Abstract
Linkage disequilibrium (LD) is the correlation among nearby genetic variants. In genetic association studies, LD is often modeled using large correlation matrices, but this approach is inefficient, especially in ancestrally diverse studies. In the present study, we introduce LD graphical models (LDGMs), which are an extremely sparse and efficient representation of LD. LDGMs are derived from genome-wide genealogies; statistical relationships among alleles in the LDGM correspond to genealogical relationships among haplotypes. We published LDGMs and ancestry-specific LDGM precision matrices for 18 million common variants (minor allele frequency >1%) in five ancestry groups, validated their accuracy and demonstrated order-of-magnitude improvements in runtime for commonly used LD matrix computations. We implemented an extremely fast multiancestry polygenic prediction method, BLUPx-ldgm, which performs better than a similar method based on the reference LD correlation matrix. LDGMs will enable sophisticated methods that scale to ancestrally diverse genetic association data across millions of variants and individuals.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
LDGMs, LDGM precision matrices and tree sequences are available from Zenodo (ref. 84; https://doi.org/10.5281/zenodo.8157131). High-coverage phased 1000 Genomes genotype data are available at http://ftp.1000genomes.ebi.ac.uk/vol1/ftp/data_collections/1000G_2504_high_coverage/working/20201028_3202_phased. LD-independent blocks are available at https://github.com/jmacdon/LDblocks_GRCh38. UK Biobank summary statistics and LD are available at s3://broad-alkesgroup-ukbb-ld/UKBB_LD/. Ancestral states are available via Ensembl release 100 and can be downloaded from ftp://ftp.ensembl.org/pub/release-100/fasta/ancestral_alleles (ref. 83).
Code availability
We have released an open-source software package, ldgm v.0.1, implemented in python and MATLAB. ldgm allows inference of LDGMs and LDGM precision matrices, as well as computationally efficient analyses of GWAS summary statistics using LDGMs. It is available at https://github.com/awohns/ldgm and is deposited to Zenodo85 (https://doi.org/10.5281/zenodo.8161389). All the functions for analyzing GWAS summary statistics with LDGMs, including BLUPx-ldgm, are currently implemented in MATLAB; a Python implementation is planned. BLUPx-ldgm is also implemented in bcftools, available at https://github.com/freeseek/score; tskit is available at https://github.com/tskit-dev/tskit. Scripts to reproduce the results of this manuscript are available at https://github.com/awohns/ldgm_paper.
References
International HapMap Consortium. A haplotype map of the human genome. Nature 437, 1299–1320 (2005).
Reich, D. E. et al. Linkage disequilibrium in the human genome. Nature 411, 199–204 (2001).
Abecasis, G. R. et al. Extent and distribution of linkage disequilibrium in three genomic regions. Am. J. Hum. Genet. 68, 191–197 (2001).
Finucane, H. K. et al. Partitioning heritability by functional annotation using genome-wide association summary statistics. Nat. Genet. 47, 1228–1235 (2015).
Zhu, X. & Stephens, M. Large-scale genome-wide enrichment analyses identify new trait-associated genes and pathways across 31 human phenotypes. Nat. Commun. 9, 4361 (2018).
Vilhjálmsson, B. J. et al. Modeling linkage disequilibrium increases accuracy of polygenic risk scores. Am. J. Hum. Genet. 97, 576–592 (2015).
Weissbrod, O. et al. Leveraging fine-mapping and multipopulation training data to improve cross-population polygenic risk scores. Nat. Genet. 54, 450–458 (2022).
Wang, G., Sarkar, A., Carbonetto, P. & Stephens, M. A simple new approach to variable selection in regression, with application to genetic fine mapping. J. R. Stat. Soc. B Stat. Methodol. 82, 1273–1300 (2020).
Weissbrod, O. et al. Functionally informed fine-mapping and polygenic localization of complex trait heritability. Nat. Genet. 52, 1355–1363 (2020).
Martin, A. R. et al. Clinical use of current polygenic risk scores may exacerbate health disparities. Nat. Genet. 51, 584 (2019).
