Abstract
Human genetic syndromes are often challenging to diagnose clinically. Facial phenotype is a key diagnostic indicator for hundreds of genetic syndromes and computer-assisted facial phenotyping is a promising approach to assist diagnosis. Most previous approaches to automated face-based syndrome diagnosis have analyzed different datasets of either 2D images or surface mesh-based 3D facial representations, making direct comparisons of performance challenging. In this work, we developed a set of subject-matched 2D and 3D facial representations, which we then analyzed with the aim of comparing the performance of 2D and 3D image-based approaches to computer-assisted syndrome diagnosis. This work represents the most comprehensive subject-matched analyses to date on this topic. In our analyses of 1907 subject faces representing 43 different genetic syndromes, 3D surface-based syndrome classification models significantly outperformed 2D image-based models trained and evaluated on the same subject faces. These results suggest that the clinical adoption of 3D facial scanning technology and continued collection of syndromic 3D facial scan data may substantially improve face-based syndrome diagnosis.
Similar content being viewed by others
Introduction
Genetic syndromes represent an unusually difficult diagnostic challenge for even the most experienced clinicians, due to the large number, complexity, variability, and rarity of these disorders. Diagnoses are frequently delayed, are often initially incorrect, and patients often must proceed without even basic information regarding health and developmental outcomes, let alone tailored clinical care [1]. While advancements in gene-based technologies have greatly improved poor diagnostic rates, accurate testing is often unavailable, and complementary diagnostic modalities thus remain of great importance. Computer-assisted facial phenotyping is a complementary diagnostic modality that makes use of inexpensive, portable, and widely available facial imaging technologies, along with image processing and statistical methods. Dysmorphic (abnormal) facial features are associated with many human genetic syndromes [2] [3]. Several face-based approaches to genetic syndrome diagnosis have been developed to provide precision diagnostic assistance in a clinical context [4, 5].
Previous work
A variety of approaches for face-based syndrome diagnosis have been proposed using different types of facial representation as input [4, 6,7,8,9]. The most common forms of facial representation used for this purpose are 2D color images, 2.5D depth images, and 3D surface scans. To our knowledge, only one previous study has performed subject matched comparisons of 2D and 3D facial images for syndrome classification [10]. However, this study was limited to a single syndrome class (22q11.2 deletion) and the 2D images were colorless renderings of 3D surface scans that are not an accurate proxy for real 2D color or grayscale images.
Outside the application of genetic syndrome diagnosis, several studies have compared 2D and 3D facial representations for different purposes. Anas et al. [11] found that using 2D images and 3D surface scans for measuring facial morphology produced significantly different results, concluding that 3D imaging is a better approach to quantify facial morphologic phenotypes. Likewise, Zogheib et al. [12] found that 3D facial scanning produced more reliable facial measurements that were closer to the clinical standard than did 2D photographs. For the task of facial action unit detection, Savran et al. [13] found that, in general, 3D outperformed 2D facial representations. For the task of facial recognition, Chang et al. [14] found that 3D and 2D facial representations performed similarly. Nevertheless, many facial recognition researchers have turned their focus to 3D to overcome the recognized and inherent limitations of 2D photography [15]. Thus, many modern proprietary face-based bio-metric security applications (such as Face ID developed by Apple Inc.) now rely on 3D information [16].
2D color images
Most syndrome recognition models developed thus far rely on 2D color images. This is largely due to the relative ease of collecting 2D facial images using widely available digital camera technologies. As a result, databases of syndromic 2D images tend to be larger than those of syndromic 3D scans. Gurovich et al. [7] and Matthews et al. [4] provide detailed surveys of 2D image-based approaches and the different facial representations that they use. Anatomical landmarks as well as local geometry and texture features captured around each landmark are popular types of facial representations derived from 2D images. State-of-the-art approaches use deep convolutional neural networks (CNN) that take 2D color images as direct inputs [7, 17].
Despite the ease of acquisition, 2D facial images have some notable disadvantages for face-based syndrome classification. 2D facial images are projections of intrinsically 3D structures and, as such, may discard diagnostically relevant 3D morphologic information. 2D facial images are also highly sensitive to variations in illumination and pose. Even when controlling for illumination and pose, varying the distance from which a frontal 2D image is captured results in perspective distortions of the resultant 2D image (see Fig. 1) that could affect a diagnostic model. Finally, 2D images generally do not contain information about overall facial size, which may be an important diagnostic indicator for some syndromes. Therefore, additional calibration procedures or measurements must be used to capture true facial size information within or alongside 2D facial images.
