Abstract
Several molecular and phenotypic algorithms exist that establish genotype–phenotype correlations, including facial recognition tools. However, no unified framework that investigates both facial data and other phenotypic data directly from individuals exists. We developed PhenoScore: an open-source, artificial intelligence-based phenomics framework, combining facial recognition technology with Human Phenotype Ontology data analysis to quantify phenotypic similarity. Here we show PhenoScore’s ability to recognize distinct phenotypic entities by establishing recognizable phenotypes for 37 of 40 investigated syndromes against clinical features observed in individuals with other neurodevelopmental disorders and show it is an improvement on existing approaches. PhenoScore provides predictions for individuals with variants of unknown significance and enables sophisticated genotype–phenotype studies by testing hypotheses on possible phenotypic (sub)groups. PhenoScore confirmed previously known phenotypic subgroups caused by variants in the same gene for SATB1, SETBP1 and DEAF1 and provides objective clinical evidence for two distinct ADNP-related phenotypes, already established functionally.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The used dataset in this study is not publicly available due to both IRB and General Data Protection Regulation (EU GDPR) restrictions because the data might be (partially) traceable. However, access to the data may be requested from the data availability committee by contacting the corresponding authors via e-mail with a research proposal, who will respond within 14 d.
Code availability
The code of PhenoScore version 1.0.0 created during this study is freely available at https://github.com/ldingemans/PhenoScore ref. 83, to enable anyone to apply PhenoScore to their own dataset. Included in PhenoScore are the following two examples: the data for the SATB1 subgroups (positive example) and random data (negative example).
References
Vissers, L. E. L. M. et al. A de novo paradigm for mental retardation. Nat. Genet. 42, 1109–1112 (2010).
de Ligt, J. et al. Diagnostic exome sequencing in persons with severe intellectual disability. N. Engl. J. Med. 367, 1921–1929 (2012).
Rauch, A. et al. Range of genetic mutations associated with severe non-syndromic sporadic intellectual disability: an exome sequencing study. Lancet 380, 1674–1682 (2012).
Gilissen, C. et al. Genome sequencing identifies major causes of severe intellectual disability. Nature 511, 344–347 (2014).
Richards, S. et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet. Med. 17, 405–424 (2015).
Beaumont, R. N. & Wright, C. F. Estimating diagnostic noise in panel-based genomic analysis. Genet. Med. 24, 2042–2050 (2022).
McGuire, A. L. et al. The road ahead in genetics and genomics. Nat. Rev. Genet. 21, 581–596 (2020).
Logsdon, G. A., Vollger, M. R. & Eichler, E. E. Long-read human genome sequencing and its applications. Nat. Rev. Genet. 21, 597–614 (2020).
100,000 Genomes Project Pilot Investigators. et al.100,000 genomes pilot on rare-disease diagnosis in health care—preliminary report. N. Engl. J. Med. 385, 1868–1880 (2021).
Neveling, K. et al. Next-generation cytogenetics: comprehensive assessment of 52 hematological malignancy genomes by optical genome mapping. Am. J. Hum. Genet. 108, 1423–1435 (2021).
Mantere, T. et al. Optical genome mapping enables constitutional chromosomal aberration detection. Am. J. Hum. Genet. 108, 1409–1422 (2021).
Schwarz, J. M., Rödelsperger, C., Schuelke, M. & Seelow, D. MutationTaster evaluates disease-causing potential of sequence alterations. Nat. Methods 7, 575–576 (2010).
Adzhubei, I. A. et al. A method and server for predicting damaging missense mutations. Nat. Methods 7, 248–249 (2010).
Kumar, P., Henikoff, S. & Ng, P. C. Predicting the effects of coding non-synonymous variants on protein function using the SIFT algorithm. Nat. Protoc. 4, 1073–1081 (2009).
Kircher, M. et al. A general framework for estimating the relative pathogenicity of human genetic variants. Nat. Genet. 46, 310–315 (2014).
Robinson, P. N. et al. The Human Phenotype Ontology: a tool for annotating and analyzing human hereditary disease. Am. J. Hum. Genet. 83, 610–615 (2008).
