Abstract
Transposable elements (TEs) are parasitic DNA sequences accounting for over half of the human genome. Tight control of the repression and activation states of TEs is critical for genome integrity, development, immunity and diseases, including cancer. However, precisely how this regulation is achieved remains unclear. Here we develop a targeted proteomic proximity labeling approach to capture TE-associated proteins in human embryonic stem cells (hESCs). We find that the RNA N6-methyladenosine (m6A) reader, YTHDC2, occupies genomic loci of the primate-specific TE, LTR7/HERV-H, specifically through its interaction with m6A-modified HERV-H RNAs. Unexpectedly, YTHDC2 recruits the DNA 5-methylcytosine (5mC)-demethylase, TET1, to remove 5mC from LTR7/HERV-H and prevent epigenetic silencing. Functionally, the YTHDC2/LTR7 axis inhibits neural differentiation of hESCs. Our results reveal both an underappreciated crosstalk between RNA m6A and DNA 5mC, the most abundant regulatory modifications of RNA and DNA in eukaryotes, and the fact that in hESCs this interplay controls TE activity and cell fate.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Sequencing data have been deposited to the NCBI Gene Expression Omnibus (GEO) (http://www.ncbi.nlm.nih.gov/geo) under accession number GSE210867. Publicly available RNA-seq data for WT and TET1 knockout H1 cells (GSM5183601, GSM5183602, GSM5183607, GSM5183608), methyl-capture sequencing (GSM5183585, GSM5183586, GSM5183589, GSM5183590), H3K4me1 (GSM409307), H3K4me3 (GSM409308) and MeRIP-seq (GSM1272365, GSM1272366, GSM1272367, GSM1272368) were obtained from the GEO. H3K27ac (ENCFF103PND) and H3K9me3 (ENCFF385ZBQ) were obtained from the ENCODE project (https://www.encodeproject.org/). Proteomics data are available in the MassIVE database (Massive.ucsd.edu) under accession MSV000091407. Additional experimental materials used in this study, including plasmids and engineered cell lines, are available upon request. Source data are provided with this paper.
Code availability
All the data were analyzed using published pipelines with parameters described in the Methods section. No custom codes were developed in this study.
References
Bourque, G. et al. Ten things you should know about transposable elements. Genome Biol. 19, 199 (2018).
Chuong, E. B., Elde, N. C. & Feschotte, C. Regulatory activities of transposable elements: from conflicts to benefits. Nat. Rev. Genet. 18, 71–86 (2017).
Senft, A. D. & Macfarlan, T. S. Transposable elements shape the evolution of mammalian development. Nat. Rev. Genet. https://doi.org/10.1038/s41576-021-00385-1 (2021).
Fueyo, R., Judd, J., Feschotte, C. & Wysocka, J. Roles of transposable elements in the regulation of mammalian transcription. Nat. Rev. Mol. Cell Biol. 23, 481–497 (2022).
Deniz, Ö., Frost, J. M. & Branco, M. R. Regulation of transposable elements by DNA modifications. Nat. Rev. Genet. 20, 417–431 (2019).
Padeken, J., Methot, S. P. & Gasser, S. M. Establishment of H3K9-methylated heterochromatin and its functions in tissue differentiation and maintenance. Nat. Rev. Mol. Cell Biol. https://doi.org/10.1038/s41580-022-00483-w (2022).
Hermant, C. & Torres-Padilla, M.-E. TFs for TEs: the transcription factor repertoire of mammalian transposable elements. Genes Dev. 35, 22–39 (2021).
Hermann, A., Goyal, R. & Jeltsch, A. The Dnmt1 DNA-(cytosine-C5)-methyltransferase methylates DNA processively with high preference for hemimethylated target sites. J. Biol. Chem. 279, 48350–48359 (2004).
Okano, M., Bell, D. W., Haber, D. A. & Li, E. DNA methyltransferases Dnmt3a and Dnmt3b are essential for de novo methylation and mammalian development. Cell 99, 247–257 (1999).
Ito, S. et al. Role of Tet proteins in 5mC to 5hmC conversion, ES-cell self-renewal and inner cell mass specification. Nature 466, 1129–1133 (2010).
Tahiliani, M. et al. Conversion of 5-methylcytosine to 5-hydroxymethylcytosine in mammalian DNA by MLL partner TET1. Science 324, 930–935 (2009).