Depth images
In between 2D color photography and 3D surface scanning is depth imaging (sometimes called 2.5D imaging). Depth images are 2D grid-like representations, like 2D color images. However, each pixel in a color image corresponds to the color and intensity of light reflected by the subject, whereas each pixel in a depth image corresponds to the distance of the subject from the imaging device. Depth images are usually obtained using infrared or time-of-flight depth sensors and/or multiple cameras that can compute depth images using stereoscopic methods. Many modern smartphones have embedded hardware that supports depth imaging [16]. Depth images have been explored for face-based syndrome classification but are less common than 2D color images and 3D surface scans. Previous studies have used principal components analysis (PCA) and a naive Bayes classifier to analyze syndromic facial depth images [10]. Many of the limitations of 2D color images also apply to depth images (such as sensitivity to changes in subject pose). Nevertheless, unlike 2D color images, depth images are generally robust to differences in subject illumination.
3D surface representations
3D surface scanning is the most sophisticated approach for facial imaging. The typical output format of 3D surface scanning systems is a discrete 3D surface mesh consisting of 3D vertices connected by polygons, such as triangles. Surface color is sometimes captured along with a polygonal surface mesh, using per-vertex color information, or UV mapped texture images. Because 3D facial surface meshes are a loosely structured data type, it is common practice to identify facial landmarks or dense point correspondences to a reference surface mesh facilitating model training and inference (see “Facial Representations” for details).
3D surface scanning is rapidly becoming more widely accessible and user friendly, although not yet as widespread and easy to use as 2D color photography. Specialized structured light imaging systems, such as those sold by 3dMDFootnote 1 produce scans of the highest quality and accuracy but are not easily portable. Facial surface scans can also be acquired using some of the newest smartphones or by less expensive handheld devices. In general, 3D scan-based syndrome diagnosis approaches are less common than 2D color image-based approaches, but several advanced models have been developed that use 3D representations [8, 9].
In theory, a 3D surface representation should result in better syndrome classification performance than comparable 2D images. 3D surface representations can intrinsically capture 3D human facial structures without discarding information through a 2D projection. 3D surface scans are robust to variation in subject illumination and pose. Finally, 3D scanning systems are typically calibrated to accurately capture size information.
Contributions
Although intuition suggests that 3D surface representations should be superior to 2D representations for face-based syndrome diagnosis, there is a lack of quantitative evidence to support this claim. It is also unclear whether the performance benefits from 3D imaging (if any) are sufficient to justify the increased effort of scan acquisition. This gap largely exists because previously published 2D and 3D face-based syndrome classification approaches have been trained and evaluated on different datasets with different numbers of subjects, different genetic syndrome classes and composition of the overall data set, and different demographic distributions within the data. Thus, it is difficult to draw any robust conclusions about facial representations by comparing previously published results. In this work, we describe the creation of parallel 2D and 3D facial representations from matched subjects in a large and diverse syndromic population. We also report the results from subject-matched analyses of four different 2D and 3D facial representations.
In summary, our experiments are the first to directly compare 2D and 3D face-based syndrome diagnosis models using identical patient faces for training and evaluation. This is important for two reasons. The first reason is that the use of different evaluation data can affect the metrics used to evaluate diagnostic models. The second is that the performance of diagnostic models is influenced by the composition, quality, and amount of data used to train the model. Thus, results from models trained on different patient data may reflect differences in the amount, quality, and composition of the training data, rather than the differences between 2D and 3D imaging modalities. Our experiments isolate the effects of using different facial representations on diagnostic model accuracy, providing empirical justification for continued research, data-collection, and model development using 3D facial imaging modalities.
