Abstract
Tissues are organized in cellular niches, the composition and interactions of which can be investigated using spatial omics technologies. However, systematic analyses of tissue composition are challenged by the scale and diversity of the data. Here we present CellCharter, an algorithmic framework to identify, characterize, and compare cellular niches in spatially resolved datasets. CellCharter outperformed existing approaches and effectively identified cellular niches across datasets generated using different technologies, and comprising hundreds of samples and millions of cells. In multiple human lung cancer cohorts, CellCharter uncovered a cellular niche composed of tumor-associated neutrophil and cancer cells expressing markers of hypoxia and cell migration. This cancer cell state was spatially segregated from more proliferative tumor cell clusters and was associated with tumor-associated neutrophil infiltration and poor prognosis in independent patient cohorts. Overall, CellCharter enables systematic analyses across data types and technologies to decode the link between spatial tissue architectures and cell plasticity.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The DLPFC Visium dataset was downloaded from the project’s GitHub page (https://github.com/LieberInstitute/HumanPilot). The mouse spleen CODEX dataset was obtained from the original publication5 (https://data.mendeley.com/datasets/zjnpwh8m5b/1). The NSCLC CosMx dataset was obtained from the original publication41 (https://nanostring.com/products/cosmx-spatial-molecular-imager/nsclc-ffpe-dataset/). The lung cancer MERFISH data was downloaded from Vizgen’s website (https://info.vizgen.com/ffpe-showcase). The lung cancer IMC dataset was obtained upon request to the corresponding author. The extended single-cell lung cancer atlas (LuCA)63 was downloaded from cellxgene (https://cellxgene.cziscience.com/collections/edb893ee-4066-4128-9aec-5eb2b03f8287). The mouse brain RNA + ATAC multiomics dataset was downloaded from the UCSC Cell and Genome Browser (https://brain-spatial-omics.cells.ucsc.edu/). The lung cancer bulk RNA-seq datasets were retrieved from the original publications11,64,65,66,67,68,69,70,71 through the following links and Gene Expression Omnibus accession numbers: https://doi.org/10.5281/zenodo.3941896, https://gdac.broadinstitute.org/ (LUAD, rnaseqv2-RSEM_genes), GSE68465, GSE72094, GSE31210, GSE50081, https://src.gisapps.org/OncoSG_public/ (GIS031), GSE37745 and https://xenabrowser.net/datapages/.
Code availability
CellCharter is released as an open-source Python library on GitHub at https://github.com/CSOgroup/cellcharter. Code to reproduce all the analyses presented in this manuscript is available at https://github.com/CSOgroup/cellcharter_analyses.
References
Chappell, L., Russell, A. J. C. & Voet, T. Single-cell (multi)omics technologies. Annu. Rev. Genomics Hum. Genet. 19, 15–41 (2018).
Marx, V. Method of the year: spatially resolved transcriptomics. Nat. Methods 18, 9–14 (2021).
Moffitt, J. R., Lundberg, E. & Heyn, H. The emerging landscape of spatial profiling technologies. Nat. Rev. Genet. 23, 741–759 (2022).
Moses, L. & Pachter, L. Museum of spatial transcriptomics. Nat. Methods 19, 534–546 (2022).
Goltsev, Y. et al. Deep profiling of mouse splenic architecture with CODEX multiplexed imaging. Cell 174, 968–981 (2018).
Ståhl, P. L. et al. Visualization and analysis of gene expression in tissue sections by spatial transcriptomics. Science 353, 78–82 (2016).
Callaway, E. M. et al. A multimodal cell census and atlas of the mammalian primary motor cortex. Nature 598, 86–102 (2021).
Bassiouni, R., Gibbs, L. D., Craig, D. W., Carpten, J. D. & McEachron, T. A. Applicability of spatial transcriptional profiling to cancer research. Mol. Cell 81, 1631–1639 (2021).
Hunter, M. V., Moncada, R., Weiss, J. M., Yanai, I. & White, R. M. Spatially resolved transcriptomics reveals the architecture of the tumor-microenvironment interface. Nat. Commun. 12, 6278 (2021).