Liu, J. et al. N6-methyladenosine of chromosome-associated regulatory RNA regulates chromatin state and transcription. Science 367, 580–586 (2020).
Xu, W. et al. METTL3 regulates heterochromatin in mouse embryonic stem cells. Nature https://doi.org/10.1038/s41586-021-03210-1 (2021).
Liu, J. et al. The RNA m6A reader YTHDC1 silences retrotransposons and guards ES cell identity. Nature 591, 322–326 (2021).
Wei, J. et al. FTO mediates LINE1 m6A demethylation and chromatin regulation in mESCs and mouse development. Science 376, 968–973 (2022).
Li, Y. et al. N6-Methyladenosine co-transcriptionally directs the demethylation of histone H3K9me2. Nat. Genet. 52, 870–877 (2020).
Lee, J.-H. et al. Enhancer RNA m6A methylation facilitates transcriptional condensate formation and gene activation. Mol. Cell 81, 3368–3385.e9 (2021).
Shi, H., Wei, J. & He, C. Where, when, and how: context-dependent functions of RNA methylation writers, readers, and erasers. Mol. Cell 74, 640–650 (2019).
Yang, Y., Hsu, P. J., Chen, Y.-S. & Yang, Y.-G. Dynamic transcriptomic m6A decoration: writers, erasers, readers and functions in RNA metabolism. Cell Res. 28, 616–624 (2018).
Michalak, E. M., Burr, M. L., Bannister, A. J. & Dawson, M. A. The roles of DNA, RNA and histone methylation in ageing and cancer. Nat. Rev. Mol. Cell Biol. 20, 573–589 (2019).
Zhao, S., Allis, C. D. & Wang, G. G. The language of chromatin modification in human cancers. Nat. Rev. Cancer 21, 413–430 (2021).
Jambhekar, A., Dhall, A. & Shi, Y. Roles and regulation of histone methylation in animal development. Nat. Rev. Mol. Cell Biol. 20, 625–641 (2019).
Deng, S. et al. RNA m6A regulates transcription via DNA demethylation and chromatin accessibility. Nat. Genet. 54, 1427–1437 (2022).
Hsu, P. J. et al. Ythdc2 is an N6-methyladenosine binding protein that regulates mammalian spermatogenesis. Cell Res. 27, 1115–1127 (2017).
Fuentes, D. R., Swigut, T. & Wysocka, J. Systematic perturbation of retroviral LTRs reveals widespread long-range effects on human gene regulation. eLife 7, e35989 (2018).
Gu, B. et al. Transcription-coupled changes in nuclear mobility of mammalian cis-regulatory elements. Science 359, 1050–1055 (2018).
Branon, T. C. et al. Efficient proximity labeling in living cells and organisms with TurboID. Nat. Biotechnol. 36, 880–887 (2018).
Kim, D. I. et al. Probing nuclear pore complex architecture with proximity-dependent biotinylation. Proc. Natl Acad. Sci. USA 111, E2453–E2461 (2014).
Schmidtmann, E., Anton, T., Rombaut, P., Herzog, F. & Leonhardt, H. Determination of local chromatin composition by CasID. Nucleus 7, 476–484 (2016).
Gao, X. D., Rodríguez, T. C. & Sontheimer, E. J. Adapting dCas9-APEX2 for subnuclear proteomic profiling. Methods Enzymol. 616, 365–383 (2019).
Liu, X. et al. Multiplexed capture of spatial configuration and temporal dynamics of locus-specific 3D chromatin by biotinylated dCas9. Genome Biol. 21, 59 (2020).
Ugur, E., Bartoschek, M. D. & Leonhardt, H. Locus-specific chromatin proteome revealed by mass spectrometry-based CasID. Methods Mol. Biol. 2175, 109–121 (2020).
Xie, W. et al. Base-resolution analyses of sequence and parent-of-origin dependent DNA methylation in the mouse genome. Cell 148, 816–831 (2012).
Leung, D. et al. Integrative analysis of haplotype-resolved epigenomes across human tissues. Nature 518, 350–354 (2015).
Zhang, Y. et al. Transcriptionally active HERV-H retrotransposons demarcate topologically associating domains in human pluripotent stem cells. Nat. Genet. 51, 1380–1388 (2019).
