Abstract
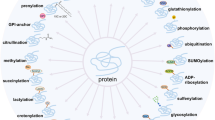
Protein lipidation, which regulates numerous biological pathways and plays crucial roles in the pharmaceutical industry, is not encoded by the genetic code but synthesized post-translationally. In the present study, we report a computational approach for designing lipidation mimics that fully recapitulate the biochemical properties of natural lipidation in membrane association and albumin binding. Furthermore, we establish an engineered system for co-translational incorporation of these lipidation mimics into virtually any desired position of proteins in Escherichia coli and mammalian cells. We demonstrate the utility of these length-tunable lipidation mimics in diverse applications, including improving the half-life and activity of therapeutic proteins in living mice, anchoring functional proteins to membrane by substituting natural lipidation, functionally characterizing proteins carrying different lengths of lipidation and determining the plasma membrane-binding capacity of a given compound. Our strategy enables gain-of-function studies of lipidation in hundreds of proteins and facilitates the creation of superior therapeutic candidates.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Supplementary information (methods and figures) and chemical compound information are available in the online version of the paper. Correspondence and requests for materials should be addressed to S.L. Source data are provided with this paper.
Code availability
The source codes for python used for clog(P) and ΔG plotting are available in Supplementary Information.
References
Janda, C. Y., Waghray, D., Levin, A. M., Thomas, C. & Garcia, K. C. Structural basis of Wnt recognition by Frizzled. Science 337, 59–64 (2012).
Jiang, H. et al. Protein lipidation: occurrence, mechanisms, biological functions, and enabling technologies. Chem. Rev. 118, 919–988 (2018).
Ping, Y. Q. et al. Structures of the glucocorticoid-bound adhesion receptor GPR97-G(o) complex. Nature 589, 620–626 (2021).
Casey, P. J. Protein lipidation in cell signaling. Science 268, 221–225 (1995).
Rocks, O. et al. An acylation cycle regulates localization and activity of palmitoylated Ras isoforms. Science 307, 1746–1752 (2005).
Ferre, G. et al. Structure and dynamics of G protein-coupled receptor-bound ghrelin reveal the critical role of the octanoyl chain. Proc. Natl Acad. Sci. USA 116, 17525–17530 (2019).
Takada, R. et al. Monounsaturated fatty acid modification of Wnt protein: its role in Wnt secretion. Dev. Cell 11, 791–801 (2006).
Jiang, H. et al. SIRT6 regulates TNF-alpha secretion through hydrolysis of long-chain fatty acyl lysine. Nature 496, 110–113 (2013).
Zha, J. P., Weiler, S., Oh, K. J., Wei, M. C. & Korsmeyer, S. J. Posttranslational N-myristoylation of BID as a molecular switch for targeting mitochondria and apoptosis. Science 290, 1761–1765 (2000).
Bersuker, K. et al. The CoQ oxidoreductase FSP1 acts parallel to GPX4 to inhibit ferroptosis. Nature 575, 688–692 (2019).
Doll, S. et al. FSP1 is a glutathione-independent ferroptosis suppressor. Nature 575, 693–698 (2019).
Mukai, K. et al. Activation of STING requires palmitoylation at the Golgi. Nat. Commun. 7, 11932 (2016).
Zhang, M. et al. A STAT3 palmitoylation cycle promotes TH17 differentiation and colitis. Nature 586, 434–439 (2020).
Das, T., Yount, J. S. & Hang, H. C. Protein S-palmitoylation in immunity. Open Biol. 11, 200411 (2021).
Kokame, K., Fukada, Y., Yoshizawa, T., Takao, T. & Shimonishi, Y. Lipid modification at the N-terminus of photoreceptor G-protein alpha-subunit. Nature 359, 749–752 (1992).
Resh, M. D. Fatty acylation of proteins: the long and the short of it. Prog. Lipid Res. 63, 120–131 (2016).
Losada de la Lastra, A., Hassan, S. & Tate, E. W. Deconvoluting the biology and druggability of protein lipidation using chemical proteomics. Curr. Opin. Chem. Biol. 60, 97–112 (2021).
Knudsen, L. B. et al. Potent derivatives of glucagon-like peptide-1 with pharmacokinetic properties suitable for once daily administration. J. Med. Chem. 43, 1664–1669 (2000).
Lau, J. et al. Discovery of the once-weekly glucagon-like peptide-1 (GLP-1) analogue semaglutide. J. Med. Chem. 58, 7370–7380 (2015).