Keren, L. et al. A structured tumor-immune microenvironment in triple negative breast cancer revealed by multiplexed ion beam imaging. Cell 174, 1373–1387 (2018).
Tavernari, D. et al. Nongenetic evolution drives lung adenocarcinoma spatial heterogeneity and progression. Cancer Discov. 11, 1490–1507 (2021).
Karras, P. et al. A cellular hierarchy in melanoma uncouples growth and metastasis. Nature 610, 190–198 (2022).
Lomakin, A. et al. Spatial genomics maps the structure, nature and evolution of cancer clones. Nature 611, 594–602 (2022).
Lundberg, E. & Borner, G. H. H. Spatial proteomics: a powerful discovery tool for cell biology. Nat. Rev. Mol. Cell Biol. 20, 285–302 (2019).
Tian, L., Chen, F. & Macosko, E. Z. The expanding vistas of spatial transcriptomics.Nat. Biotechnol. 41, 773–782 (2022).
Rao, A., Barkley, D., França, G. S. & Yanai, I. Exploring tissue architecture using spatial transcriptomics. Nature 596, 211–220 (2021).
Liu, Y. et al. High-spatial-resolution multi-omics sequencing via deterministic barcoding in tissue. Cell 183, 1665–1681 (2020).
Vickovic, S. et al. High-definition spatial transcriptomics for in situ tissue profiling. Nat. Methods 16, 987–990 (2019).
Cho, C.-S. et al. Microscopic examination of spatial transcriptome using Seq-Scope. Cell 184, 3559–3572 (2021).
Chen, A. et al. Spatiotemporal transcriptomic atlas of mouse organogenesis using DNA nanoball-patterned arrays. Cell 185, 1777–1792 (2022).
Fu, X. et al. Polony gels enable amplifiable DNA stamping and spatial transcriptomics of chronic pain. Cell 185, 4621–4633 (2022).
Zhang, D. et al. Spatial epigenome–transcriptome co-profiling of mammalian tissues. Nature 616, 113–122 (2023).
Jackson, H. W. et al. The single-cell pathology landscape of breast cancer. Nature 578, 615–620 (2020).
Sorin, M. et al. Single-cell spatial landscapes of the lung tumour immune microenvironment. Nature 614, 548–554 (2023).
Chatzis, S. P. & Tsechpenakis, G. The infinite hidden Markov random field model. IEEE Trans. Neural Netw. 21, 1004–1014 (2010).
Scarselli, F., Gori, M., Tsoi, A. C., Hagenbuchner, M. & Monfardini, G. The graph neural network model. IEEE Trans. Neural Netw. 20, 61–80 (2009).
Kipf, T. N. & Welling, M. Semi-supervised classification with graph convolutional networks. Preprint at https://doi.org/10.48550/arXiv.1609.02907 (2017).
Zhao, E. et al. Spatial transcriptomics at subspot resolution with BayesSpace. Nat. Biotechnol. 39, 1375–1384 (2021).
Liu, W. et al. Joint dimension reduction and clustering analysis of single-cell RNA-seq and spatial transcriptomics data. Nucleic Acids Res. 50, e72 (2022).
Hu, J. et al. SpaGCN: integrating gene expression, spatial location and histology to identify spatial domains and spatially variable genes by graph convolutional network. Nat. Methods 18, 1342–1351 (2021).
Fu, H. et al. Unsupervised spatially embedded deep representation of spatial transcriptomics. Preprint at bioRxiv https://doi.org/10.1101/2021.06.15.448542 (2021).
Dong, K. & Zhang, S. Deciphering spatial domains from spatially resolved transcriptomics with an adaptive graph attention auto-encoder. Nat. Commun. 13, 1739 (2022).
Wu, Z. et al. Graph deep learning for the characterization of tumour microenvironments from spatial protein profiles in tissue specimens. Nat. Biomed. Eng. 6, 1435–1448 (2022).
Kim, J. et al. Unsupervised discovery of tissue architecture in multiplexed imaging. Nat. Methods 19, 1653–1661 (2022).