Glinsky, G. V. Transposable elements and DNA methylation create in embryonic stem cells human-specific regulatory sequences associated with distal enhancers and noncoding RNAs. Genome Biol. Evol. 7, 1432–1454 (2015).
Gorkin, D. U., Leung, D. & Ren, B. The 3D genome in transcriptional regulation and pluripotency. Cell Stem Cell 14, 762–775 (2014).
Shi, J. & Vakoc, C. R. The mechanisms behind the therapeutic activity of BET bromodomain inhibition. Mol. Cell 54, 728–736 (2014).
Dhalluin, C. et al. Structure and ligand of a histone acetyltransferase bromodomain. Nature 399, 491–496 (1999).
Ernst, J. & Kellis, M. Discovery and characterization of chromatin states for systematic annotation of the human genome. Nat. Biotechnol. 28, 817–825 (2010).
Ernst, J. et al. Mapping and analysis of chromatin state dynamics in nine human cell types. Nature 473, 43–49 (2011).
Jao, C. Y. & Salic, A. Exploring RNA transcription and turnover in vivo by using click chemistry. Proc. Natl Acad. Sci. USA 105, 15779–15784 (2008).
Saito, Y. et al. YTHDC2 control of gametogenesis requires helicase activity but not m6A binding. Genes Dev. 36, 180–194 (2022).
Li, L. et al. The XRN1-regulated RNA helicase activity of YTHDC2 ensures mouse fertility independently of m6A recognition. Mol. Cell 82, 1678–1690.e12 (2022).
Larivera, S. & Meister, G. Domain confusion 2: m6A-independent role of YTHDC2. Mol. Cell 82, 1608–1609 (2022).
Lu, X. et al. The retrovirus HERVH is a long noncoding RNA required for human embryonic stem cell identity. Nat. Struct. Mol. Biol. 21, 423–425 (2014).
Wang, J. et al. Primate-specific endogenous retrovirus-driven transcription defines naive-like stem cells. Nature 516, 405–409 (2014).
Ohnuki, M. et al. Dynamic regulation of human endogenous retroviruses mediates factor-induced reprogramming and differentiation potential. Proc. Natl Acad. Sci. USA 111, 12426–12431 (2014).
Koyanagi-Aoi, M. et al. Differentiation-defective phenotypes revealed by large-scale analyses of human pluripotent stem cells. Proc. Natl Acad. Sci. USA 110, 20569–20574 (2013).
Takahashi, K. et al. The pluripotent stem cell-specific transcript ESRG is dispensable for human pluripotency. PLoS Genet. 17, e1009587 (2021).
Chambers, S. M. et al. Highly efficient neural conversion of human ES and iPS cells by dual inhibition of SMAD signaling. Nat. Biotechnol. 27, 275–280 (2009).
Verma, N. et al. TET proteins safeguard bivalent promoters from de novo methylation in human embryonic stem cells. Nat. Genet. 50, 83–95 (2018).
Ahmad, R. et al. Functional neuronal cells generated by human parthenogenetic stem cells. PLoS ONE 7, e42800 (2012).
Zheng, Y. et al. Single-cell analysis of embryoids reveals lineage diversification roadmaps of early human development. Cell Stem Cell 29, 1402–1419.e8 (2022).
Thakore, P. I. et al. Highly specific epigenome editing by CRISPR-Cas9 repressors for silencing of distal regulatory elements. Nat. Methods 12, 1143–1149 (2015).
Loewer, S. et al. Large intergenic non-coding RNA-RoR modulates reprogramming of human induced pluripotent stem cells. Nat. Genet. 42, 1113–1117 (2010).
Cheng, E.-C. & Lin, H. Repressing the repressor: a lincRNA as a microRNA sponge in embryonic stem cell self-renewal. Dev. Cell 25, 1–2 (2013).
Wang, Y. et al. Endogenous miRNA sponge lincRNA-RoR regulates Oct4, Nanog, and Sox2 in human embryonic stem cell self-renewal. Dev. Cell 25, 69–80 (2013).
Konermann, S. et al. Transcriptome engineering with RNA-targeting type VI-D CRISPR effectors. Cell 173, 665–676.e14 (2018).
Yan, W. X. et al. Cas13d is a compact RNA-targeting type VI CRISPR effector positively modulated by a WYL-domain-containing accessory protein. Mol. Cell 70, 327–339.e5 (2018).