Menacho-Melgar, R., Decker, J. S., Hennigan, J. N. & Lynch, M. D. A review of lipidation in the development of advanced protein and peptide therapeutics. J. Control. Release 295, 1–12 (2019).
Garst, E. H., Das, T. & Hang, H. C. Chemical approaches for investigating site-specific protein S-fatty acylation. Curr. Opin. Chem. Biol. 65, 109–117 (2021).
Wang, L., Brock, A., Herberich, B. & Schultz, P. G. Expanding the genetic code of Escherichia coli. Science 292, 498–500 (2001).
Neumann, H., Peak-Chew, S. Y. & Chin, J. W. Genetically encoding N-epsilon-acetyllysine in recombinant proteins. Nat. Chem. Biol. 4, 232–234 (2008).
Fu, C. Y. et al. Genetically encoding a lipidated amino acid for extension of protein half-life in vivo. Angew. Chem. Int. Ed. 58, 1392–1396 (2019).
Zhang, Z. J., Pedicord, V. A., Peng, T. & Hang, H. C. Site-specific acylation of a bacterial virulence regulator attenuates infection. Nat. Chem. Biol. 16, 95–103 (2020).
Forli, S. et al. Computational protein-ligand docking and virtual drug screening with the AutoDock suite. Nat. Protoc. 11, 905–919 (2016).
Leo, A., Hansch, C. & Elkins, D. Partition coefficients and their uses. Chem. Rev. 71, 525–616 (1971).
Ng, C. A. & Hungerbuehler, K. Exploring the use of molecular docking to identify bioaccumulative perfluorinated alkyl acids (PFAAs). Environ. Sci. Technol. 49, 12306–12314 (2015).
Dumas, A., Lercher, L., Spicer, C. D. & Davis, B. G. Designing logical codon reassignment—expanding the chemistry in biology. Chem. Sci. 6, 50–69 (2015).
Ding, W. et al. Chimeric design of pyrrolysyl-tRNA synthetase/tRNA pairs and canonical synthetase/tRNA pairs for genetic code expansion. Nat. Commun. 11, 3154 (2020).
Zhao, H. et al. Directed-evolution of translation system for efficient unnatural amino acids incorporation and generalizable synthetic auxotroph construction. Nat. Commun. 12, 7039 (2021).
Bhattacharya, A. A., Grune, T. & Curry, S. Crystallographic analysis reveals common modes of binding of medium and long-chain fatty acids to human serum albumin. J. Mol. Biol. 303, 721–732 (2000).
Silva, D. A. et al. De novo design of potent and selective mimics of IL-2 and IL-15. Nature 565, 186–191 (2019).
Hannoush, A. N. & Sun, J. The chemical toolbox for monitoring protein fatty acylation and prenylation. Nat. Chem. Biol. 6, 498–506 (2010).
Bivona, T. G. et al. PKC regulates a farnesyl-electrostatic switch on K-Ras that promotes its association with Bcl-XL on mitochondria and induces apoptosis. Mol. Cell 21, 481–493 (2006).
Michaelson, D., Ahearn, I., Bergo, M., Young, S. & Philips, M. Membrane trafficking of heterotrimeric G proteins via the endoplasmic reticulum and Golgi. Mol. Biol. Cell 13, 3294–3302 (2002).
Hurd, T. et al. The retinitis pigmentosa protein RP2 interacts with polycystin 2 and regulates cilia-mediated vertebrate development. Hum. Mol. Genet. 19, 4330–4344 (2010).
Yasuda, K. et al. Serine 6 of Lck tyrosine kinase: a critical site for Lck myristoylation, membrane localization, and function in T lymphocytes. J. Immunol. 165, 3226–3231 (2000).
Peitzsch, R. M. & McLaughlin, S. Binding of acylated peptides and fatty acids to phospholipid vesicles: pertinence to myristoylated proteins. Biochemistry 32, 10436–10443 (1993).
Velazquez-Libera, J. L., Duran-Verdugo, F., Valdes-Jimenez, A., Nunez-Vivanco, G. & Caballero, J. LigRMSD: a web server for automatic structure matching and RMSD calculations among identical and similar compounds in protein-ligand docking. Bioinformatics 36, 2912–2914 (2020).
Acknowledgements
We thank the National Key R&D Program of China (grant no. 2019YFA09006600), the National Natural Science Foundation of China (grant nos. 22222705, 92253302, 91953113 and 21877096), Young Scientists Fund of the National Natural Science Foundation of China (grant no. 22207095 for W.D.), China Postdoctoral Science Foundation (grant no. 2019M652072 to W.D.) and the Fundamental Research Funds for the Central Universities (grant no. K20220228) for financial support. We are grateful to the core facility of the Life Sciences Institute and X. He for helpful discussions.
