Abstract
Focal adhesion kinase (FAK) relays integrin signaling from outside to inside cells and contributes to cell adhesion and motility. However, the spatiotemporal dynamics of FAK activity in single FAs is unclear due to the lack of a robust FAK reporter, which limits our understanding of these essential biological processes. Here we have engineered a genetically encoded FAK activity sensor, dubbed FAK–separation of phases-based activity reporter of kinase (SPARK), which visualizes endogenous FAK activity in living cells and vertebrates. Our work reveals temporal dynamics of FAK activity during FA turnover. Most importantly, our study unveils polarized FAK activity at the distal tip of newly formed single FAs in the leading edge of a migrating cell. By combining FAK–SPARK with DNA tension probes, we show that tensions applied to FAs precede FAK activation and that FAK activity is proportional to the strength of tension. These results suggest tension-induced polarized FAK activity in single FAs, advancing the mechanistic understanding of cell migration.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All data are available in the article, including the source data section. Source data are provided with this paper.
References
Kechagia, J. Z., Ivaska, J. & Roca-Cusachs, P. Integrins as biomechanical sensors of the microenvironment. Nat. Rev. Mol. Cell Biol. 20, 457–473 (2019).
Burridge, K. Focal adhesions: a personal perspective on a half century of progress. FEBS J. 284, 3355–3361 (2017).
Mitra, S. K., Hanson, D. A. & Schlaepfer, D. D. Focal adhesion kinase: in command and control of cell motility. Nat. Rev. Mol. Cell Biol. 6, 56–68 (2005).
Ilic, D. et al. Reduced cell motility and enhanced focal adhesion contact formation in cells from FAK-deficient mice. Nature 377, 539–544 (1995).
Parsons, J. T., Horwitz, A. R. & Schwartz, M. A. Cell adhesion: integrating cytoskeletal dynamics and cellular tension. Nat. Rev. Mol. Cell Biol. 11, 633–643 (2010).
Seong, J. et al. Detection of focal adhesion kinase activation at membrane microdomains by fluorescence resonance energy transfer. Nat. Commun. 2, 406 (2011).
Chung, C.-I., Zhang, Q. & Shu, X. Dynamic imaging of small molecule induced protein–protein interactions in living cells with a fluorophore phase transition based approach. Anal. Chem. 90, 14287–14293 (2018).
Zhang, Q. et al. Visualizing dynamics of cell signaling in vivo with a phase separation-based kinase reporter. Mol. Cell 69, 334–345.e5 (2018).
Schepis, A. et al. Protease signaling regulates apical cell extrusion, cell contacts, and proliferation in epithelia. J. Cell Biol. 217, 1097–1112 (2018).
Zouq, N. K. et al. FAK engages multiple pathways to maintain survival of fibroblasts and epithelia: differential roles for paxillin and p130Cas. J. Cell Sci. 122, 357–367 (2009).
Mastop, M. et al. Characterization of a spectrally diverse set of fluorescent proteins as FRET acceptors for mTurquoise2. Sci. Rep. 20, 11999 (2017).
Tsutsui, H., Karasawa, S., Okamura, Y. & Miyawaki, A. Improving membrane voltage measurements using FRET with new fluorescent proteins. Nat. Methods 5, 683–685 (2008).
Yu, D. et al. Rational design of a monomeric and photostable far-red fluorescent protein for fluorescence imaging in vivo. Protein Sci. 25, 308–315 (2015).
Yu, D. et al. A naturally monomeric infrared fluorescent protein for protein labeling. Nat. Methods 12, 763–765 (2015).
Yu, D. et al. An improved monomeric infrared fluorescent protein for neuronal and tumour brain imaging. Nat. Commun. 5, 3626 (2014).
Shu, X. et al. Mammalian expression of infrared fluorescent proteins engineered from a bacterial phytochrome. Science 324, 804–807 (2009).