Batista, P. J. et al. m6A RNA modification controls cell fate transition in mammalian embryonic stem cells. Cell Stem Cell 15, 707–719 (2014).
Van Nostrand, E. L. et al. Robust transcriptome-wide discovery of RNA-binding protein binding sites with enhanced CLIP (eCLIP). Nat. Methods 13, 508–514 (2016).
Sridhar, B. et al. Systematic mapping of RNA-chromatin interactions in vivo. Curr. Biol. 27, 602–609 (2017).
Yankova, E. et al. Small-molecule inhibition of METTL3 as a strategy against myeloid leukaemia. Nature 593, 597–601 (2021).
Zheng, G. et al. ALKBH5 is a mammalian RNA demethylase that impacts RNA metabolism and mouse fertility. Mol. Cell 49, 18–29 (2013).
Xia, Z. et al. Epitranscriptomic editing of the RNA N6-methyladenosine modification by dCasRx conjugated methyltransferase and demethylase. Nucleic Acids Res. 49, 7361–7374 (2021).
Dixon, G. et al. QSER1 protects DNA methylation valleys from de novo methylation. Science 372, eabd0875 (2021).
Liu, X. S. et al. Editing DNA methylation in the mammalian genome. Cell 167, 233–247.e17 (2016).
Ziller, M. J. et al. Charting a dynamic DNA methylation landscape of the human genome. Nature 500, 477–481 (2013).
Robertson, K. D. DNA methylation and human disease. Nat. Rev. Genet. 6, 597–610 (2005).
Xu, W. & Shen, H. When RNA methylation meets DNA methylation. Nat. Genet. 54, 1261–1262 (2022).
McKenna, A. & Shendure, J. FlashFry: a fast and flexible tool for large-scale CRISPR target design. BMC Biol. 16, 74 (2018).
Huang, D. W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57 (2009).
Sherman, B. T. et al. DAVID: a web server for functional enrichment analysis and functional annotation of gene lists (2021 update). Nucleic Acids Res. https://doi.org/10.1093/nar/gkac194 (2022).
Krämer, A., Green, J., Pollard, J. Jr & Tugendreich, S. Causal analysis approaches in Ingenuity Pathway Analysis. Bioinformatics 30, 523–530 (2014).
Sanjana, N. E., Shalem, O. & Zhang, F. Improved vectors and genome-wide libraries for CRISPR screening. Nat. Methods 11, 783–784 (2014).
Diao, Y., Wang, X. & Wu, Z. SOCS1, SOCS3, and PIAS1 promote myogenic differentiation by inhibiting the leukemia inhibitory factor-induced JAK1/STAT1/STAT3 pathway. Mol. Cell. Biol. 29, 5084–5093 (2009).
Chelmicki, T. et al. m6A RNA methylation regulates the fate of endogenous retroviruses. Nature https://doi.org/10.1038/s41586-020-03135-1 (2021).
Zeng, Y. et al. Refined RIP-seq protocol for epitranscriptome analysis with low input materials. PLoS Biol. 16, e2006092 (2018).
Picelli, S. et al. Full-length RNA-seq from single cells using Smart-seq2. Nat. Protoc. 9, 171–181 (2014).
Chen, S., Zhou, Y., Chen, Y. & Gu, J. fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34, i884–i890 (2018).
Kim, D., Paggi, J. M., Park, C., Bennett, C. & Salzberg, S. L. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat. Biotechnol. 37, 907–915 (2019).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Tarasov, A., Vilella, A. J., Cuppen, E., Nijman, I. J. & Prins, P. Sambamba: fast processing of NGS alignment formats. Bioinformatics 31, 2032–2034 (2015).
Feng, J., Liu, T., Qin, B., Zhang, Y. & Liu, X. S. Identifying ChIP-seq enrichment using MACS. Nat. Protoc. 7, 1728–1740 (2012).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Ramírez, F. et al. deepTools2: a next generation web server for deep-sequencing data analysis. Nucleic Acids Res. 44, W160–W165 (2016).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Acknowledgements
We thank B. Hogan and K. Poss for the discussion, S. Soderling and C. Nicchitta for advising on the BioID experiments, S. Endow for recombinant protein purification, and D. Huangfu and C. He for sharing reagents. Y.D. acknowledges support from the Duke Regeneration Center, the Duke Whitehead Scholarship, the Glenn Foundation for Medical Research and AFAR Grants for Junior Faculty, grant nos. NIH U01HL156064 and R35HG011328. Y. Xiang is supported by a Duke Regeneration Center postdoctoral fellowship. Y. Xu is supported by the Center for Advanced Genomic Technologies and the Duke Regeneration Center Fellowships.