Leite, A. J. D. C. et al. Diagnostic yield of patients with undiagnosed intellectual disability, global developmental delay and multiples congenital anomalies using karyotype, microarray analysis, whole exome sequencing from Central Brazil. PLoS ONE 17, e0266493 (2022).
Clift, K. et al. Patients’ views on variants of uncertain significance across indications. J. Community Genet. 11, 139–145 (2020).
Makhnoon, S., Garrett, L. T., Burke, W., Bowen, D. J. & Shirts, B. H. Experiences of patients seeking to participate in variant of uncertain significance reclassification research. J. Community Genet. 10, 189–196 (2019).
Van Dijk, S. et al. Clinical characteristics affect the impact of an uninformative DNA test result: the course of worry and distress experienced by women who apply for genetic testing for breast cancer. J. Clin. Oncol. 24, 3672–3677 (2006).
Murray, M. L., Cerrato, F., Bennett, R. L. & Jarvik, G. P. Follow-up of carriers of BRCA1 and BRCA2 variants of unknown significance: variant reclassification and surgical decisions. Genet. Med. 13, 998–1005 (2011).
Hamburg, M. A. & Collins, F. S. The path to personalized medicine. N. Engl. J. Med. 363, 301–304 (2010).
Ashley, E. A. Towards precision medicine. Nat. Rev. Genet. 17, 507–522 (2016).
Brittain, H. K., Scott, R. & Thomas, E. The rise of the genome and personalised medicine. Clin. Med. 17, 545–551 (2017).
Coudray, N. et al. Classification and mutation prediction from non-small cell lung cancer histopathology images using deep learning. Nat. Med. 24, 1559–1567 (2018).
Hosny, A., Parmar, C., Quackenbush, J., Schwartz, L. H. & Aerts, H. J. W. L. Artificial intelligence in radiology. Nat. Rev. Cancer 18, 500–510 (2018).
Killock, D. AI outperforms radiologists in mammographic screening. Nat. Rev. Clin. Oncol. 17, 134 (2020).
Lu, M. Y. et al. AI-based pathology predicts origins for cancers of unknown primary. Nature 594, 106–110 (2021).
Poplin, R. et al. A universal SNP and small-indel variant caller using deep neural networks. Nat. Biotechnol. 36, 983–987 (2018).
Sundaram, L. et al. Predicting the clinical impact of human mutation with deep neural networks. Nat. Genet. 50, 1161–1170 (2018).
Wick, R. R., Judd, L. M. & Holt, K. E. Performance of neural network basecalling tools for Oxford Nanopore sequencing. Genome Biol. 20, 129 (2019).
Köhler, S. et al. Clinical diagnostics in human genetics with semantic similarity searches in ontologies. Am. J. Hum. Genet. 85, 457–464 (2009).
Robinson, P. N. et al. Improved exome prioritization of disease genes through cross-species phenotype comparison. Genome Res. 24, 340–348 (2014).
Zemojtel, T. et al. Effective diagnosis of genetic disease by computational phenotype analysis of the disease-associated genome. Sci. Transl. Med. 6, 252ra123 (2014).
Smedley, D. & Robinson, P. N. Phenotype-driven strategies for exome prioritization of human Mendelian disease genes. Genome Med. 7, 81 (2015).
Smedley, D. et al. Next-generation diagnostics and disease-gene discovery with the Exomiser. Nat. Protoc. 10, 2004–2015 (2015).
Hsieh, T.-C. et al. PEDIA: prioritization of exome data by image analysis. Genet. Med. 21, 2807–2814 (2019).
Robinson, P. N. et al. Interpretable clinical genomics with a likelihood ratio paradigm. Am. J. Hum. Genet. 107, 403–417 (2020).
Ferry, Q. et al. Diagnostically relevant facial gestalt information from ordinary photos. eLife 3, e02020 (2014).
Dudding-Byth, T. et al. Computer face-matching technology using two-dimensional photographs accurately matches the facial gestalt of unrelated individuals with the same syndromic form of intellectual disability. BMC Biotechnol. 17, 90 (2017).
Van der Donk, R. et al. Next-generation phenotyping using computer vision algorithms in rare genomic neurodevelopmental disorders. Genet. Med. 21, 1719–1725 (2019).