Banaszynski, L. A., Liu, C. W. & Wandless, T. J. Characterization of the FKBP.rapamycin.FRB ternary complex. J. Am. Chem. Soc. 127, 4715–4721 (2005).
Case, L. B. & Waterman, C. M. Integration of actin dynamics and cell adhesion by a three-dimensional, mechanosensitive molecular clutch. Nat. Cell Biol. 17, 955–963 (2015).
McGrath, J. L. Cell spreading: the power to simplify. Curr. Biol. 17, R357–R358 (2007).
Tomar, A., Lim, S.-T., Lim, Y. & Schlaepfer, D. D. A FAK-p120RasGAP-p190RhoGAP complex regulates polarity in migrating cells. J. Cell Sci. 122, 1852–1862 (2009).
Ezratty, E. J., Partridge, M. A. & Gundersen, G. G. Microtubule-induced focal adhesion disassembly is mediated by dynamin and focal adhesion kinase. Nat. Cell Biol. 7, 581–590 (2005).
Ballestrem, C., Hinz, B., Imhof, B. A. & Wehrle-Haller, B. Marching at the front and dragging behind. J. Cell Biol. 155, 1319–1332 (2001).
Laukaitis, C. M., Webb, D. J., Donais, K. & Horwitz, A. F. Differential dynamics of alpha 5 integrin, paxillin, and alpha-actinin during formation and disassembly of adhesions in migrating cells. J. Cell Biol. 153, 1427–1440 (2001).
Gardel, M. L. et al. Traction stress in focal adhesions correlates biphasically with actin retrograde flow speed. J. Cell Biol. 183, 999–1005 (2008).
Feliks, M., Lafaye, C., Shu, X., Royant, A. & Field, M. Structural determinants of improved fluorescence in a family of bacteriophytochrome-based infrared fluorescent proteins: insights from continuum electrostatic calculations and molecular dynamics simulations. Biochemistry 55, 4263–4274 (2016).
Le Coq, J., Acebrón, I., Rodrigo Martin, B., López Navajas, P. & Lietha, D. New insights into FAK structure and function in focal adhesions. J. Cell Sci. 135, jcs259089 (2022).
Swaminathan, V., Fischer, R. S. & Waterman, C. M. The FAK-Arp2/3 interaction promotes leading edge advance and haptosensing by coupling nascent adhesions to lamellipodia actin. Mol. Biol. Cell 27, 1085–1100 (2016).
Serrels, B. et al. Focal adhesion kinase controls actin assembly via a FERM-mediated interaction with the Arp2/3 complex. Nat. Cell Biol. 9, 1046–1056 (2007).
Goñi, G. M. et al. Phosphatidylinositol 4,5-bisphosphate triggers activation of focal adhesion kinase by inducing clustering and conformational changes. Proc. Natl Acad. Sci. USA 111, E3177–E3186 (2014).
Seong, J. et al. Distinct biophysical mechanisms of focal adhesion kinase mechanoactivation by different extracellular matrix proteins. Proc. Natl Acad. Sci. USA 110, 19372–19377 (2013).
Orgovan, N. et al. Dependence of cancer cell adhesion kinetics on integrin ligand surface density measured by a high-throughput label-free resonant waveguide grating biosensor. Sci. Rep. 4, 4034 (2014).
Sundaram, A. et al. Targeting integrin α5β1 ameliorates severe airway hyperresponsiveness in experimental asthma. J. Clin. Invest. 127, 365–374 (2017).
Pasapera, A. M., Schneider, I. C., Rericha, E., Schlaepfer, D. D. & Waterman, C. M. Myosin II activity regulates vinculin recruitment to focal adhesions through FAK-mediated paxillin phosphorylation. J. Cell Biol. 188, 877–890 (2010).
Labouesse, C., Verkhovsky, A. B., Meister, J.-J., Gabella, C. & Vianay, B. Cell shape dynamics reveal balance of elasticity and contractility in peripheral arcs. Biophys. J. 108, 2437–2447 (2015).