Author information
Authors and Affiliations
Contributions
T.S. and Y.D. conceived the idea for this study. T.S. performed all the experiments and data analysis with the help from Y. Xu, Y. Xiang and J.O. for bioinformatics. T.S. and E.S. performed LC–MS/MS proteomics analysis. T.S. and Y.D. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
All authors declare no competing interests.
Peer review
Peer review information
Nature Genetics thanks Yun-Gui Yang, Todd Macfarlan and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 CARGO targets LTR7 sequences in human embryonic stem cells.
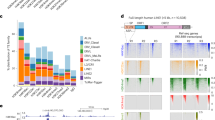
a, A total of 15 sgRNAs targeting the conserved LTR7 sequences were designed for CARGO cloning. The predicted coverage ratio of all LTR7 loci is calculated based on the sequence similarity between sgRNA and LTR7 sequences. b, Classification of LTR7 loci based on their histone modification signature in H1 hESCs. H1 hESC ChIP-seq datasets were obtained from ENCODE data portal. c, Histone H3K27ac ChIP-seq signals of LTR7 loci in H1 hESCs and H1-derived differentiated lineages, including mesendoderm (ME), mesenchymal stem cell (MSC), neural progenitor cells (NPC), and trophoblast-like cells (TBL). The ChIP-seq datasets are generated by Epigenome Roadmap projects. P values determined by two-sided Wilcoxon signed-rank test. n = 1 biologically independent experiment. In the plots, center lines represent the median value, box limits the 25th and 75th percentiles, and whiskers denote minima and maxima (1.5× the interquartile range). d, Illustration of the predicated TF binding motifs (NANOG, KLF4, SOX3) and the CARGO sgRNA target sites across all LTR7 sequences. Color key: The darkness indicates counts of predicted sgRNA target sequences (red) and the number of predicated TF motifs (orange, purple, and green). In the heatmap (bottom), each row in gray color represents one LTR7 element, with pink color highlighting the sequences targeted by CARGO sgRNAs. The LTR7 elements are first sorted based on their histone modifications (H3K27ac, H3K9me3, H3K4me3, and other histone marks) and then by the number of sgRNA targeting sites. e, f, CARGO-BioID does not affect the expression of HERV-H RNA and its retroviral genes (e, RT-qPCR), or histone H3K9me3 and H3K27ac modifications on LTR7 DNA sequences (f, ChIP-qPCR). Data represent mean ± s.e.m. from three independent experiments. ns, not significant. P values were calculated by two-tailed Student′s test. In the box plots, center lines represent the median value, box limits the 25th and 75th percentiles, and whiskers denote minima and maxima (1.5× the interquartile range). g, h, The purified biotinylated nuclear proteins prepared from LTR7 CARGO-BioID experiment and subjected to proteomics analysis were analyzed by western blot (g) and silver staining (h). n = 3 biologically independent experiments.
Extended Data Fig. 2 YTHDC2 occupies active genomic loci and limits transcriptional silencing of LTR7/HERV-H loci in hESCs.