Shang, L. & Zhou, X. Spatially aware dimension reduction for spatial transcriptomics. Nat. Commun. 13, 7203 (2022).
Yuan, Z. et al. SOTIP is a versatile method for microenvironment modeling with spatial omics data. Nat. Commun. 13, 7330 (2022).
Danenberg, E. et al. Breast tumor microenvironment structures are associated with genomic features and clinical outcome. Nat. Genet. 54, 660–669 (2022).
Karimi, E. et al. Single-cell spatial immune landscapes of primary and metastatic brain tumours. Nature 614, 555–563 (2023).
Ali, H. R. et al. Imaging mass cytometry and multiplatform genomics define the phenogenomic landscape of breast cancer. Nat. Cancer 1, 163–175 (2020).
Zhang, M. et al. Spatially resolved cell atlas of the mouse primary motor cortex by MERFISH. Nature 598, 137–143 (2021).
He, S. et al. High-plex imaging of RNA and proteins at subcellular resolution in fixed tissue by spatial molecular imaging. Nat. Biotechnol. 40, 1794–1806 (2022).
Kingma, D. P. & Welling, M. Auto-encoding variational Bayes. Preprint at https://doi.org/10.48550/arXiv.1312.6114 (2014).
Lopez, R., Regier, J., Cole, M. B., Jordan, M. I. & Yosef, N. Deep generative modeling for single-cell transcriptomics. Nat. Methods 15, 1053–1058 (2018).
Lotfollahi, M. et al. Mapping single-cell data to reference atlases by transfer learning. Nat. Biotechnol. 40, 121–130 (2022).
Fowlkes, E. B. & Mallows, C. L. A method for comparing two hierarchical clusterings. J. Am. Stat. Assoc. 78, 553–569 (1983).
Palla, G. et al. Squidpy: a scalable framework for spatial omics analysis. Nat. Methods 19, 171–178 (2022).
Maynard, K. R. et al. Transcriptome-scale spatial gene expression in the human dorsolateral prefrontal cortex. Nat. Neurosci. 24, 425–436 (2021).
Akaike, H. A new look at the statistical model identification. IEEE Trans. Automat. Contr. 19, 716–723 (1974).
Schwarz, G. Estimating the dimension of a model. Ann. Stat. 6, 461–464 (1978).
Allen Institute for Brain Science. Adult mouse—coronal sections. Allen Brain Atlas https://atlas.brain-map.org (2011).
Ramiscal, R. R. & Vinuesa, C. G. T-cell subsets in the germinal center. Immunol. Rev. 252, 146–155 (2013).
Pusztaszeri, M. P., Seelentag, W. & Bosman, F. T. Immunohistochemical expression of endothelial markers CD31, CD34, von Willebrand Factor, and Fli-1 in normal human tissues. J. Histochem. Cytochem. 54, 385–395 (2006).
Greaves, M. & Maley, C. C. Clonal evolution in cancer. Nature 481, 306–313 (2012).
Yuan, S., Norgard, R. J. & Stanger, B. Z. Cellular plasticity in cancer. Cancer Discov. 9, 837–851 (2019).
Swanton, C. Intratumor heterogeneity: evolution through space and time. Cancer Res. 72, 4875–4882 (2012).
Black, J. R. M. & McGranahan, N. Genetic and non-genetic clonal diversity in cancer evolution. Nat. Rev. Cancer 21, 379–392 (2021).
Wang, Y. et al. N-myc downstream regulated gene 1 (NDRG1) promotes the stem-like properties of lung cancer cells through stabilized c-Myc. Cancer Lett. 401, 53–62 (2017).
Ma, J., Gao, Q., Zeng, S. & Shen, H. Knockdown of NDRG1 promote epithelial–mesenchymal transition of colorectal cancer via NF-κB signaling. J. Surg. Oncol. 114, 520–527 (2016).
Zhu, H. & Zhang, S. Hypoxia inducible factor-1α/vascular endothelial growth factor signaling activation correlates with response to radiotherapy and its inhibition reduces hypoxia-induced angiogenesis in lung cancer. J. Cell. Biochem. 119, 7707–7718 (2018).