Petridou, N. I., Stylianou, P. & Skourides, P. A. A dominant-negative provides new insights into FAK regulation and function in early embryonic morphogenesis. Development 140, 4266–4276 (2013).
Zheng, Y. et al. FAK phosphorylation by ERK primes Ras-induced tyrosine dephosphorylation of FAK mediated by PIN1 and PTP-PEST. Mol. Cell 35, 11–25 (2009).
Blanchard, A. et al. Turn-key mapping of cell receptor force orientation and magnitude using a commercial structured illumination microscope. Nat. Commun. 12, 4693 (2021).
Glazier, R., Shinde, P., Ogasawara, H. & Salaita, K. Spectroscopic analysis of a library of DNA tension probes for mapping cellular forces at fluid interfaces. ACS Appl. Mater. Interfaces 13, 2145–2164 (2021).
Glazier, R. et al. DNA mechanotechnology reveals that integrin receptors apply pN forces in podosomes on fluid substrates. Nat. Commun. 10, 4507 (2019).
Stabley, D. R., Jurchenko, C., Marshall, S. S. & Salaita, K. S. Visualizing mechanical tension across membrane receptors with a fluorescent sensor. Nat. Methods 9, 64–67 (2011).
Zhang, Y. et al. Platelet integrins exhibit anisotropic mechanosensing and harness piconewton forces to mediate platelet aggregation. Proc. Natl Acad. Sci. USA 115, 325–330 (2018).
Wang, X. & Ha, T. Defining single molecular forces required to activate integrin and notch signaling. Science 340, 991–994 (2013).
Pfaff, M. et al. Selective recognition of cyclic RGD peptides of NMR defined conformation by αIIbβ3, αVβ3, and α5β1 integrins. J. Biol. Chem. 269, 20233–20238 (1994).
Liu, Y. et al. Nanoparticle tension probes patterned at the nanoscale: impact of integrin clustering on force transmission. Nano Lett. 14, 5539–5546 (2014).
Liu, Y., Yehl, K., Narui, Y. & Salaita, K. Tension sensing nanoparticles for mechano-imaging at the living/nonliving interface. J. Am. Chem. Soc. 135, 5320–5323 (2013).
Rashid, S. A. et al. DNA tension probes show that cardiomyocyte maturation is sensitive to the piconewton traction forces transmitted by integrins. ACS Nano 16, 5335–5348 (2022).
Pérez, L. A. et al. An outside-in switch in integrin signaling caused by chemical and mechanical signals in reactive astrocytes. Front. Cell Dev. Biol. 9, 712627 (2021).
Jo, M. H., Cottle, W. T. & Ha, T. Real-time measurement of molecular tension during cell adhesion and migration using multiplexed differential analysis of tension gauge tethers. ACS Biomater. Sci. Eng. 5, 3856–3863 (2019).
Ritt, M., Guan, J.-L. & Sivaramakrishnan, S. Visualizing and manipulating focal adhesion kinase regulation in live cells. J. Biol. Chem. 288, 8875–8886 (2013).
Cai, X. et al. Spatial and temporal regulation of focal adhesion kinase activity in living cells. Mol. Cell. Biol. 28, 201–214 (2008).
Li, X. et al. ATM-SPARK: a GFP phase separation-based activity reporter of ATM. Sci. Adv. 9, eade3760 (2023).
Shu, X. Imaging dynamic cell signaling in vivo with new classes of fluorescent reporters. Curr. Opin. Chem. Biol. 54, 1–9 (2020).
Wakatsuki, T., Wysolmerski, R. B. & Elson, E. L. Mechanics of cell spreading: role of myosin II. J. Cell Sci. 116, 1617–1625 (2003).
Bell, S. & Terentjev, E. M. Focal adhesion kinase: the reversible molecular mechanosensor. Biophys. J. 112, 2439–2450 (2017).
