Supplementary information
Supplementary Information
Supplementary Figs. 1–10 and Notes 1–8.
Supplementary Data 1
Statistical source data for Supplementary Figs. 1 and 5–10.
Supplementary Tables 1–4
Supplementary Tables 1–4.
Source data
Source Data Figs. 1–5 and Extended Data Figs. 1, 4, 6, 8 and 9
Statistical source data.
Source Data Extended Data Fig. 3
Unprocessed gel.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Qi, T., Wang, Y., Yang, Y. et al. Phage-assisted evolution of compact Cas9 variants targeting a simple NNG PAM. Nat Chem Biol 20, 344–352 (2024). https://doi.org/10.1038/s41589-023-01481-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-023-01481-5