Materials and methods
Data description
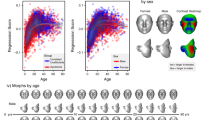
1907 3D surface scans from subjects with 43 different genetic syndromes were used in our experiments. Each syndrome was represented by at least 20 subjects. The 3D scans were acquired using a 3DMD facial imaging system from patients across the United States and Canada and are available through application to the Face Base consortiumFootnote 2. All scans were in the format of polygonal meshes with additional per-vertex color information. The demographic distribution of the subjects is shown in Fig. 2 and Table 1.
Facial representations
The following subsections describe the creation of the different 3D and 2D facial representations for each subject used in our comparative analyses. All representations were derived using a single raw 3D surface scan from each subject so that any variability in the imaging conditions (e.g., facial expression and illumination) is constant across the different representations. Figure 3 shows the image-like facial representations used in this study for an example subject.
3D surfaces
Just as 2D cameras can produce images with different numbers of pixels, raw 3D surface scans may have different numbers of vertices that are connected in different ways by different polygonal faces. Therefore, prior to model training and inference, raw 3D surface scans are typically processed to produce standardized 3D surface representations with a uniform number of vertices and a common mesh topology. To achieve this, dense vertex correspondences are identified between a reference facial mesh and each raw facial scan.
To facilitate the processing of raw 3D surface scans, eight anatomical landmarks were first identified on each scan using the automatic approach described in [18]. Next, dense vertex correspondences were estimated between a reference facial mesh and each raw facial scan using the non-rigid iterative closest point algorithm [19] guided by the anatomical landmarks. The reference mesh used in this work contains 26,649 vertices and the facial region that is covered by the reference mesh is shown in Fig. 3. Next, the standardized vertex configurations extracted from each raw facial scan were rigidly aligned to one another employing Procrustes alignment. Thus, diagnostically irrelevant information related to facial position and orientation was removed from the data while information about facial size and shape was retained. Finally, the aligned vertex coordinate information for each subject was flattened into a vector of length 3 × 26649 to be used for model training and inference.
2D color images
2D color images were created from 3D surface scans by rendering each scan from a frontal position. First, the raw scans were rigidly aligned to one another as described in the previous subsection. Next, a virtual camera was positioned to capture each scan from the same frontal position. Each scan was then rendered using per-vertex color information to produce realistic surface shading. To ensure that both 2D and 3D representations capture the same facial region as one another, the standardized 3D representations described in the previous subsection were used to mask regions of each 2D image that were not also captured by the 3D surface representations. Finally, the images were cropped to a bounding square that captures the full facial region and resized to a resolution of 128 × 128 pixels (slightly larger than the resolution used by [7]).
2D colorless images
2D colorless images were created from 3D surface scans using a slight modification of the rendering process for 2D color images. For colorless images, a uniform grey color was applied to the surface mesh instead of the available per-vertex color information. Although this colorless 2D representation is not comparable to any 2D imaging technique (including black and white photography), we included it due to its use in previous studies [10], and to investigate the effect of surface color information on syndrome classification performance.
Depth images
Depth images were created from 3D surface scans using another slight modification of the rendering process for 2D color images. For depth images, pixel values were set using the Z-buffer instead of using surface shading information. Z-buffer values represent the distance of rendered objects from a particular camera perspective.
Experiments
The primary aim of this study was to compare syndrome classification performance across different 3D and 2D representations using the same subject faces for training and evaluation. Therefore, we created five cross-validation folds by randomly splitting subjects with 70% of subject faces as training samples and 30% of subject faces as testing samples. Random sampling was stratified by syndrome class to ensure that an adequate number of subjects from each syndrome was included in each test set. The same subject splits were used for all experiments.
2D classifier model
We used the same convolutional neural network (CNN) model architecture for all 2D representations. The CNN (described in Table 2) was designed to emulate the model proposed by [7]. Unlike [7], we do not employ any 2D data augmentation or patch-based ensemble strategies to make the 2D classification experiments as comparable as possible to the 3D classification experiments. All CNN models were trained for 100 epochs using a batch size of 128 and an Adam optimizer with learning rate 10−3.
In addition to training 2D CNNs from scratch, we explored the use of pre-trained facial recognition CNN models provided by the DeepFace python library [20]. In these experiments, a CNN pre-trained on non-syndromic 2D facial images was used to extract embedding vectors for each syndromic facial image. The embedding vectors were then used to train and evaluate a MLP classification model with the same structure as the 3D classifier model described below. We found that the pre-trained ArcFace [21] model performed best for our application.