Zhou, J. et al. Mechanism of focal adhesion kinase mechanosensing. PLoS Comput. Biol. 11, e1004593 (2015).
Zhou, J., Bronowska, A., Le Coq, J., Lietha, D. & Gräter, F. Allosteric regulation of focal adhesion kinase by PIP2 and ATP. Biophys. J. 108, 698–705 (2015).
Plotnikov, S. V., Pasapera, A. M., Sabass, B. & Waterman, C. M. Force fluctuations within focal adhesions mediate ECM-rigidity sensing to guide directed cell migration. Cell 151, 1513–1527 (2012).
Pollard, T. D. & Cooper, J. A. Actin, a central player in cell shape and movement. Science 326, 1208–1212 (2009).
Stutchbury, B., Atherton, P., Tsang, R., Wang, D.-Y. & Ballestrem, C. Distinct focal adhesion protein modules control different aspects of mechanotransduction. J. Cell Sci. 130, 1612–1624 (2017).
Acebrón, I. et al. Structural basis of focal adhesion kinase activation on lipid membranes. EMBO J. 39, e104743 (2020).
Panagiotakopoulou, M. et al. Cell cycle-dependent force transmission in cancer cells. Mol. Biol. Cell 29, 2528–2539 (2018).
Bauer, M. S. et al. Structural and mechanistic insights into mechanoactivation of focal adhesion kinase. Proc. Natl Acad. Sci. USA 116, 6766–6774 (2019).
Zhou, D. W. et al. Force-FAK signaling coupling at individual focal adhesions coordinates mechanosensing and microtissue repair. Nat. Commun. 12, 2359 (2021).
Hail, M. E., Elliott, B. & Anderson, K. High-throughput analysis of oligonucleotides using automated electrospray ionization mass spectrometry. Am. Biotechnol. Lab. 22, 12–14 (2004).
Zhang, Y., Ge, C., Zhu, C. & Salaita, K. DNA-based digital tension probes reveal integrin forces during early cell adhesion. Nat. Commun. 5, 5167 (2014).
Liu, Y., Galior, K., Ma, V. P.-Y. & Salaita, K. Molecular tension probes for imaging forces at the cell surface. Acc. Chem. Res. 50, 2915–2924 (2017).
Gidi, Y., Bayram, S., Ablenas, C. J., Blum, A. S. & Cosa, G. Efficient one-step PEG-silane passivation of glass surfaces for single-molecule fluorescence studies. ACS Appl. Mater. Interfaces 10, 39505–39511 (2018).
Woodside, M. T. et al. Nanomechanical measurements of the sequence-dependent folding landscapes of single nucleic acid hairpins. Proc. Natl Acad. Sci. USA 103, 6190–6195 (2006).
Preibisch, S., Saalfeld, S., Schindelin, J. & Tomancak, P. Software for bead-based registration of selective plane illumination microscopy data. Nat. Methods 7, 418–419 (2010).
Acknowledgements
We would like to thank S. Narum for carrying out FLIM and analysis, W. Weiss for sharing cell lines, W. Degrado and D. Sheppard for sharing integrin inhibitors and antibodies and O. Weiner and D. Sheppard for constructive comments. Funding for this work was provided by NIH NIGMS R35GM131766 to X.S. and NIH NIGMS R01GM131099 and R01GM124472 to K.S.
Author information
Authors and Affiliations
Contributions
X.S. initiated the project. X.L. and X.S. designed the experiments and analyzed the data. X.L. conducted the experiments. D.C. and K.S. designed the tension experiments and analyzed the data. X.L., D.C., K.S. and X.S. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Chemical Biology thanks Daniel Lietha, Adam W. Smith and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 FAK-SPARK forms droplets after removal of the FAK inhibitor.
Data are mean ± SEM (n = 3 biological replicates). Scale bar, 10 μm.