Peterson, R. E. et al. Genome-wide association studies in ancestrally diverse populations: opportunities, methods, pitfalls, and recommendations. Cell 179, 589–603 (2019).
Coram, M. A., Fang, H., Candille, S. I., Assimes, T. L. & Tang, H. Leveraging multi-ethnic evidence for risk assessment of quantitative traits in minority populations. Am. J. Hum. Genet. 101, 218–226 (2017).
Kichaev, G. & Pasaniuc, B. Leveraging functional-annotation data in trans-ethnic fine-mapping studies. Am. J. Hum. Genet. 97, 260–271 (2015).
Asgari, S. et al. A positively selected FBN1 missense variant reduces height in Peruvian individuals. Nature 582, 234–239 (2020).
Conrad, D. F. et al. A worldwide survey of haplotype variation and linkage disequilibrium in the human genome. Nat. Genet. 38, 1251–1260 (2006).
Li, N. & Stephens, M. Modeling linkage disequilibrium and identifying recombination hotspots using single-nucleotide polymorphism data. Genetics 165, 2213–2233 (2003).
Kelleher, J., Etheridge, A. M. & McVean, G. Efficient coalescent simulation and genealogical analysis for large sample sizes. PLoS Comput. Biol. 12, e1004842 (2016).
Wilder, W. A. et al. A unified genealogy of modern and ancient genomes. Science 375, eabi8264 (2022).
Kelleher, J. et al. Inferring whole-genome histories in large population datasets. Nat. Genet. 51, 1330–1338 (2019).
Speidel, L., Forest, M., Shi, S. & Myers, S. R. A method for genome-wide genealogy estimation for thousands of samples. Nat. Genet. 51, 1321–1329 (2019).
1000 Genomes Project Consortium. et al. A global reference for human genetic variation. Nature 526, 68–74 (2015).
Scheib, C. L. et al. East Anglian early Neolithic monument burial linked to contemporary Megaliths. Ann. Hum. Biol. 46, 145–149 (2019).
Schaefer, N. K., Shapiro, B. & Green, R. E. An ancestral recombination graph of human, Neanderthal, and Denisovan genomes. Sci. Adv. 7, eabc0776 (2022).
Hubisz, M. J., Williams, A. L. & Siepel, A. Mapping gene flow between ancient hominins through demography-aware inference of the ancestral recombination graph. PLoS Genet. 16, e1008895 (2020).
Stern, A. J., Wilton, P. R. & Nielsen, R. An approximate full-likelihood method for inferring selection and allele frequency trajectories from DNA sequence data. PLoS Genet. 15, e1008384 (2019).
Stern, A. J., Speidel, L., Zaitlen, N. A. & Nielsen, R. Disentangling selection on genetically correlated polygenic traits via whole-genome genealogies. Am. J. Hum. Genet. 108, 219–239 (2021).
Fan, C., Mancuso, N. & Chiang, C. W. K. A genealogical estimate of genetic relationships. Am. J. Hum. Genet. 109, 812–824 (2022).
Zhang, B. C., Biddanda, A., Gunnarsson, A. F., Cooper, F. & Palamara, P. F. Biobank-scale inference of ancestral recombination graphs enables genealogical analysis of complex traits. Nat. Genet. 55, 768–776 (2023).
Ernst, J. & Kellis, M. ChromHMM: automating chromatin-state discovery and characterization. Nat. Methods 9, 215–216 (2012).
Border, R. et al. Assortative mating biases marker-based heritability estimators. Nat. Commun 13, 660 (2022).
Border, R. et al. Cross-trait assortative mating is widespread and inflates genetic correlation estimates. Science 378, 754–761 (2022).
Mazumder, R. & Hastie, T. The graphical lasso: new insights and alternatives. Electron J. Stat. 6, 2125–2149 (2012).
Byrska-Bishop, M. et al. High coverage whole genome sequencing of the expanded 1000 Genomes Project cohort including 602 trios. Cell 185, 3426–3440 (2021).