Gurovich, Y. et al. Identifying facial phenotypes of genetic disorders using deep learning. Nat. Med. 25, 60–64 (2019).
Dingemans, A. J. M. et al. Quantitative facial phenotyping for Koolen-de Vries and 22q11.2 deletion syndrome. Eur. J. Hum. Genet. 29, 1418–1423 (2021).
Hsieh, T.-C. et al. GestaltMatcher facilitates rare disease matching using facial phenotype descriptors. Nat. Genet. 54, 349–357 (2022).
Claes, P. et al. Genome-wide mapping of global-to-local genetic effects on human facial shape. Nat. Genet. 50, 414–423 (2018).
White, J. D. et al. Insights into the genetic architecture of the human face. Nat. Genet. 53, 45–53 (2021).
Naqvi, S. et al. Shared heritability of human face and brain shape. Nat. Genet. 53, 830–839 (2021).
Zhang, M. et al. Genetic variants underlying differences in facial morphology in East Asian and European populations. Nat. Genet. 54, 403–411 (2022).
Vulto-van Silfhout, A. T. et al. Clinical significance of de novo and inherited copy-number variation. Hum. Mutat. 34, 1679–1687 (2013).
Brier, G. W. Verification of forecasts expressed in terms of probability. Mon. Weather Rev. 78, 1–3 (1950).
Koolen, D. A. et al. Mutations in the chromatin modifier gene KANSL1 cause the 17q21.31 microdeletion syndrome. Nat. Genet. 44, 639–641 (2012).
Zollino, M. et al. Mutations in KANSL1 cause the 17q21.31 microdeletion syndrome phenotype. Nat. Genet. 44, 636–638 (2012).
Koolen, D. A. et al. The Koolen-de Vries syndrome: a phenotypic comparison of patients with a 17q21.31 microdeletion versus a KANSL1 sequence variant. Eur. J. Hum. Genet. 24, 652–659 (2016).
Köhler, S. et al. Expansion of the Human Phenotype Ontology (HPO) knowledge base and resources. Nucleic Acids Res. 47, D1018–D1027 (2019).
den Hoed, J. et al. Mutation-specific pathophysiological mechanisms define different neurodevelopmental disorders associated with SATB1 dysfunction. Am. J. Hum. Genet. 108, 346–356 (2021).
Nabais Sá, M. J. et al. De novo and biallelic DEAF1 variants cause a phenotypic spectrum. Genet. Med. 21, 2059–2069 (2019).
Hoischen, A. et al. De novo mutations of SETBP1 cause Schinzel-Giedion syndrome. Nat. Genet. 42, 483–485 (2010).
Filges, I. et al. Reduced expression by SETBP1 haploinsufficiency causes developmental and expressive language delay indicating a phenotype distinct from Schinzel-Giedion syndrome. J. Med. Genet. 48, 117–122 (2011).
Bend, E. G. et al. Gene domain-specific DNA methylation episignatures highlight distinct molecular entities of ADNP syndrome. Clin. Epigenetics 11, 64 (2019).
Breen, M. S. et al. Episignatures stratifying Helsmoortel-Van Der Aa syndrome Show modest correlation with phenotype. Am. J. Hum. Genet. 107, 555–563 (2020).
Jagadeesh, K. A. et al. Phrank measures phenotype sets similarity to greatly improve Mendelian diagnostic disease prioritization. Genet. Med. 21, 464–470 (2019).
Lyra Jr, P. C. M. et al. Integration of functional assay data results provides strong evidence for classification of hundreds of BRCA1 variants of uncertain significance. Genet. Med. 23, 306–315 (2021).
Frederiksen, J. H., Jensen, S. B., Tümer, Z. & Hansen, T. V. O. Classification of MSH6 variants of uncertain significance using functional assays. Int. J. Mol. Sci. 22, 8627 (2021).
Caswell, R. C., Gunning, A. C., Owens, M. M., Ellard, S. & Wright, C. F. Assessing the clinical utility of protein structural analysis in genomic variant classification: experiences from a diagnostic laboratory. Genome Med. 14, 77 (2022).