Rajarathnam, K., Schnoor, M., Richardson, R. M. & Rajagopal, S. How do chemokines navigate neutrophils to the target site: dissecting the structural mechanisms and signaling pathways. Cell. Signal. 54, 69–80 (2019).
Behrens, C. et al. EZH2 protein expression associates with the early pathogenesis, tumor progression, and prognosis of non-small cell lung carcinoma. Clin. Cancer Res. 19, 6556–6565 (2013).
Valadez-Cosmes, P., Raftopoulou, S., Mihalic, Z. N., Marsche, G. & Kargl, J. Myeloperoxidase: growing importance in cancer pathogenesis and potential drug target. Pharmacol. Ther. 236, 108052 (2022).
Salcher, S. et al. High-resolution single-cell atlas reveals diversity and plasticity of tissue-resident neutrophils in non-small cell lung cancer. Cancer Cell 40, 1503–1520 (2022).
Collisson, E. A. et al. Comprehensive molecular profiling of lung adenocarcinoma. Nature 511, 543–550 (2014).
Shedden, K. et al. Gene expression-based survival prediction in lung adenocarcinoma: a multi-site, blinded validation study. Nat. Med. 14, 822–827 (2008).
Schabath, M. B. et al. Differential association of STK11 and TP53 with KRAS mutation-associated gene expression, proliferation and immune surveillance in lung adenocarcinoma. Oncogene 35, 3209–3216 (2016).
Okayama, H. et al. Identification of genes upregulated in ALK-positive and EGFR/KRAS/ALK-negative lung adenocarcinomas. Cancer Res. 72, 100–111 (2012).
Der, S. D. et al. Validation of a histology-independent prognostic gene signature for early-stage, non–small-cell lung cancer including stage IA patients. J. Thorac. Oncol. 9, 59–64 (2014).
Chen, J. et al. Genomic landscape of lung adenocarcinoma in East Asians. Nat. Genet. 52, 177–186 (2020).
Mezheyeuski, A. et al. Multispectral imaging for quantitative and compartment-specific immune infiltrates reveals distinct immune profiles that classify lung cancer patients. J. Pathol. 244, 421–431 (2018).
Ding, L. et al. Somatic mutations affect key pathways in lung adenocarcinoma. Nature 455, 1069–1075 (2008).
Virshup, I. et al. The scverse project provides a computational ecosystem for single-cell omics data analysis. Nat. Biotechnol. 41, 604–606 (2023).
Deng, Y. et al. Spatial-CUT&Tag: spatially resolved chromatin modification profiling at the cellular level. Science 375, 681–686 (2022).
Zhao, T. et al. Spatial genomics enables multi-modal study of clonal heterogeneity in tissues. Nature 601, 85–91 (2022).
Deng, Y. et al. Spatial profiling of chromatin accessibility in mouse and human tissues. Nature 609, 375–383 (2022).
Vickovic, S. et al. SM-Omics is an automated platform for high-throughput spatial multi-omics. Nat. Commun. 13, 795 (2022).
Ben-Chetrit, N. et al. Integration of whole transcriptome spatial profiling with protein markers.Nat. Biotechnol. 41, 788–793 (2023).
Chen, R. J. et al. Scaling vision transformers to gigapixel images via hierarchical self-supervised learning. In Proc. IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR) 16144–16155, (IEEE, 2022).
Blaisdell, A. et al. Neutrophils oppose uterine epithelial carcinogenesis via debridement of hypoxic tumor cells. Cancer Cell 28, 785–799 (2015).
Yee, P. P. et al. Neutrophil-induced ferroptosis promotes tumor necrosis in glioblastoma progression. Nat. Commun. 11, 5424 (2020).
Su, H. et al. Identification of hub genes associated with neutrophils infiltration in colorectal cancer. J. Cell. Mol. Med. 25, 3371–3380 (2021).
Bruni, D., Angell, H. K. & Galon, J. The immune contexture and Immunoscore in cancer prognosis and therapeutic efficacy. Nat. Rev. Cancer 20, 662–680 (2020).