Kelleher, J. et al. Inferring whole-genome histories in large population datasets. Nat. Genet. 51, 1330–1338 (2019).
Berisa, T. & Pickrell, J. K. Approximately independent linkage disequilibrium blocks in human populations. Bioinformatics 32, 283–285 (2016).
MacDonald, J. W., Harrison, T., Bammler, T. K., Mancuso, N. & Lindström, S. An updated map of GRCh38 linkage disequilibrium blocks based on European ancestry data. Preprint at bioRxiv https://doi.org/10.1101/2022.03.04.483057 (2022).
Atkinson, E. G. et al. Tractor uses local ancestry to enable the inclusion of admixed individuals in GWAS and to boost power. Nat. Genet. 53, 195–204 (2021).
Bycroft, C. et al. The UK Biobank resource with deep phenotyping and genomic data. Nature 562, 203–209 (2018).
Kelleher, J., Etheridge, A. M. & McVean, G. Efficient coalescent simulation and genealogical analysis for large sample sizes. PLoS Comput. Biol. 12, e1004842 (2016).
Wen, X. & Stephens, M. Using linear predictors to impute allele frequencies from summary or pooled genotype data. Ann. Appl. Stat. 4, 1158–1182 (2010).
Shi, H., Kichaev, G. & Pasaniuc, B. Contrasting the genetic architecture of 30 complex traits from summary association data. Am. J. Hum. Genet. 99, 139–153 (2016).
Shi, H., Mancuso, N., Spendlove, S. & Pasaniuc, B. Local genetic correlation gives insights into the shared genetic architecture of complex traits. Am. J. Hum. Genet. 101, 737–751 (2017).
Ning, Z., Pawitan, Y. & Shen, X. High-definition likelihood inference of genetic correlations across human complex traits. Nat Genet 52, 859–864 (2020).
Werme, J., van der Sluis, S., Posthuma, D. & de Leeuw, C. A. An integrated framework for local genetic correlation analysis. Nat. Genet. 54, 274–282 (2022).
Spence, J. P., Sinnott-Armstrong, N., Assimes, T. L. & Pritchard, J. K. A flexible modeling and inference framework for estimating variant effect sizes from GWAS summary statistics. Preprint at bioRxiv https://doi.org/10.1101/2022.04.18.488696 (2022).
Pasaniuc, B. & Price, A. L. Dissecting the genetics of complex traits using summary association statistics. Nat. Rev. Genet. 18, 117–127 (2017).
Conneely, K. N. & Boehnke, M. So many correlated tests, so little time! Rapid adjustment of P values for multiple correlated tests. Am. J. Hum. Genet. 81, 1158–1168 (2007).
Zhu, X. & Stephens, M. Bayesian large-scale multiple regression with summary statistics from genoe-wide association studies. Ann. Appl. Stat. 11, 1561–1592 (2017).
Henderson, C. R. Estimation of variance and covariance components. Biometrics 9, 226–252 (1953).
Henderson, C. R. Rapid method for computing the inverse of a relationship matrix. J. Dairy Sci. 58, 1727–1730 (1975).
Henderson, C. R. A simple method for computing the inverse of a numerator relationship matrix used in prediction of breeding values. Biometrics 32, 69–83 (1976).
Ge, T., Chen, C. Y., Ni, Y., Feng, Y. C. A. & Smoller, J. W. Polygenic prediction via Bayesian regression and continuous shrinkage priors. Nat. Commun. 10, 1776 (2019).
Cai, M. et al. A unified framework for cross-population trait prediction by leveraging the genetic correlation of polygenic traits. Am. J. Hum. Genet. 108, 632–655 (2021).
Loh, P.-R. et al. Efficient Bayesian mixed-model analysis increases association power in large cohorts. Nat. Genet. 47, 284–290 (2015).
Lango Allen, H. et al. Hundreds of variants clustered in genomic loci and biological pathways affect human height. Nature 467, 832–838 (2010).