Extended Data Fig. 2 FAK-SPARK is applicable to various cancer cells in imaging FAK activity using a lentivirus expressing FAK-SPARK.
Data are mean ± SEM, n = 5 biological replicates, two-sided nonpaired t-test. p = 5.52 × 10−7 for MDA-MB-231, p = 0.00056 for Kelly, p = 0.0026 for MEF, p = 2.64 × 10−8 for SHEP, p = 0.0019 for SKNAS and p = 8.17 × 10−6 for U2OS). **: p value < 0.01. ***: p value < 0.001. Scale bar, 10 μm.
Extended Data Fig. 3 FAK-SPARK visualizes FAK activity with spatiotemporal resolution by targeting active FAK into specific locations.
a Fluorescence images showing that FAK-SPARK visualizes FAK activity in the centrosome when constitutively active FAK (CA-FAK) is fused to aurora A kinase that is located in the centrosome (left panels). FAK activity is absent in the centrosome without CA-FAK (right panels). b Fluorescence images showing that nucleus-located FAK-SPARK-NLS (nuclear localization signal) visualizes FAK activity in the nuclear HOTag1 condensates that are tagged with CA-FAK (left panels). FAK activity is absent the CA-FAK-absent HOTag1 condensates. a and b were repeated three times independently with similar results. (c) Cartoon showing experimental procedure of measuring FAK-SPARK temporal resolution. d Time-lapse images after addition of rapamycin. This experiment was repeated three times with similar results. e Normalized fluorescence along the dash line in (D). f Normalized SPARK signal over time. Scale bar, 5 μm (a, b, d).
Extended Data Fig. 4 FAK-SPARK expression does not perturb dynamics of focal adhesion assembly and disassembly.
a, b, Representative images of HeLa cells expressing mApple-paxillin + FAK-SPARK and mApple-paxillin + GFP, respectively. a and b were repeated for three times independently with similar results. c, d, Tracking of FAs using Focal Adhesion Analysis Server {Steenkiste:2021iv, Berginski:2013dj} in a & b. e, A typical trace of assembly and disassembly rate calculation. f, g, Assembly rate and disassembly rate in HeLa cells expressing FAK-SPARK or GFP. two-sided nonpaired t-test, p = 0.12 and 0.10 for f and g respectively. Data represent mean ± SEM (n indicates FA numbers, 305 and 298 for GFP and FAK-SAPRK in f, 331 for both GFP and FAK-SPARK in g). Data represent mean ± SEM. n.s., not significant. Scale= 10 μm.
Extended Data Fig. 5 FAK-SPARK expression does not perturb cell migration during wound healing using wound scratch assay.
Wound was induced in HeLa cells expressing FAK-SPARK using a scratcher. a, b, Representative images of HeLa cells expressing FAK-SPARK or GFP, respectively. c, d, Typical images showing wound healing right after scratching (day 0), one day (day 1) and two days (day 2) after scratching. The red line marks cell boundary. a, b, c and d were repeated three times independently with similar results. e, Quantification of wound healing, n = 3 biological replicates. two-sided nonpaired t-test, p = 0.65 and 0.46 for Day 1 and Day 2. Data represent mean ± SEM. n.s., not significant. Scale bar, 20 μm for A & B and 200 μm for c & d.
Extended Data Fig. 6 Integrin-ECM ligand tension precedes FAK activity.
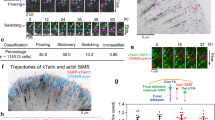
a Diagram of 19 pN DNA-based MTFM probes and FAK-SPARK activation mechanism. b Representative cell RICM (gray) and fluorescence micrographs of 19 pN hairpin tension(red), IFP2-Paxillin(magenta), and FAK-SPARK(green) at t = 0.0 min. Scale bar, 5 μm. c Overlayed kymographs, yellow line in d, of tension, FAK-SPARK, and paxillin channels. Yellow arrows denote the point of tension and paxillin recruitment and FAK-SPARK droplet formation respectively. d Individual fluorescence micrographs of yellow inset in b at different timepoints, yellow dotted line denotes the linear ROI used for kymograph analysis in (c). Scale bar, 2 μm. e Plot of normalized maximum fluorescence over time derived from the kymograph, with threshold points used to derive the time delay denoted with black circles f Histogram of time delay measurements yielding 21.0 ± 9.0 min average delay between 19 pN integrin tension and FAK-SPARK droplet formation. n = 19 focal adhesions from 4 cells on 4 different surfaces (4 biological replicates).