3D classifier model
For the 3D classification experiment, we used principal component analysis (PCA) to reduce the dimensionality of the 3D surface data from 3 × 26649 down to 100 following [9] to avoid overfitting. A multi-layer perceptron classifier model (MLP) was then trained on the dimensionality reduced data. The MLP architecture contains a single hidden layer of size 100 with ReLU activation. All MLP models were trained for 100 epochs using a using a batch size of 128 and an Adam optimizer with learning rate 10−3.
Results and discussion
Classification results
Table 3 shows the mean top-1 and top-3 accuracy scores across all syndrome classes for the cross-validated syndrome classification experiments. The 3D surface-based model produced the best performance of all experiments (40.7% mean top-1 sensitivity) by a margin of over 6%. Among the 2D representations, color images produced the best results (34.2% mean top-1 sensitivity), followed by colorless images (26.3% mean top-1 sensitivity). Depth images produced the worst classification results of all models evaluated (24.8% mean top-1 sensitivity).
We also investigated performance differences between facial representations within specific genetic syndrome classes (Table 1). Overall, the performance advantage of 3D representations is relatively consistent across syndrome classes although some differences were observed. Due to the much smaller sample sizes of individual syndrome classes, performance estimates are less precise and comparisons on a per-syndrome basis are more difficult.
Discussion
Overall, the experimental results support the proposition that 3D surface representations are superior to 2D representations for face-based syndrome diagnosis given equivalent subject faces for model training and evaluation. Compared to the best performing 2D representation (color images), the mean top-1 sensitivity of a diagnostic model trained using 3D surface representations was roughly 6% higher. All experiments were performed using the exact same subject faces for training and evaluation to control for performance variability from using different subjects, with different demographic and syndrome class distributions. By using a single 3D surface scan to derive all 3D and 2D representations, we were also able to ensure equivalent facial expressions and facial regions across the different facial representations.
Although our color renderings were created as a proxy for 2D color photographs, there are some relevant differences between our renderings and real color photographs. The backgrounds of our rendered images are uniformly black, unlike real images with complex and diverse backgrounds. Furthermore, our 3D alignment procedure removes pose variation from the image data and all images are captured using the same fixed camera distance from each subject face. As shown in Fig. 1, camera distance can have a noticeable effect on the appearance of frontal 2D photographs. Our rendering pipeline also uses the same virtual lighting conditions for each subject, although lighting conditions during the process of 3D scanning may still affect the per-vertex color information of each raw 3D scan. Typically, real world 2D facial photographs are captured under less controlled circumstances compared to the rendering process used for our simulated 2D color image representations. More precisely, real 2D images are often taken from different perspectives, with different lighting conditions and backgrounds. For our experiments, we chose not to simulate real life variation in imaging conditions so that our results simulate 2D photography in a controlled environment. Thus, it may be expected that real-world results for 2D images would lead to worse results than described here.
Although the 3D model outperformed 2D models for most syndromes (Table 1), some syndromes (Phelan McDermid, Mucopolysaccharidosis, Kabuki, Cohen, and Crouzon) showed lower performance for the 3D approach. This suggests that the methods used to measure 3D facial phenotype could still be improved. One potential source of noise in the 3D data is the process of estimating vertex correspondences between raw 3D scans and a reference facial mesh. Using improved anatomical landmark estimation algorithms to guide dense vertex correspondence estimation could lead to more robust 3D representations. Another potential direction to improve 3D models would be to use 3D surface texture as predictor of genetic syndrome diagnosis in addition to 3D geometry. 3D surface mesh textures can be encoded as per-vertex colors, or as 2D images that are mapped onto a surface mesh by assigning 2D image coordinates to each 3D vertex. Either encoding could be passed as an additional input to a syndrome classification model. The proposition that facial complexion carries diagnostically relevant information is also supported by a comparison of the 2D model results. In our experiments, 2D color representations (34.2% mean top-1 sensitivity) outperformed 2D colorless representations (26.3% mean top-1 sensitivity) that had surface color information removed. One interpretation of this result is that facial complexion carries diagnostically relevant information for some genetic syndromes. Some syndromes are known to be associated with unique features that manifest in the complexion of the skin rather than in the geometry of the facial structure. For example, neurofibromatosis is characterized by café au lait spots and Prader-Willi syndrome is characterized by hypopigmentation. These features can only be detected by imaging modalities that capture skin complexion information such as color-based 2D classification models or 3D models that include texture information. Another possibility is that our dataset contains spurious correlations between facial complexion or illumination at the time of 3D scanning. While all model development efforts should carefully consider the demographic distribution of their data, special care should be taken for data representations that include complexion information so that models are not ethnically biased. One additional advantage of 3D surface scanning is the ability to explicitly separate geometric information and information about facial complexion as they are acquired by scanning systems through different mechanisms. Although 3D scanning offers the possibility to isolate information about facial geometry, facial shape does vary across ethnic groups [22, 23] and 3D models can still be ethnically biased when trained on imbalanced data.