Locke, A. E. et al. Genetic studies of body mass index yield new insights for obesity biology. Nature 518, 197–206 (2015).
Morris, A. P. et al. Large-scale association analysis provides insights into the genetic architecture and pathophysiology of type 2 diabetes. Nat. Genet. 44, 981–990 (2012).
Chen, W. et al. Improved analyses of GWAS summary statistics by reducing data heterogeneity and errors. Nat. Commun. 12, 7117 (2021).
Wray, N. R. et al. Pitfalls of predicting complex traits from SNPs. Nat. Rev. Genet. 14, 507–515 (2013).
O’Connor, L. J. The distribution of common-variant effect sizes. Nat. Genet. 53, 1243–1249 (2021).
Benner, C. et al. Prospects of fine-mapping trait-associated genomic regions by using summary statistics from genome-wide association studies. Am. J. Hum. Genet. 101, 539–551 (2017).
Pasaniuc, B. et al. Fast and accurate imputation of summary statistics enhances evidence of functional enrichment. Bioinformatics 30, 2906–2914 (2014).
Lee, D., Bigdeli, T. B., Riley, B. P., Fanous, A. H. & Bacanu, S.-A. DIST: direct imputation of summary statistics for unmeasured SNPs. Bioinformatics 29, 2925–2927 (2013).
Kanai, M. et al. Meta-analysis fine-mapping is often miscalibrated at single-variant resolution. Cell Genom. 2, 100210 (2022).
Loh, P.-R., Kichaev, G., Gazal, S., Schoech, A. P. & Price, A. L. Mixed-model association for biobank-scale datasets. Nat. Genet. 50, 906–908 (2018).
Zhou, W. et al. Global Biobank Meta-analysis Initiative: powering genetic discovery across human disease. Cell Genom. 2, 100192 (2022).
Scutari, M., Mackay, I. & Balding, D. Using genetic distance to infer the accuracy of genomic prediction. PLoS Genet 12, e1006288 (2016).
Majara, L. et al. Low and differential polygenic score generalizability among African populations due largely to genetic diversity. HGG Adv. 4, 100184 (2021).
Browning, B. L. & Browning, S. R. Genotype imputation with millions of reference samples. Am. J. Hum. Genet. 98, 116–126 (2016).
Loh, P.-R. et al. Reference-based phasing using the Haplotype Reference Consortium panel. Nat. Genet. 48, 1443–1448 (2016).
Ralph, P., Thornton, K. & Kelleher, J. Efficiently summarizing relationships in large samples: a general duality between statistics of genealogies and genomes. Genetics 215, 779–797 (2020).
Zhang, B. C., Biddanda, A., Gunnarsson, Á. F., Cooper, F. & Palamara, P. F. Biobank-scale inference of ancestral recombination graphs enables genealogical analysis of complex traits. Nat. Genet. 55, 768–776 (2023).
Minichiello, M. J. & Durbin, R. Mapping trait loci by use of inferred ancestral recombination graphs. Am. J. Hum. Genet. 79, 910–922 (2006).
Link, V. et al. Tree-based QTL mapping with expected local genetic relatedness matrices. Preprint at bioRxiv https://doi.org/10.1101/2023.04.07.536093 (2023).
Ruan, Y. et al. Improving polygenic prediction in ancestrally diverse populations. Nat. Genet. 54, 573–580 (2022).
Privé, F., Arbel, J. & Vilhjálmsson, B. J. LDpred2: better, faster, stronger. Bioinformatics 36, 5424–5431 (2020).
Gazal, S. et al. Linkage disequilibrium–dependent architecture of human complex traits shows action of negative selection. Nat. Genet. 49, 1421–1427 (2017).
Orliac, E. J. et al. Improving GWAS discovery and genomic prediction accuracy in biobank data. Proc. Natl Acad. Sci. USA 119, e2121279119 (2022).
Lewis, A. C. F. et al. Getting genetic ancestry right for science and society. Science 376, 250–252 (2022).