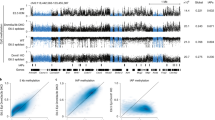
a, Validation of YTHDC2 binding to LTR7 loci using Western blot. n = 3 biologically independent experiments. b, A permutation test was conducted to determine the enriched repeat families overlapping with YTHDC2 ChIP-seq peaks. Exact p values are in Supplementary Table 3,a. c, The indicated genomic datasets are presented as heatmaps centered on YTHDC2 ChIP-seq peaks. Each row represents one YTHDC2 peak. For all the heatmaps, the rows are sorted in the same order based on YTHDC2 ChIP-seq signal. d, Pie chart displaying the proportion of 15 chromatin states (defined by ChromHMM in H1 hESCs) in the YTHDC2 occupies DNA sequences. e, A permutation test was carried out in a similar way described in (b) to identify the enriched chromatin states in the YTHDC2-occupied DNAs. Exact p values are in Supplementary Table 3,b. f, YTHDC2 knock-out in the two mutant clones revealed by Western blot. g, h, YTHDC2 KO did not alter the expression of pluripotency markers at RNA (g) and protein (h) levels. KO no. 1, KO no. 2: two independent YTHDC2 KO clones. i, HERV-H RNA decay rate in the indicated hESCs. j, The protein expressions of wild-type and two mutant forms of YTHDC2 was restored in the YTHDC2 KO hESCs. n = 3 biologically independent experiments. k, l, HERV-H expression in control and mutant cells analyzed by RNA-seq are shown as boxplot (k) and volcano plot (l). P value determined by two-sided Wilcoxon signed-rank test. n = 2 biologically independent experiments. In the plots, center lines represent the median value, box limits the 25th and 75th percentiles, and whiskers denote minima and maxima (1.5× the interquartile range) (k). Blue and red dots: the downregulated or upregulated HERV-H RNAs in YTHDC2 KO compared to wild-type cells(l). Data represent mean ± s.e.m. from three experiments. ns, not significant; *p < 0.05, **p < 0.01, ***p < 0.001. P values were calculated by two-tailed Student′s test (g, i). Exact p values are in Source Data 1.
Extended Data Fig. 3 The YTHDC2/LTR7 axis prevents the neuronal fate of hESC via LINC-ROR.
a, Gene Ontology (GO) analysis of the genes differentially expressed in YTHDC2 KO versus control hESCs. b, GO analysis of the genes whose promoters are located within 20kb of YTHDC2 ChIP-seq peaks. c, YTHDC2 KO promotes neuronal fate of hESC-derived neuronal cells and embryoid bodies (EBs). The protein levels of the indicated neuronal markers were analyzed by Western blot. n = 3 biologically independent experiments. d, RT-qPCR analysis of gene expression of EBs derived from wild-type control versus YTHDC2 KO hESCs. The marker genes for neuronal, endoderm, mesoderm, and trophectoderm lineages were analyzed. e–h, Epigenetic silencing of LTR7 loci by dCas9-KRAB/CARGO (CRISPRi) lead to: increased histone H3K9me3 and decreased H3K27ac modifications on LTR7 loci, determined by ChIP-seq(e, f); reduction of HERV-H transcripts, determined by RNA-seq (g) and RT-qPCR (h). i-k, LTR7 CRISPRi did not affect hESCs morphology (i) or the expression of pluripotency markers at RNA (j) and protein (k) levels. l, LTR7 CRISPRi promotes neuronal fate of hESC-derived neuronal cells and embryoid bodies (EBs). The protein levels of the indicated neuronal markers were analyzed by Western blot as indicated. n = 3 biologically independent experiments. m, LINC-ROR expression is substantially decreased in YTHDC2 KO compared to the wild-type control (Ctrl) cells, determined by RT-qPCR. n, RT-qPCR analysis reveals a notable reduction of LINC-ROR in hESCs upon epigenetic silencing of LTR7s. o, RT-qPCR analysis showing efficient knockdown (KD) of LINC-ROR by two gRNAs targeting LINC-ROR sequence. p, CasRx-mediated KD of LINC-ROR did not alter the expression of pluripotency markers in hESCs, determined by RT-qPCR analysis. Data represent mean ± s.e.m. from three (h, j, m-p) or four (d)independent experiments. ns, not significant; *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001. P values were calculated by two-tailed Student′s test in (d, h, j, m-p). Exact p values are in Source Data 1.
Extended Data Fig. 4 YTHDC2, m6A-modified HERV-H RNAs, and LTR7 DNA physically interact with each other.