Our experiments also revealed that using a pre-trained CNN to extract facial embedding vectors (34.2% mean top-1 sensitivity) performed as well or better than training a CNN from scratch (32.6% mean top-1 sensitivity). However, even when additional non-syndromic facial images were used to pre-train a CNN, the 2D models did not achieve the performance level of the 3D model (40.7% mean top-1 sensitivity). Nevertheless, this result suggests that syndromic and non-syndromic faces are similar enough to make transfer-learning between non-syndromic and syndromic datasets viable. Furthermore, it suggests that images rendered from 3D surface scans are similar enough to real 2D facial images to make transfer-learning between 3D and 2D datasets viable. We believe that using 3D facial scans to investigate and improve 2D facial diagnosis models is also a very promising avenue for future research. High quality 3D scans can be used to generate 2D facial images of the same subject from various perspectives, and with different lighting conditions and backgrounds. These images could be used to evaluate how sensitive existing 2D models are to different imaging conditions. Furthermore, images of the same subject under different conditions could be used to train models that are invariant to differences in those conditions.
Conclusion
For a variety of reasons, it can be concluded that 3D scans are superior to 2D images for face-based genetic syndrome diagnosis. As demonstrated in this work, 3D surface scans can be converted into 2D images taken from any perspective using a configurable rendering pipeline. Furthermore, our subject-matched experiments revealed that 3D surface representations produce improved syndrome classification performance compared to 2D representations. We believe that investment in clinical 3D imaging systems and syndromic 3D data collection is likely to result in continued improvements to both 3D and 2D face-based syndrome diagnosis models.
Data availability
The 3D facial scan data used in this work is available through application to the Face Base consortium (www.facebase.org).
References
Thevenot J, Lopez M, Hadid A. A survey on computer vision for assistive medical diagnosis from faces. IEEE J Biomed Health Inform. 2018:22:1497–511.
Hennekam R, Allanson J, Krantz I. Gorlin’s Syndromes of the Head and Neck, Oxford University Press, 2010.
Hart T, Hart P. Genetic studies of craniofacial anomalies: clinical implications and applications. Orthod Craniofacial Res. 2009;12:212–20.
Matthews H, de Jong G, Maal T, Claes P. Static and motion facial analysis for craniofacial assessment and diagnosing diseases. Annu Rev Biomed Data Sci. 2022;5:19–42.
MacEachern SJ, Forkert ND. Machine learning for precision medicine. Genome. 2021;64:416–25.
Alsafar MCER, Werghi N, Al-Muhairi H, Alsafar H. Using facial images for the diagnosis of genetic syndromes: a survey. in International Conference on Communications, Signal Processing, and their Applications, 2015.
Gurovich Y, Hanani Y, Bar O, Nadav G, Fleischer N, Gelbman D, et al. Identifying facial phenotypes of genetic disorders using deep learning. Nat Med. 2019;25:60–4.
Hallgrímsson B, Aponte JD, Katz DC, Bannister JJ, Riccardi SL, Mahasuwan N, et al. Automated syndrome diagnosis by three-dimensional facial imaging. Genet Med. 2020;22:1–12.
Bannister J, Wilms M, Aponte D, Katz D, Klein OD, Bernier FP, et al. A deep invertible 3D facial shape model for interpretable genetic syndrome diagnosis. IEEE J Biomed Health Inform. 2022;26:3229–39.