Speed, D., Hemani, G., Johnson, M. R. & Balding, D. J. Improved heritability estimation from genome-wide SNPs. Am. J. Hum. Genet. 91, 1011–1021 (2012).
Yang, J., Zaitlen, N. A., Goddard, M. E., Visscher, P. M. & Price, A. L. Advantages and pitfalls in the application of mixed-model association methods. Nat. Genet. 46, 100–106 (2014).
Yengo, L. et al. Imprint of assortative mating on the human genome. Nat. Hum. Behav. 2, 948–954 (2018).
Hunt, S. E. et al. Ensembl variation resources. Database 2018, bay119 (2018).
Salehi Nowbandegani, P. et al. Data for extremely sparse models of linkage disequilibrium in ancestrally diverse association studies. Zenodo https://doi.org/10.5281/zenodo.8157131 (2023).
Wohns, A. W., O’Connor, L. J. & Salehi Nowbandegani, P. Software for linkage disequilibrium graphical models. Zenodo https://doi.org/10.5281/zenodo.8161389 (2023).
Acknowledgements
We thank D. J. Weiner, A. Nadig, A. L. Price, R. Walters, H. Finucane, X. Lin, H. Li, B. Lehmann, P. Ralph, G. Gorjanc, J. Kelleher and R. Mazumder for helpful discussions. We also thank G. Genovese for his implementation of BLUPx-ldgm in bcftools.
Author information
Authors and Affiliations
Contributions
A.W.W., P.S.N. and L.J.O. developed the methods. E.S.L., B.M.N. and A.B. suggested analyses. A.W.W., P.S.N., J.L.B. and L.J.O. performed the experiments. A.W.W., P.S.N., J.L.B., B.M.N. and L.J.O. wrote the paper. L.J.O. supervised the research.
Corresponding authors
Ethics declarations
Competing interests
B.M.N. is a member of the scientific advisory board at Deep Genomics and Neumora. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Genetics thanks the anonymous reviewers for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 LD between adjacent pairs of LD blocks.
For all blocks of chromosomes 1 to 22, we evaluated the mean r2 between every pair of SNPs (‘LDsum’) in consecutive LD blocks (n = 1,360 pairs), within each ancestry group. The expected mean r2 is around 1 × 10−3, that is 1/2n. The lower whisker, lower hinge, center, upper hinge and upper whisker correspond to (lower hinge − 1.5× interquartile range (IQR)) and the 25th percentile, median, 75th percentile, and (upper hinge + 1.5× IQR), respectively.
Extended Data Fig. 2 Within-sample accuracy of LDGM precision matrices.
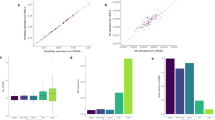
a-c, Boxplots showing the error of the LDGM precision matrix. d-f, Boxplots showing the error of the identity matrix. Three different error metrics were used. Boxplots indicate the median, quartiles and range for the 20 LD blocks on chromosome 22, for each 1000 Genomes ancestry group. a and d show the mean-squared error (see Methods). b and e show the mean-squared error after restricting to SNP pairs with a correlation of r2 > 0.01. c and f show the alternative mean-squared error, defined as m−2 Tr((I−PR)(I−RP)). This measures the difference between PR, the product of the LD correlation matrix and the LDGM precision matrix, and the identity matrix (see Supplementary Note, section 4). Compared with the MSE, the alternative MSE is less sensitive to large eigenvalues of R, probably explaining why it is not elevated for AMR. For the identity matrix, the alternative MSE and the MSE are identical. In all plots, the lower whisker, lower hinge, center, upper hinge and upper whisker correspond to (lower hinge − 1.5× interquartile range (IQR)) and the 25th percentile, median, 75th percentile, and (upper hinge + 1.5× IQR), respectively.
Extended Data Fig. 3 Comparison of LDGM precision matrices with Wen-Stephens shrinkage estimator.