Author information
Authors and Affiliations
Contributions
S.L. conceived the idea and supervised the study. W.D. and C.L. conducted most experiments and analyzed the data together. Y.C. and C.F. helped with the virtual screening, J.G. and L.H. assisted with mice experiments. L.Z., Y.Y. and X.-H.F. provided technical advice. S.L. and W.D. wrote the manuscript. All authors commented on the final draft of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare the following competing interests: S.L., W.D., C.L. and Y.C. are listed as inventors on a patent application describing the development and application of genetically encoded lipidation mimics (Chinese patent application no. 202310095754.9). All remaining authors declare no competing interests.
Peer review
Peer review information
Nature Chemical Biology thanks Jeremy Mills and Xu Wu for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 The computationally assisted design and screening of lipidation mimics.
(a) The procedure for predicting the HSA binding affinity (ΔG) of lipidation mimics. Lipidation mimics were docked to 7 sites of HSA (highlighted with red dash boxes) by AutoDock Vina. (b) The procedure for calculating RMSD. RMSD, defined as the average distance between the coordinates of the lipidation mimics obtained by docking and the coordinates of the reference ligand generated from the crystal structure, was calculated by LigRMSD. (c) The chemical structures of selected lipidation mimics and myristate whose linear carbon chains were numbered and colored in red. (d) The predicted cLog(P) of lipidation mimics in (C). (e) The HSA binding affinity of selected lipid mimics and fatty acids in the 7 binding sites of HSA. (f) The docking structure of 4HexyF (white) in the 7 binding sites of HSA. The amino acids surrounding 4HexyF for π-π interactions were highlighted by magenta. (g) Surface plasmon resonance analysis of HSA binding affinity of selected lipidation mimics. Data were fitted to single affinity mode and the binding affinities were shown.
Extended Data Fig. 2 The detailed information of Fig. 1b.
Every symbol was numbered as Supplementary Data 1. HepoK, 4HexyF, 4OctyF, 4NonyF and 4DecyF were numbered as K2, F1, F2, F3 and F4 respectively.
Extended Data Fig. 3 Computationally aided design of functional lipidation mimics.
(a) Candidate lipid mimics with -ΔG > 8.2 and cLogP > 4.0 were plotted based on their ΔG and cLog(P) values. The dash line indicated the RMSD threshold of around 2.20 for Fs, 2.29 for Ws, and 2.93 for As and Ks. UAAs with RMSD above the threshold were colored with grey. UAAs with RMSD values below the threshold on the left side of the dash line had a high chance of being recognized by orthogonal aaRSs. (b) The chemical structure of candidate lipid mimics. (c) Phenylalanine derivatives in (B) were plotted based on RMSD and aliphatic carbon chains length. The RMSD value was dramatically increased when length of aliphatic carbon chains was greater than 9.
Extended Data Fig. 4 The chemical synthesis and genetic encoding of lipidation mimics.
(a) The chemical synthesis of 4HexyF, 4OctyF, 4NonyF and 4DecyF. A three-step synthetic route was developed with the isolation yield of each synthetic step shown under the products. (b) The model structure of human mitochondrial PheRS in complex with 4HexyF. The gatekeeper residues colored in magenta displayed steric hindrance with 4HexyF. (c) The model structure of LipRS-1 in complex with 4HexyF (cyan). Residues around 4HexyF were shown as sticks with three key mutations highlighted in magenta and the others in blue. (d) The amber suppression assay of lipidation mimics by the indicated LipRS-2 variants (n = 2 biologically independent experiments). (e) The gate strategy of FACS used in Fig. 4a.
Extended Data Fig. 5 The high HSA binding affinity of proteins incorporated with lipidation mimics.
(a) Coomassie blue staining gel analysis of purified GLP-1 variants incorporated with indicated lipidation mimics. The experiment in the figure was repeated twice with similar results. (b) Mass spectrometry characterization of the fidelity of lipid mimics on GLP-1. The expected MW (-Met) of 4HexyF and 4OctyF incorporation were 15463 and 15491 Da, and the observed MW (-Met) were 15461 and 15490 Da, respectively. (c) MST analysis of affinity of GLP-1 variants for HSA. The data were fitted with Kd model by MO.Affinity analysis software and the binding affinity were shown. Error bars represented ± standard error of the mean (n = 3 biologically independent experiments). (d, e) ITC analysis of the affinity of GFP variants (D) or Neo-2/15 variants (E) for HSA. The data were fit to a single binding site model, and fitting thermostability parameters were shown in the figure.
