Abstract
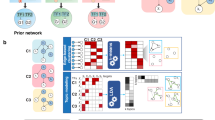
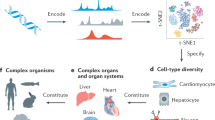
Despite extensive efforts to generate and analyze reference genomes, genetic models to predict gene regulation and cell fate decisions are lacking for most species. Here, we generated whole-body single-cell transcriptomic landscapes of zebrafish, Drosophila and earthworm. We then integrated cell landscapes from eight representative metazoan species to study gene regulation across evolution. Using these uniformly constructed cross-species landscapes, we developed a deep-learning-based strategy, Nvwa, to predict gene expression and identify regulatory sequences at the single-cell level. We systematically compared cell-type-specific transcription factors to reveal conserved genetic regulation in vertebrates and invertebrates. Our work provides a valuable resource and offers a new strategy for studying regulatory grammar in diverse biological systems.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Raw data files for the RNA sequencing analysis reported in this paper have been deposited in is the NCBI Gene Expression Omnibus under accession number GSE178151. Digital expression matrices are available at https://figshare.com/s/ecc05b1051fb5678fd3e. Nvwa data can be accessed at http://bis.zju.edu.cn/nvwa/.
Code availability
The source code for reproducing our analysis and running and training the Nvwa models is available at GitHub (https://github.com/JiaqiLiZju/Nvwa/) and Zenodo (https://zenodo.org/record/6806748) (JiaqiLiZju/Nvwa: release v.1.0, 2022).
References
Han, X. et al. Construction of a human cell landscape at single-cell level. Nature 581, 303–309 (2020).
Tabula Sapiens, C. et al. The Tabula Sapiens: a multiple-organ, single-cell transcriptomic atlas of humans. Science 376, eabl4896 (2022).
Han, X. et al. Mapping the Mouse Cell Atlas by Microwell-Seq. Cell 173, 1307 (2018).
Tabula Muris, C. et al. Single-cell transcriptomics of 20 mouse organs creates a Tabula Muris. Nature 562, 367–372 (2018).
Jiang, M. M. et al. Characterization of the zebrafish cell landscape at single-cell resolution. Front. Cell Dev. Biol. 9, 743421 (2021).
Cao, J. Y. et al. Comprehensive single-cell transcriptional profiling of a multicellular organism. Science 357, 661–667 (2017).
Cao, C. et al. Comprehensive single-cell transcriptome lineages of a proto-vertebrate. Nature 571, 349–354 (2019).
Fincher, C. T., Wurtzel, O., de Hoog, T., Kravarik, K. M. & Reddien, P. W. Cell type transcriptome atlas for the planarian Schmidtea mediterranea. Science 360, eaaq1736 (2018).
Sebe-Pedros, A. et al. Cnidarian cell type diversity and regulation revealed by whole-organism single-cell RNA-seq. Cell 173, 1520–1534.e20 (2018).
Li, H. et al. Fly Cell Atlas: A single-nucleus transcriptomic atlas of the adult fruit fly. Science 375, eabk2432 (2022).
Arendt, D. et al. The origin and evolution of cell types. Nat. Rev. Genet. 17, 744–757 (2016).
Wang, J. et al. Tracing cell-type evolution by cross-species comparison of cell atlases. Cell Rep. 34, 108803 (2021).
Agarwal, V. & Shendure, J. Predicting mRNA abundance directly from genomic sequence using deep convolutional neural networks. Cell Rep. 31, 107663 (2020).
Zhou, J. & Troyanskaya, O. G. Predicting effects of noncoding variants with deep learning-based sequence model. Nat. Methods 12, 931–934 (2015).
Kelley, D. R., Snoek, J. & Rinn, J. L. Basset: learning the regulatory code of the accessible genome with deep convolutional neural networks. Genome Res. 26, 990–999 (2016).
Avsec, Z. et al. Effective gene expression prediction from sequence by integrating long-range interactions. Nat. Methods 18, 1196–1203 (2021).