The comparison was performed in EUR, on chromosome 22 only (n = 20 LD blocks). To vary the amount of shrinkage, we changed the sample size parameter in the Wen-Stephens estimator (actual sample size: 1,006). a, Mean-squared error between the Wen-Stephens estimator and the LDGM precision matrix inverse. Dotted line denotes the median MSE between the LD sample correlation matrix and the LDGM precision matrix inverse. b, Mean-squared error between the Wen-Stephens estimator and the sample correlation matrix. Values are larger than the corresponding numbers in a for sample size parameters up to 40, and smaller for sample size parameters of 201 or higher. c, Number of nonzero entries per SNP in the Wen-Stephens estimator. Correlations with absolute value less than 1 × 10−8 are set to zero (consistent with the original paper), resulting in slightly increased sparsity for small values of the sample size parameter. At larger parameter values, no SNP pairs are below the threshold within LD blocks, but this approach can still be used to produce a sparse, banded diagonal matrix when it is not desired to use discrete blocks. Somewhat more sparsity can be achieved by relaxing the 1 × 10−8 threshold, but not without causing increased error. In all plots, the lower whisker, lower hinge, center, upper hinge and upper whisker correspond to (lower hinge − 1.5× interquartile range (IQR)) and the 25th percentile, median, 75th percentile, and (upper hinge + 1.5× IQR), respectively.
Extended Data Fig. 4 Comparison of LDGM precision matrices vs. rank-k approximations for the local LD matrix.
The comparison was performed in EUR, on chromosome 22 only (20 LD blocks), and we considered different values of k. a, To quantify the accuracy of the rank-k approximation to the LD correlation matrix, we computed its MSE at different values of k. k = 10 corresponds most closely to the density of the LDGM precision matrix, which is a symmetric matrix with 20 nonzero entries per SNP (10 per SNP in its upper triangle). The MSE at k = 10 was about 3 times higher than that of the LDGM precision matrix; it was most similar at k = 50. MSE is always zero when k is greater than or equal to the sample size (that is, when k = 1,006). b, To quantify the accuracy of the rank-k approximation to the LD precision matrix, we computed the alternative MSE ratio, which quantifies whether the approximate precision matrix multiplied by the correlation matrix is close to the identity (see Extended Data Fig. 2 and Supplementary Note). By this metric, the LDGM performs much better than a rank-k approximation even at k = 500. c, We calculated the MSE between the rank-k approximation and the inverse of the LDGM precision matrix. This was never significantly smaller than the MSE of the LDGM precision matrix inverse with the sample correlation matrix. In all plots, the lower whisker, lower hinge, center, upper hinge and upper whisker correspond to (lower hinge − 1.5× interquartile range (IQR)) and the 25th percentile, median, 75th percentile, and (upper hinge + 1.5× IQR), respectively.
Extended Data Fig. 5 Performance of LDGM precision matrices in cross-validation.
For each LD block on chromosome 22 (n = 20 LD blocks), we randomly split the 1000 Genomes EUR haploid samples into two subsets of equal size. We computed an LDGM precision matrix from one of the two subsets (the LDGM was constructed from all samples in 1000 Genomes). We computed the MSE for three comparisons: the precision matrix vs. the correlation matrix from the same sample; the precision matrix vs. the correlation matrix from the opposite sample; and the correlation matrix from one sample vs. the correlation matrix from the opposite sample. The lower whisker, lower hinge, center, upper hinge and upper whisker correspond to (lower hinge − 1.5× interquartile range (IQR)) and the 25th percentile, median, 75th percentile, and (upper hinge + 1.5× IQR), respectively.