Extended Data Fig. 6 Evaluating the therapeutic effect of lipidation mimics and Neo-2/15.
CT26 tumor growth (a) and Kaplan-Meier survival curves (b) in mice after treatment with 4OctyF, Neo-2/15-F and 4OctyF plus Neo-2/15-F, respectively, were quantified. Statistical significance of tumor growth was quantified with unpaired two-tailed t-test (n = 4). Mice survival was analyzed with a log-rank test (n = 4 biologically independent experiments). Error bars represents ± standard error of the mean (n = 4 biologically independent experiments).
Extended Data Fig. 7 Genetically encoded lipidation mimic for efficient membrane association.
(a) The representative images (n = 3) of G2F mutation of indicated proteins showed cytoplasmic localization. (b, d) The representative images (n = 3) of XRP2 variants (B) and Lck variants (D). (c, e) Fluorescence intensity profiles across the yellow line in (B, D) indicated the P/C ratio of XRP2 variants (C) and Lck variants (D), respectively. (f) The representative images (n = 3) of XRP2 and Lck variants incorporated with 4HexyF. The Scale bar: 10 μm.
Extended Data Fig. 8 The 4OctyF incorporation mimic the function of Gα subunit.
(a) the assembly model of G-protein αβγ heterotrimers. (b) Representative images (n = 3) showed that 4OctyF incorporation of Gα subunit could bind and recruit Gβγ complex to plasma membrane. Scale bar: 10 μm.
Extended Data Fig. 9 The membrane association capability of length-tunable lipidation mimics.
(a) The representative images (n = 3) of protein with the construct cartoon shown was incorporated with length-tunable lipidation mimics. Scale bar: 10 μm. (b) Fluorescence intensity profiles across the yellow lines in (A) showed perinuclear area accumulation of protein in the 4OctyF group. (c) Mass spectrometry characterization of the fidelity of lipidation mimics on polyK-EGFP. The expected MW (-Met) of 4HexyF and 4OctyF incorporation were 29648 and 29676 Da, and the observed MW (-Met) were 29649 and 29677 Da, respectively. (d) The cartoon procedure of in vitro binding affinity with liposomes. The liposome binding affinity of protein was determined by measuring the fraction of liposome-protein complex with liposomes co-sedimentation assay. (e) The cLogP threshold (x) for efficient membrane anchoring was between 5.07 (laurate) and 5.3 (4OctyF) for the peptide substrates with similar sequence. The peptide modified by a compound with cLogP value larger than x (green hexagon) tended to enrich at plasma membrane, while compound with cLogP value less than x (blue square) distributed diffusely in cytoplasm.
Supplementary information
Supplementary Information
Supplementary Notes 1 and 2, and Figs. 1–18.
Supplementary Data 1
The chemical structure and biophysical properties of lipidation mimics that we designed.
Source data
Source Data Fig. 1
Source data of clog(P) and ΔG of lipidation mimics in Fig. 1.
Source Data Fig. 2
Raw plate reader data and raw MS data in Fig. 2.
Source Data Fig. 3
Raw MS data, ITC data and tumor data in Fig. 3
Source Data Fig. 4
Raw FACS statistical data.
Source Data Fig. 4
Unprocessed western blots in Fig. 4.
Source Data Fig. 5
Raw micrograph profiling data and plate reader data in Fig. 5.
Source Data Extended Data Fig. 1
Raw SPR data in Extended Data Fig. 1.
Source Data Extended Data Fig. 3
Raw data for RMSD plotting in Extended Data Fig. 3.
Source Data Extended Data Fig. 4
Raw plate reader data in Extended Data Fig. 4.
Source Data Extended Data Fig. 5
Raw MS data and MST data in Extended Data Fig. 5.
Source Data Extended Data Fig. 5
Unprocessed SDS–PAGE gel in Extended Data Fig. 5.
Source Data Extended Data Fig. 6
Raw data for tumor growth in Extended Data Fig. 6.
Source Data Extended Data Fig. 7
Raw micrograph profiling data in Extended Data Fig. 7.
Source Data Extended Data Fig. 9
Raw micrograph profiling data and MS data in Extended Data Fig. 9.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Ding, W., Liu, C., Chen, Y. et al. Computational design and genetic incorporation of lipidation mimics in living cells. Nat Chem Biol 20, 42–51 (2024). https://doi.org/10.1038/s41589-023-01400-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-023-01400-8
This article is cited by
-
Greasy proteins made easy
Nature Chemical Biology (2024)