Dingemans, A. J. M. et al. Human disease genes website series: an international, open and dynamic library for up-to-date clinical information. Am. J. Med. Genet. A 185, 1039–1046 (2021).
McKusick, V. A. Mendelian inheritance in man and its online version, OMIM. Am. J. Hum. Genet. 80, 588–604 (2007).
Firth, H. V. et al. DECIPHER: database of chromosomal imbalance and phenotype in humans using ensembl resources. Am. J. Hum. Genet. 84, 524–533 (2009).
Adam, M. P. et al. GeneReviews (Univ. Washington, 2010).
Helsmoortel, C. et al. A SWI/SNF-related autism syndrome caused by de novo mutations in ADNP. Nat. Genet. 46, 380–384 (2014).
Côté, R. A. & Robboy, S. Progress in medical information management. Systematized nomenclature of medicine (SNOMED). JAMA 243, 756–762 (1980).
Karras, T., Laine, S. & Aila, T. A style-based generator architecture for generative adversarial networks. IEEE Trans. Pattern Anal. Mach. Intell. 43, 4217–4228 (2021).
Manders, P., Lutomski, J. E., Smit, C., Swinkels, D. W. & Zielhuis, G. A. The Radboud biobank: a central facility for disease-based biobanks to optimise use and distribution of biomaterial for scientific research in the Radboud university medical center, Nijmegen. Open J. Bioresour. 5, 2 (2018).
Parkhi, O. M., Vedaldi, A. & Zisserman, A. Deep face recognition. Proceedings of the British Machine Vision Conference (eds Xianghua X. et al.) 41.1–41.12 (BMVA Press, 2015).
Cao, Q. Shen, L., Xie, W. Parkhi, O. M. & Zisserman, A. VGGFace2: a dataset for recognising faces across pose and age. Proceedings of 13th IEEE International Conference on Automatic Face & Gesture Recognition (F&G) pp. 67–74 (IEEE, 2018).
Dingemans, A. J. M., de Vries, B. B. A., Vissers, L. E. L., van Gerven, M. A. J. & Hinne, M. Comparing facial feature extraction methods in the diagnosis of rare genetic syndromes. Preprint at medRxiv https://doi.org/10.1101/2022.08.26.22279217 (2022).
Resnik, P. Semantic similarity in a taxonomy: an information-based measure and its application to problems of ambiguity in natural language. J. Artif. Intell. Res. 11, 95–130 (1999).
Pesquita, C. et al. Metrics for GO based protein semantic similarity: a systematic evaluation. BMC Bioinformatics 9, S4 (2008).
Arvai, K., Gainullin, V. & Borroto, C. GeneDx/phenopy. Zenodo https://doi.org/10.5281/zenodo.4587231 (2019).
Ribeiro, M. T., Singh, S. & Guestrin, C. ‘Why should I trust you?’ Explaining the predictions of any classifier. Proceedings of the 22nd ACM SIGKDD International Conference on Knowledge Discovery and DATA MINIng 1135–1144 (Association for Computing Machinery, 2016).
Ras, G., Xie, N., van Gerven, M. & Doran, D. Explainable deep learning: a field guide for the uninitiated. J. Artif. Intell. Res. 73, 329–396 (2022).
Köhler, S. et al. The human phenotype ontology in 2017. Nucleic Acids Res. 45, D865–D876 (2017).
Yuan, X. et al. Evaluation of phenotype-driven gene prioritization methods for Mendelian diseases. Brief. Bioinform. 23, bbac019 (2022).
Dingemans, L. ldingemans/PhenoScore: v1.0.0. Zenodo https://doi.org/10.5281/zenodo.7892317 (2023).