Howard, R., Kanetsky, P. A. & Egan, K. M. Exploring the prognostic value of the neutrophil-to-lymphocyte ratio in cancer. Sci. Rep. 9, 19673 (2019).
Hagberg, A., Swart, P. J. & Schult, D. A. Exploring network structure, dynamics, and function using networkX. In Proceedings of the 7th Python in Science conference (SciPy 08) (eds Varoquaux, G., Vaught, T. & Millman, J.) 11–15 (OSTI.GOV, 2008)
Martens, L. D., Fischer, D. S., Theis, F. J. & Gagneur, J. Modeling fragment counts improves single-cell ATAC-seq analysis. Preprint at bioRxiv https://doi.org/10.1101/2022.05.04.490536 (2022).
Edelsbrunner, H., Kirkpatrick, D. & Seidel, R. On the shape of a set of points in the plane. IEEE Trans. Inf. Theory 29, 551–559 (1983).
Zhang, T. Y. & Suen, C. Y. A fast parallel algorithm for thinning digital patterns. Commun. ACM 27, 236–239 (1984).
Walt van der, S. et al. scikit-image: image processing in Python. PeerJ 2, e453 (2014).
Luecken, M. D. et al. Benchmarking atlas-level data integration in single-cell genomics. Nat. Methods 19, 41–50 (2022).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Johnson, W. E., Li, C. & Rabinovic, A. Adjusting batch effects in microarray expression data using empirical Bayes methods. Biostatistics 8, 118–127 (2007).
Acknowledgements
We thank R. Gottardo (Centre hositalier universitaire vaudois, Lausanne), D. Gfeller (University of Lausanne), G. La Manno and E. Oricchio (École Polytechnique Fédérale de Lausanne) for their precious feedback during the development of this project. This project was in part supported by the Swiss Institute for Experimental Cancer Research TANDEM Research Project (G.C.). D.T. is supported by the Personalized Health and Related Technologies (PHRT) iPostdoc Project (grant no. 2022-476). A.S.-M. is supported by the PHRT Technology transfer project (grant no. 2021-553).
Author information
Authors and Affiliations
Contributions
M.V. designed the algorithm and performed the computational analyses. D.T. analyzed the RNA-seq datasets. A.S.-M. contributed to the interpretation of the CODEX mouse spleen results. L.A.W. provided the data for the IMC lung cancer dataset and contributed to the interpretation of the results. M.V. and G.C. designed the project and wrote the manuscript with contributions from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Genetics thanks Naveed Ishaque, Hamid Ali, Shashwat Sahay and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 CellCharter’s features and benchmarking of spatial clustering methods.
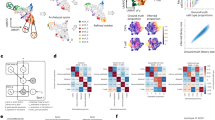
a, Example of 3 spatial clusters (color coded) in a tissue sample of mouse spleen analyzed by CODEX (left) and symmetric (top right) vs. asymmetric (bottom right) neighborhood enrichment analysis. b, Schematic representation of the four metrics (curl, elongation, linearity, and purity) implemented in CellCharter to describe the shape of spatial clusters. c, Example of spatial cluster components (color coded) in a tissue sample of mouse spleen analyzed by CODEX (left). Heatmap representation of the shape metric values for each cluster component (right). Cluster components are grouped in representative shape classes: linear, round, circular, and irregular. d, Manual annotations of the Visium DLPFC samples. e, Previously adopted strategy based on hyperparameter tuning and clustering testing on the same dataset. f, strategy proposed in this study based on independent tuning and testing datasets. g, Mean Adjusted Rand Index (ARI) for each DLPFC sample (over 10 repetitions) obtained by the listed methods upon performing individual spatial clustering of the samples (n = 9 samples). The boxes show the quartiles of the dataset while the whiskers extend to points that lie within 1.5 inter-quartile ranges (IQRs) of the lower and upper quartiles. h, ARI for each DLPFC sample obtained by the listed methods upon performing joint clustering of all samples (n = 9 samples). The boxes show the quartiles of the dataset while the whiskers extend to points that lie within 1.5 IQRs of the lower and upper quartiles.