Maslova, A. et al. Deep learning of immune cell differentiation. Proc. Natl Acad. Sci. USA 117, 25655–25666 (2020).
Satija, R., Farrell, J. A., Gennert, D., Schier, A. F. & Regev, A. Spatial reconstruction of single-cell gene expression data. Nat. Biotechnol. 33, 495–502 (2015).
Buchon, N., Silverman, N. & Cherry, S. Immunity in Drosophila melanogaster–from microbial recognition to whole-organism physiology. Nat. Rev. Immunol. 14, 796–810 (2014).
Krausgruber, T. et al. Structural cells are key regulators of organ-specific immune responses. Nature 583, 296–302 (2020).
van Dijk, D. et al. Recovering gene interactions from single-cell data using data diffusion. Cell 174, 716 (2018).
Tarashansky, A. J. et al. Mapping single-cell atlases throughout Metazoa unravels cell type evolution. eLife 10, e66747 (2021).
Rui, L., Schmitz, R., Ceribelli, M. & Staudt, L. M. Malignant pirates of the immune system. Nat. Immunol. 12, 933–940 (2011).
Zhou, J. et al. Deep learning sequence-based ab initio prediction of variant effects on expression and disease risk. Nat. Genet. 50, 1171–1179 (2018).
Kelley, D. R. et al. Sequential regulatory activity prediction across chromosomes with convolutional neural networks. Genome Res. 28, 739–750 (2018).
de Souza, N. The ENCODE project. Nat. Methods 9, 1046 (2012).
Srivastava, A. K. & Schlessinger, D. Structure and organization of ribosomal DNA. Biochimie 73, 631–638 (1991).
Suzuki, H., Moriwaki, K. & Sakurai, S. Sequences and evolutionary analysis of mouse 5S rDNAs. Mol. Biol. Evol. 11, 704–710 (1994).
Zentner, G. E., Balow, S. A. & Scacheri, P. C. Genomic characterization of the mouse ribosomal DNA locus. G3 4, 243–254 (2014).
Aibar, S. et al. SCENIC: single-cell regulatory network inference and clustering. Nat. Methods 14, 1083–1086 (2017).
Gupta, S., Stamatoyannopoulos, J. A., Bailey, T. L. & Noble, W. S. Quantifying similarity between motifs. Genome Biol. 8, R24 (2007).
Cusanovich, D. A. et al. A single-cell atlas of in vivo mammalian chromatin accessibility. Cell 174, 1309–1324.e18 (2018).
Hannenhalli, S. & Kaestner, K. H. The evolution of Fox genes and their role in development and disease. Nat. Rev. Genet. 10, 233–240 (2009).
Shafer, M. E. R. Cross-species analysis of single-cell transcriptomic data. Front. Cell Dev. Biol. 7, 175 (2019).
Picelli, S. et al. Full-length RNA-seq from single cells using Smart-seq2. Nat. Protoc. 9, 171–181 (2014).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Zheng, G. X. et al. Massively parallel digital transcriptional profiling of single cells. Nat. Commun. 8, 14049 (2017).
McGinnis, C. S., Murrow, L. M. & Gartner, Z. J. DoubletFinder: doublet detection in single-cell RNA sequencing data using artificial nearest neighbors. Cell Syst. 8, 329–337.e4 (2019).
Wolf, F. A., Angerer, P. & Theis, F. J. SCANPY: large-scale single-cell gene expression data analysis. Genome Biol. 19, 15 (2018).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902.e21 (2019).
Shao, Y. et al. Genome and single-cell RNA-sequencing of the earthworm Eisenia andrei identifies cellular mechanisms underlying regeneration. Nat. Commun. 11, 2656 (2020).
Rozanski, A. et al. PlanMine 3.0–improvements to a mineable resource of flatworm biology and biodiversity. Nucleic Acids Res. 47, D812–D820 (2019).
Satou, Y., Kawashima, T., Shoguchi, E., Nakayama, A. & Satoh, N. An integrated database of the ascidian, Ciona intestinalis: towards functional genomics. Zool. Sci. 22, 837–843 (2005).