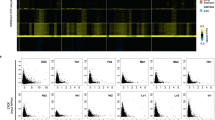
a, Heatmap showing enriched YTHDC2 binding (normalized against input control) along all the HERV-H transcripts, determined by RIP-seq. b, Aggregated reads counts of YTDHC2 eCLIP-seq data showing enriched YTHDC2 binding across all HERV-H RNAs. Red line: YTDHC2 pulldown, blue line: Input signal. c, A representative genome browser snapshot illustrating the binding of YTHDC2 on one represent LTR7/HERV-H locus revealed by YTHDC2 eCLIP-seq and RIP-seq. d, The 1,119 mRNAs associated with enriched YTHDC2 binding (YTHDC2 RIP/input fold enrichment > 2, p-value<0.05) are expressed at a comparable level with those mRNAs with less enriched YTHDC2 binding. P values determined by two-sided Wilcoxon signed-rank test. n = 2 biologically independent experiments. In the plots, center lines represent the median value, box limits the 25th and 75th percentiles, and whiskers denote minima and maxima (1.5× the interquartile range). e, A ‘CCGGACGC’ motif harboring the conserved m6A motif GGACU is enriched in the 1,119 mRNA sequences associated with enriched YTHDC2 binding. f, Coomassie blue staining showing the recombinant protein of the YTH-domain of human YTHDC2 (1288-1418aa) purified from E.coli. g-i, MAGRI data showing the patterns of HERV-H RNA interacting with the indicated DNA sequences. HERV-H RNAs interact with the entire LTR7/HERV-H elements while exhibiting strongest interactions with the 5′- promoter LTR7s (TSS LTR7) (g). HERV-H RNAs were separated based on whether they are bound by YTHDC2 (h, blue line) or modified by m6A (i, blue line). The YTHDC2-bound and m6A-modified HERV-H RNAs exhibited stronger interactions with their TSS LTRs compared to those without YTHDC2 binding or m6A modification.
Extended Data Fig. 5 YTHDC2 binds to LTR7/HERV-H loci and prevents their silencing in a m6A dependent manner.
a, Two-day STM2457 treatment effectively decreased the m6A modification of HERV-H RNAs, determined by m6A MeRIP-qPCR. b, HERV-H RNA degradation detection by qRT-PCR. The cells were treated by STM2457 or control DMSO for 2 days before sample collection. Actinomycin D was added to the cell culture at the indicated times. c, Detection of METTL3 protein level in hESCs treated with STM2457. n = 3 biologically independent experiments. d, e, YTHDC2 RIP-qPCR (d) and ChIP-qPCR (e) revealing decreased YTHDC2 binding to HERV-H RNAs and LTR7 DNA after 2-day STM2457 treatment compared to cells treated by DMSO. In the box plots, center lines represent the median value, box limits the 25th and 75th percentiles, and whiskers denote minima and maxima (1.5× the interquartile range). f, LINC-ROR degradation determined by qRT-PCR. g-i, The dCasRx-ALHKB5 was expressed in the wild-type or YTHDC2 KO hESCs, together with control (Ctrl, black) gRNA or two independent gRNAs targeting the m6A modified sequence of LINC-ROR (gRNA1, gRNA2, blue and red), to remove the m6A modification of LINC-ROR. m6A MeRIP-qPCR analysis showing a substantial reduction of m6A modification on LINC-ROR in both WT and KO cells(g). RT-qPCR analysis shows reduced LINC-ROR RNA level in WT cells, but no further reduction of LINC-ROR in the KO cells upon LINC-ROR m6A removal(h). Erasing m6A from LINC-ROR only deposits H3K9me3 modification on the two LTR7 loci flanking LINC-ROR locus in WT cells, but not in KO cells(i). In the box plots, center lines represent the median value, box limits the 25th and 75th percentiles, and whiskers denote minima and maxima (1.5× the interquartile range). Data represent mean ± s.e.m. from three (b, d, e, f, i) or four (a, g, h) independent experiments. ns, not significant; *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001. P values were calculated by two-tailed Student′s test in (a, b, d-i). Exact p values are in Source Data 1.
Extended Data Fig. 6 YTHDC2 recruits TET1 to demethylate LTR7/HERV-H DNA.