Extended Data Fig. 2 Comparison between CellCharter STAGATE and evaluation of cluster stability analysis.
a, Runtime divided by processing step of the two best-performing methods (STAGATE and CellCharter) in clustering all 12 samples of the DLPFC dataset on GPU and CPU. b, From left to right: Bayesian Information Criterion (BIC), Akaike Information Criterion (AIC), and Negative Log-Likelihood (NLL) values (y-axis) for a number of clusters ranging between 2 and 100 (x-axis) for the pilot (12 samples) and full (42 samples) version of the DLPFC dataset. c, Purity of the n = 756 cluster components in the CODEX mouse spleen dataset for the clusters of the two best-performing methods. The boxes show the quartiles of the dataset while the whiskers extend to points that lie within 1.5 inter-quartile ranges (IQRs) of the lower and upper quartiles. d, CellCharter cluster stability for range of numbers of cluster (x-axis) using a concatenation of the embeddings from chromatin accessibility and gene expression. The most stable cluster solution is highlighted. Data presented as mean values (solid line) with a 95% confidence interval (shaded area).
Extended Data Fig. 3 Characterization of CellCharter’s spatial clusters of the CODEX mouse spleen dataset.
a, CellCharter’s spatial cluster at n = 11 clusters for all cells of the 3 healthy (BALBc) and 6 systemic lupus erythematosus samples (MRL). b, (left) Spatial distribution of cells assigned to spatial cluster C1 in three selected samples. (right) Representative images of immunofluorescence staining artifacts corresponding to C1: autofluorescence (1), missing tile (2), fluorophore accumulation (3).
Extended Data Fig. 4 Differences in the spatial organization between healthy and systemic lupus spleen.
a, Spatial distribution of CD4+ T cells, B cells, and B220+ T cells in the clusters associated to the germinal center, marginal zone, and GC-PALS boundary for representative examples of healthy (BALBc-1), early lupus (MRL-4) and intermediate lupus (MRL-8) spleen. b, Normalized intensity of ERTR7, CD31, and Ly6G markers in the intermediate lupus sample MRL-5. The MRL samples show the presence of two distinct clusters associated with trabecular structures. Insets (1 to 6) highlight the different expression of the markers in correspondence trabecular clusters C10 (insets 2, 4, 6) and C11 (insets 1, 3, 5).
Extended Data Fig. 5 Characterization of CellCharter’s spatial clusters of the CosMx non-small cell lung cancer (NSCLC) dataset.
a, CellCharter’s spatial cluster at n = 20 clusters for all cells of the CosMx NSCLC samples. b, Entropy of patient proportion between n = 10 tumor microenvironment-enriched (TME-enriched) clusters and n = 10 tumor-enriched clusters. The thick central line of each box plot represents the median entropy, the bounding box corresponds to the 25th–75th percentiles, and the whiskers extend up to 1.5 times the interquartile range. p-values computed by two-tailed Wilcoxon test. c, Immune and stromal cell type fractions in tumor samples from 5 non-small cell lung cancer patients (LUAD: lung adenocarcinoma, LUSC: lung squamous cell carcinoma, NK-cell: natural killer cell).
Extended Data Fig. 6 Cell signature scores in two non-small cell lung cancer (NSCLC) patients.
a, Gene expression signature score of n = 41,088 tumor cells in spatial clusters C0 and C12. The boxes show the quartiles of the dataset while the whiskers extend to points that lie within 1.5 inter-quartile ranges (IQRs) of the lower and upper quartile. b-e, Gene expression signature score of tumor cells of patients LUAD-9 and LUAD-12 for four gene signatures: neutrophil chemotaxis, hypoxia, epithelial-to-mesenchymal transition (EMT-II) and cell proliferation. The boxes show the quartiles of the dataset while the whiskers extend to points that lie within 1.5 IQRs of the lower and upper quartiles.