Supplementary information
Supplementary Information
Supplementary Figs. 1–24, Tables 1 and 2, Videos 1–15, References and Supplementary Fig. 9A (uncropped data of blots).
Supplementary Video 1
FAK–SPARK achieves a spatiotemporal resolution in visualizing FAK activity in living cells. HOTag1–NLS–mKO3–Frb (blue), FAK–SPARK–NLS (green) and FKBP–IFP2–CA–FAK (red). Images were taken every 30 s per frame.
Supplementary Video 2
FAK activation at the leading of a spreading cell. FAK–SPARK (green) and mApple–paxillin (red). Images were taken every 30 s per frame.
Supplementary Video 3
FAK activity in a living cell. FAK–SPARK (green) and mApple–paxillin (red). Images were taken every 30 s per frame.
Supplementary Video 4
An ROI showing FAK activity in a living cell. FAK–SPARK (green) and mApple–paxillin (red). Images were taken every 30 s per frame.
Supplementary Video 5
FAK activation during assembly of a single FA in cells. FAK–SPARK (green) and mApple–paxillin (red). Images were taken every 30 s per frame.
Supplementary Video 6
FAK activation before disassembly of FAs in cells. FAK–SPARK (green) and mApple–paxillin (red). Images were taken every 30 s per frame.
Supplementary Video 7
Disassembly of a single FA following FAK activation. FAK–SPARK (green) and mApple–paxillin (red). Images were taken every 30 s per frame.
Supplementary Video 8
FAK activation during sliding of single FAs. FAK–SPARK (green) and mApple–paxillin (red). Images were taken every 30 s per frame.
Supplementary Video 9
FAK activation during the turnover of FAs in cells. FAK–SPARK (green) and mApple–paxillin (red). Images were taken every 30 s per frame.
Supplementary Video 10
An ROI showing FAK activation during the turnover of FAs in cells. FAK–SPARK (green) and mApple–paxillin (red). Images were taken every 30 s per frame.
Supplementary Video 11
Polarized FAK activity at the distal tip of newly formed single FAs in a migrating cell. FAK–SPARK (green) and mApple–paxillin (red). Images were taken every 30 s per frame.
Supplementary Video 12
No distal polarization of FAK activity when CA–FAK is fused to paxillin and targeted to FAs in cells with endogenous FAK knocked down by shRNA against FAK. FAK–SPARK (green) and mApple–paxillin (red). Images were taken every 30 s per frame.
Supplementary Video 13
Nineteen piconewton hairpin integrin tension, IFP2–paxillin and TIRF-imaged FAK–SPARK droplets generated at regions experiencing 19 pN tension and enriched in paxillin.
Supplementary Video 14
TGT rupture (red), IFP2–paxillin (blue) and FAK–SPARK droplets (green), generated overtime on 12 pN TGTs.
Supplementary Video 15
TGT rupture (red), IFP2–paxillin (blue) and FAK–SPARK droplets (green), generated overtime on 56 pN TGTs.
Supplementary Data
Supporting data for Supplementary Figs. 1, 3, 5–9 and 22–24.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 6
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, X., Combs, J.D., Salaita, K. et al. Polarized focal adhesion kinase activity within a focal adhesion during cell migration. Nat Chem Biol 19, 1458–1468 (2023). https://doi.org/10.1038/s41589-023-01353-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-023-01353-y