Acknowledgements
We are grateful to all families and clinicians who agreed to participate and provide clinical and genotypic information. R.F.K. acknowledges financial support from the Research Fund of the University of Antwerp (Methusalem-OEC grant GENOMED). The work of G.J.L. is supported by New York State Office for People with Developmental Disabilities (OPWDD) and NIH NIGMS R35-GM-133408. E.E.P. is supported by a National Health and Medical Research Council Investigator Grant (award 2021/GNT2008166). Furthermore, we are grateful to the Dutch Organization for Health Research and Development—ZON-MW grants 912-12-109 (to B.B.A.d.V. and L.E.L.M.V.), Donders Junior researcher grant 2019 (to B.B.A.d.V. and L.E.L.M.V.) and Aspasia grant 015.014.066 (to L.E.L.M.V.). The aims of this study contribute to the Solve-RD project (to L.E.L.M.V.), which has received funding from the European Unions Horizon 2020 research and innovation program under grant agreement 779257. The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
A.J.M.D., M.H., L.E.L.M.V., B.B.A.d.V. and M.A.J.v.G. designed the study. A.J.M.D., K.M.G.T., L.G., J.v.R., N.d.L., J.S.H., R.P., I.J.D., E.d.B., J.d.H., J.v.d.S., S.J., B.W.v.B., N.J., E.E.P., P.M.C., A.T.V.v.S., T.K., D.A.K., F.K., H.V.E., G.J.L., F.S.A., A.R., R.M., D.B., P.J.v.d.S., G.S., L.E.L.M.V. and B.B.A.d.V. collected and curated the data. A.J.M.D. and M.H. performed the formal analyses. L.E.L.M.V. and B.B.A.d.V. acquired the funding. A.J.M.D. and M.H. completed the modeling and investigations. A.J.M.D. developed the software. A.J.M.D., M.H., L.E.L.M.V., B.B.A.d.V. and M.A.J.v.G. wrote the original draft. All authors reviewed and edited the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Genetics thanks Xinran Dong and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
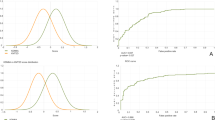
Extended Data Fig. 1 Benchmarking PhenoScore.
The predictive accuracies of LIRICAL, Phenomizer and PhenoScore [118-120] for every included genetic syndrome are displayed here, except for ACTL6A, since the associated phenotype has no OMIM number and therefore Phenomizer and LIRICAL do not include it in its predictions. For PhenoScore and LIRICAL, to calculate the accuracy, a cut-off value of 0.5 for the predictions was used, while for Phenomizer in this case, 0.05 was chosen. For almost every investigated syndrome, PhenoScore outperforms Phenomizer and LIRICAL.
Extended Data Fig. 2 AUC curves of PhenoScore per genetic syndrome.
The receiver operating characteristic curve of all 40 genetic syndromes included in this study.
Extended Data Fig. 3 UMAP plots of facial feature vectors.
The Uniform Manifold Approximation and Projection for Dimension Reduction (UMAP3) plot for the VGGFace2 vectors of all included genetic syndromes, and for the extra systematic confounder analysis for which the individuals with Koolen-de Vries syndrome seen at other centers were compared to individuals seen at our outpatient clinic. For all plots (except the KANSL1 internal/external plot), the feature vectors of all sampled controls during five iterations and the feature vectors of the included patients were provided as input to UMAP. The classes are not separable in this projected space, which provides evidence that the classification is not based on a systematic confounder.
Supplementary information
Supplementary Information
Supplementary Data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Dingemans, A.J.M., Hinne, M., Truijen, K.M.G. et al. PhenoScore quantifies phenotypic variation for rare genetic diseases by combining facial analysis with other clinical features using a machine-learning framework. Nat Genet 55, 1598–1607 (2023). https://doi.org/10.1038/s41588-023-01469-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-023-01469-w
This article is cited by
-
Clinical impact and in vitro characterization of ADNP variants in pediatric patients
Molecular Autism (2024)
-
ADNP dysregulates methylation and mitochondrial gene expression in the cerebellum of a Helsmoortel–Van der Aa syndrome autopsy case
Acta Neuropathologica Communications (2024)
-
Phenotypic similarity-based approach for variant prioritization for unsolved rare disease: a preliminary methodological report
European Journal of Human Genetics (2024)
-
Loss-of-function of activity-dependent neuroprotective protein (ADNP) by a splice-acceptor site mutation causes Helsmoortel–Van der Aa syndrome
European Journal of Human Genetics (2024)
-
Computer-aided diagnostic screen for Congenital Central Hypoventilation Syndrome with facial phenotype
Pediatric Research (2024)