Extended Data Fig. 7 Spatial organization of tumor associated neutrophils and hypoxic tumor cells in CellCharter and STAGATE.
a, Average hypoxia gene signature score for tumor cells at increasing hop-distance from all neutrophils (left), neutrophils assigned to spatial cluster C11 (center), and neutrophils not assigned in cluster C11 (right). Error bands correspond to the 95% confidence interval. b, cluster concordance between CellCharter (rows) and STAGATE (columns). c, Example of CellCharter and STAGATE clusters on LUAD-9 R1. d, Hypoxia and cell proliferation signatures in n = 128,454 cells of tumor-enriched spatial clusters determined with STAGATE (NK: natural killer cell, Treg: regulatory T cell, mDC: myeloid dendritic cell, pDC: plasmacytoid dendritic cell). The boxes show the quartiles of the dataset while the whiskers extend to points that lie within 1.5 inter-quartile ranges (IQRs) of the lower and upper quartiles. e, Cell type enrichment of tumor-enriched clusters and neutrophil-enriched clusters obtained with STAGATE. f, Coarse cell type labels in LUAD-9 R1. g, Comparison of neutrophil-enriched clusters detected by STAGATE (left, C13) and CellCharter (right, C11).
Extended Data Fig. 8 Association between tumor hypoxia and neutrophil infiltration in the MERFISH and Image Mass Cytometry (IMC) datasets.
a, Cluster stability (y-axis) for range of numbers of cluster (x-axis) on the full MERFISH dataset. Data presented as mean values (solid line) with a 95% confidence interval (shaded area). b, Runtime divided by processing step of CellCharter in clustering the MERFISH dataset (left) and IMC dataset (right) on GPU and CPU. c, Cluster stability (y-axis) for range of numbers of cluster (x-axis) on the 2 lung cancer samples of the MERFISH dataset. Data presented as mean values (solid line) with a 95% confidence interval (shaded area). d, Cell type enrichment of tumor-enriched and tumor microenvironment-enriched (TME-enriched) clusters in HumanLungCancerPatient1 obtained with CellCharter. e, Neighborhood enrichment between the TME-enriched clusters (source) and the tumor-enriched clusters (target) in HumanLungCancerPatient1. p-values computed by unpaired two-sided t-test. f, Cell type enrichment analysis (left) and hazard ratio and Cox-regression p-values (right) for the clusters C15 and C17 identified by CellCharter in n = 413 samples of the IMC dataset. Data are presented as mean values with 95% confidence interval. g, Cell type enrichment of tumor-enriched and TME-enriched clusters in the IMC dataset obtained with CellCharter. Only TME clusters for which there is a positive neighborhood enrichment with at least one tumor-enriched cluster are shown.
Extended Data Fig. 9 Association of tumor-associated neutrophils (TANs), normal-associated neutrophils (NANs), and hypoxia with overall survival.
a, Pearson correlation between hypoxia gene signature score and TAN score in 9 bulk RNA-seq datasets of lung adenocarcinoma11,64,65,66,67,68,69,70,71. Data are presented as mean values (solid line) with a 95% confidence interval (shaded area). b, Pearson correlation between hypoxia gene signature score and NAN score in 9 bulk RNA-seq datasets of lung adenocarcinoma. Data are presented as mean values (solid line) with a 95% confidence interval (shaded area). c, Association of TAN, NAN, and hypoxia score with overall survival, corrected for sex, age, and stage, in the 7 bulk RNA-seq datasets that contained survival information64,65,66,67,68,69,70. Data are presented as mean values with 95% confidence interval.
Supplementary information
Supplementary Information
Supplementary methods and legends to Supplementary Tables 1–3.
Supplementary Table 1
Supplementary Tables 1–3.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Varrone, M., Tavernari, D., Santamaria-Martínez, A. et al. CellCharter reveals spatial cell niches associated with tissue remodeling and cell plasticity. Nat Genet 56, 74–84 (2024). https://doi.org/10.1038/s41588-023-01588-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-023-01588-4
This article is cited by
-
Charting multicellular tissue structure cell-to-cell
Nature Genetics (2024)
-
Semi-automated approaches for interrogating spatial heterogeneity of tissue samples
Scientific Reports (2024)