Extended Data Fig. 6 Accuracy and sparsity of precision matrices constructed from naive LDGMs.
a,b, In 1000 Genomes EUR data from chromosome 22, we compare our inferred LDGM (derived from tree sequences) with a banded-diagonal LDGM, an r2 threshold LDGM, and a banded-diagonal LDGM with a large band for 20 LD blocks. a, MSE versus control LDGMs. b, Density versus control LDGMS. For the first banded LDGM, the band size was chosen to match the number of edges with path weight less than 4 in our tree-sequence based LDGM for each LD block (approximately 50 edges per SNP). For the r2-threshold LDGM, the threshold was chosen to produce the same number of edges. For the large banded-diagonal LDGM, we used a band size that corresponded to the number of edges with path weight less than 8 (approximately 300 edges per SNP). For each LDGM, we computed precision matrices with an L1 penalty of 0.1, and we calculated the mean squared error (a) and the number of edges per SNP in the precision matrix (b). In all plots, the lower whisker, lower hinge, center, upper hinge and upper whisker correspond to (lower hinge − 1.5× interquartile range (IQR)) and the 25th percentile, median, 75th percentile, and (upper hinge + 1.5× IQR), respectively.
Extended Data Fig. 7 Accuracy and density of LDGM precision matrices at different allele frequencies.
For the 20 EUR LDGM precision matrices on chromosome 22, we partitioned SNPs into three bins by their minor allele frequency in EUR. Each bin contained a similar number of SNPs. a, MSE across pairs of SNPs (i, j) where SNP i has the specified allele frequency (and SNP j may or may not). b, Average number of neighbors per SNP in each MAF bin (including edges with SNPs not in the bin). In all plots the lower whisker, lower hinge, center, upper hinge and upper whisker correspond to (lower hinge − 1.5× interquartile range (IQR)) and the 25th percentile, median, 75th percentile, and (upper hinge + 1.5× IQR), respectively.
Extended Data Fig. 8 Tradeoff between accuracy and sparsity with different parameter settings.
a-d, On the 20 LD blocks of chromosome 22 EUR data, we varied the path distance threshold (a, b) and the L1 penalty (c, d). Our default parameter settings are a distance threshold of 4 and an L1 penalty of 0.1. Precision matrix inference runtime also varies with parameter settings, with greater runtime for settings that produce higher density. In all plots, the lower whisker, lower hinge, center, upper hinge and upper whisker correspond to (lower hinge − 1.5× interquartile range (IQR)) and the 25th percentile, median, 75th percentile, and (upper hinge + 1.5× IQR), respectively.
Extended Data Fig. 9 MSE of CpGs compared to non CpG sites on chromosome 21 and 22.
The mutation rate at CpG sites is more than an order of magnitude greater than non-CpG sites, and thus have a higher rate of recurrent mutations (Wohns, 2022). To evaluate how disregarding recurrent mutation effects overall accuracy (see Methods), we examined the MSE of CpG sites vs. non-CpG sites on the 40 LD blocks of chromosomes 21 and 22 between the LDGM precision matrix and sample correlation matrices. The lower whisker, lower hinge, center, upper hinge and upper whisker correspond to (lower hinge − 1.5× interquartile range (IQR)) and the 25th percentile, median, 75th percentile, and (upper hinge + 1.5× IQR), respectively. For numerical results, see Supplementary Table 13.
Extended Data Fig. 10 Runtime for LDGM and precision matrix inference across chromosome 22.
The first five boxplots indicate the runtime for precision matrix inference for each ancestry group; the sixth indicates the runtime to derive the LDGM from the original tree sequence. For the LDGM inference step, we used 5 compute threads (1 for the precision matrix inference step). Runtime varies across LD blocks and ancestry groups due to variation in the number of SNPs. The lower whisker, lower hinge, center, upper hinge and upper whisker correspond to (lower hinge − 1.5× interquartile range (IQR)) and the 25th percentile, median, 75th percentile, and (upper hinge + 1.5× IQR), respectively. For numerical results, see Supplementary Table 4.
Supplementary information
Supplementary Information
Supplementary Note, Figs. 1–8 and Table captions.
Supplementary Table 1
Supplementary Tables 1–15.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Salehi Nowbandegani, P., Wohns, A.W., Ballard, J.L. et al. Extremely sparse models of linkage disequilibrium in ancestrally diverse association studies. Nat Genet 55, 1494–1502 (2023). https://doi.org/10.1038/s41588-023-01487-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-023-01487-8