Wilamowska K, Shapiro L, Heike C. Classification of 3D face shape in 22q11.2 deletion syndrome. in IEEE International Symposium on Biomedical Imaging: From Nano to Macro, 2009.
Anas I, Bamgbose B, Nuhu S. A comparison between 2D and 3D methods of quantifying facial morphology. Heliyon. 2019;5:e01880.
Zogheib T, Jacobs R, Bornstein M, Agbaje J, Anumendem D, Klazen Y, et al. Comparison of 3D scanning versus 2D photography for the identification of facial soft-tissue landmarks. Open Dent J. 2018;12:61–71.
Bilge AS, Sankur B, Taha M. Comparative evaluation of 3D vs. 2D modality for automatic detection of facial action units. Pattern Recognit. 2012;45:767–82.
Chang K, Bowyer K, Flynn P, Chen X. Multi-biometrics using facial appearance, shape and temperature. in Sixth IEEE International Conference on Automatic Face and Gesture Recognition, 2004.
Zhou S, Xiao S. 3D face recognition: a survey. Hum-centric Comput Inform Sci. 2018;8. https://hcis-journal.springeropen.com/articles/10.1186/s13673-018-0157-2.
“Face ID Security,” 2017. [Online]. Available: www.apple.com/business-docs/FaceID_Security_Guide.pdf.
Hsieh T-C, Aviram B-H, Moosa S, Ehmke N, Gripp KW, Pantel JT, et al. GestaltMatcher facilitates rare disease matching using facial phenotype descriptors. Nat Genet. 2022;54:349–57.
Bannister JJ, Crites SR, Aponte JD, Katz DC, Wilms M, Klein OD, et al. Fully automatic landmarking of syndromic 3D facial surface scans using 2D images. Sensors. 2020;20:3171.
Amberg B, Romdhani S, Vetter T. Optimal Step Nonrigid ICP Algorithms for Surface Registration. in IEEE Conference on Computer Vision and Pattern Recognition, 2007.
Serengil SI, Ozpinar A. “LightFace: a hybrid deep face recognition framework,” in 2020 Innovations in Intelligent Systems and Applications Conference (ASYU), 2020.
Deng J, Guo J, Xue N, Zafeiriou S. ArcFace: Additive Angular Margin Loss for Deep Face Recognition. in 2019 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), 2019.
Farkas LG, Katic MJ, Forrest CR. International anthropometric study of facial morphology in various ethnic groups/races. J Craniofacial Surg. 2005;16:615–46.
Berretti S, Del Bimbo A, Pala P. Distinguishing facial features for ethnicity-based 3D face recognition. ACM Trans Intell Syst Technol (TIST). 2012;3:1–20.
Funding
This research was funded by the National Institutes of Health (U01-DE024440), the Canada Research Chairs program, as well as by the River Fund at Calgary Foundation.
Author information
Authors and Affiliations
Contributions
(1) Research Project: A. Conception, B. Organization, C. Execution; (2) Statistical Analysis: A. Design, B. Execution, C. Review and Critique; (3) Manuscript: A. Writing of the First Draft, B. Review and Critique. JJB: 1A, 1B, 1C, 2A, 2B, 3A. DA: 1B, 1C, 2C, 3B. DK: 1B, 1C, 2C, 3B. ODK: 1A, 1B, 3B. FPJB: 1A, 1B, 3B. RAS: 1A, 1B, 3B. BH: 1A, 1B, 2A, 2C, 3B. NDF: 1A, 1B, 2A, 2C, 3B.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval
This study was conducted in accordance with the Declaration of Helsinki. Ethics approval was granted by the Conjoint Health Research Ethics Board (CHREB# REB14-0340_REN4) at the University of Calgary. Informed consent informed was obtained from all subjects. Consent for the publication of facial images was obtained for the example subject shown in Figs. 1 and 3.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Bannister, J.J., Wilms, M., Aponte, J.D. et al. Comparing 2D and 3D representations for face-based genetic syndrome diagnosis. Eur J Hum Genet 31, 1010–1016 (2023). https://doi.org/10.1038/s41431-023-01308-w
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41431-023-01308-w
This article is cited by
-
Why don’t we all use genomic testing?
European Journal of Human Genetics (2023)