a, Anti-V5 IP and western for TET1-V5 lines alongside parental, non-tagged WT cells. n = 3 biologically independent experiments. b, c, Comparison of the TET1-V5 ChIP-seq data generated in this study and in a prior study. The similarities of the two data sets were assessed by visualization of genome browser snapshot (b) and Pearson correlation(c). d, Protein co-immunoprecipitation experiments using lysates from cells transduced with indicated plasmids. n = 3 biologically independent experiments. e, Protein co-immunoprecipitation experiments using lysates treated with or without RNase. n = 3 biologically independent experiments. f, Protein co-immunoprecipitation experiments using lysates from cells treated with or without STM2457. n = 3 biologically independent experiments. g, YTHDC2 and TET1-V5 binding on LTR7s occupied by TET1-V5. h, Pearson correlation of YTHDC2 and TET1 ChIP-seq signals on co-occupied LTR7 loci. i, Volcano plot of RNA-seq data showing differentially expressed HERV-H transcripts in TET1 KO hESCs compared to the wild-type cells. The blue and red dots indicating substantially changed HERV-H RNAs (p-value < 0.05). j, ChIP-qPCR analysis of TET1 binding on the two LTR7 loci flanking the LINC-ROR in WT or YTHDC2 KO cells upon m6A editing. k, l, MeDIP-qPCR analyses of the two LTR7 loci flanking the LINC-ROR locus using antibodies against 5mC (k) or 5hmC (l). In the box plots (j-l), center lines represent the median value, box limits the 25th and 75th percentiles, and whiskers denote minima and maxima (1.5× the interquartile range) n = 3 biologically independent experiments. ns, not significant; *p < 0.05, **p < 0.01, ***p < 0.001. P values were calculated by two-tailed Student′s test in (j-l). Exact p values are in Source Data 1.
Supplementary information
Supplementary Information
Supplementary Discussion, Figs. 1–6 and uncropped blots for Supplementary Fig. 2.
Supplementary Tables 1–4
Table 1. Oligonucleotide sequences and antibodies used in this study. Table 2a–c. Analysis of LTR7-CARGO-BioID proteome data. The proteins enriched in LTR7 DNA sequences in hESCs (Table 2a) are subjected to Ingenuity Pathway Analysis (Table 2b) and Gene Ontology (Table 2c) analysis. Table 3a–c. Enriched repeats families and chromatin states in YTHDC2-occupied DNA and RNA sequences. Table 3a. Enrichment analysis of repeat families in YTHDC2 ChIP–seq peaks. Table 3b. Permutation test of YTHDC2-associated chromatin state features. Table 3c. Enrichment analysis of repeat families in YTHDC2 RIP-seq peaks. Table 4a–c. The differentially expressed genes in pluripotent hESCs caused by YTHDC2 KO, LTR7 epigenetic silencing and neural differentiation. Table 4a. The differentially expressed genes in YTHDC2 KO compared with WT cells. Table 4b. The differentially expressed genes in LTR7 CRISPRi compared with control cells. Table 4c. The differentially expressed genes in neural cells compared with WT cells.
Supplementary Data 1
Enrichment analysis of chromatin states in YTHDC2 ChIP–seq peaks.
Supplementary Data 2
Exact P values for supplementary figures.
Source data
Figs. 1–6 and Extended Data Figs. 2, 3, 5 and 6
Statistical source data Figs. 1–6 and Extended Data Figs. 2, 3, 5 and 6.
Source Data Fig. 1
Unprocessed western blots and/or gels for Fig. 1e.
Source Data Fig. 4
Unprocessed western blots and/or gels for Fig. 4f,g.
Source Data Fig. 6
Unprocessed western blots and/or gels for Fig. 6a.
Source Data Extended Data Fig. 1
Unprocessed western blots and/or gels for Extended Data Fig. 1g,h.
Source Data Extended Data Fig. 2
Unprocessed western blots and/or gels for Extended Data Fig. 2a,f.
Source Data Extended Data Fig. 3
Unprocessed western blots and/or gels for Extended Data Fig. 3c,l.
Source Data Extended Data Fig. 5
Unprocessed western blots and/or gels for Extended Data Fig. 5c.
Source Data Extended Data Fig. 6
Unprocessed western blots and/or gels for Extended Data Fig. 6a,d–f.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Sun, T., Xu, Y., Xiang, Y. et al. Crosstalk between RNA m6A and DNA methylation regulates transposable element chromatin activation and cell fate in human pluripotent stem cells. Nat Genet 55, 1324–1335 (2023). https://doi.org/10.1038/s41588-023-01452-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-023-01452-5
This article is cited by
-
Selection on synonymous sites: the unwanted transcript hypothesis
Nature Reviews Genetics (2024)
-
Regulation and function of transposable elements in cancer genomes
Cellular and Molecular Life Sciences (2024)
-
Transcriptome-wide profiling identifies colon cancer-associated m6A transcripts and potential RNA methyl modifiers
Molecular Biology Reports (2024)