Haas, B. J. et al. De novo transcript sequence reconstruction from RNA-seq using the Trinity platform for reference generation and analysis. Nat. Protoc. 8, 1494–1512 (2013).
Emms, D. M. & Kelly, S. OrthoFinder: solving fundamental biases in whole genome comparisons dramatically improves orthogroup inference accuracy. Genome Biol. 16, 157 (2015).
Crow, M., Paul, A., Ballouz, S., Huang, Z. J. & Gillis, J. Characterizing the replicability of cell types defined by single cell RNA-sequencing data using MetaNeighbor. Nat. Commun. 9, 884 (2018).
Fischer, S., Crow, M., Harris, B. D. & Gillis, J. Scaling up reproducible research for single-cell transcriptomics using MetaNeighbor. Nat. Protoc. 16, 4031–4067 (2021).
Hu, H. et al. AnimalTFDB 3.0: a comprehensive resource for annotation and prediction of animal transcription factors. Nucleic Acids Res. 47, D33–D38 (2019).
dos Santos, G. et al. FlyBase: introduction of the Drosophila melanogaster Release 6 reference genome assembly and large-scale migration of genome annotations. Nucleic Acids Res. 43, D690–D697 (2015).
Dubaj Price, M. & Hurd, D. D. WormBase: a model organism database. Med. Ref. Serv. Q. 38, 70–80 (2019).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Grant, C. E., Bailey, T. L. & Noble, W. S. FIMO: scanning for occurrences of a given motif. Bioinformatics 27, 1017–1018 (2011).
Fornes, O. et al. JASPAR 2020: update of the open-access database of transcription factor binding profiles. Nucleic Acids Res. 48, D87–D92 (2020).
Harris, T. W. et al. WormBase: a modern model organism information resource. Nucleic Acids Res. 48, D762–D767 (2020).
Zhao, H. et al. CrossMap: a versatile tool for coordinate conversion between genome assemblies. Bioinformatics 30, 1006–1007 (2014).
Ramirez, F., Dundar, F., Diehl, S., Gruning, B. A. & Manke, T. deepTools: a flexible platform for exploring deep-sequencing data. Nucleic Acids Res. 42, W187–W191 (2014).
Yu, G., Wang, L. G. & He, Q. Y. ChIPseeker: an R/Bioconductor package for ChIP peak annotation, comparison and visualization. Bioinformatics 31, 2382–2383 (2015).
Ramirez, F. et al. High-resolution TADs reveal DNA sequences underlying genome organization in flies. Nat. Commun. 9, 189 (2018).
Simonyan, K., Vedaldi, A. & Zisserman, A. Deep inside convolutional networks: visualising image classification models and saliency maps. Preprint at https://doi.org/10.48550/arXiv.1312.6034 (2014).
Weirauch, M. T. et al. Determination and inference of eukaryotic transcription factor sequence specificity. Cell 158, 1431–1443 (2014).
Acknowledgements
G.G. is a participant in the Human Cell Atlas Project. We thank M. Chen, Y. Zhou, F. Gu, D. Wang, P. Xu, C. Li, K. Li and H. Wu for support on the project. We thank M20 (Hangzhou), G-BIO (Hangzhou), BGI (Shenzhen) and CNGB (Shenzhen) for supporting the sequencing experiments; and Vazyme (Nanjing) for supplying the customized enzymes used in the study. We also thank the core facility platform of Zhejiang University School of Medicine and the Center of Cryo-Electron Microscopy at Zhejiang University for computational resources, and the core facilities of Zhejiang University Medical Center and the Liangzhu Laboratory for technical support. This work was supported by National Natural Science Foundation of China grants 31930028 to G.G., 31922049 to X.H., 91842301 to G.G., 32000461 to J.W. and 62088101 to J.C.; National Key Research and Development Program grants 2018YFA0800503 to G.G., 2018YFA0107804 to G.G. and 2018YFA0107801 to X.H.; Fundamental Research Funds for the Central Universities (G.G.); and Alibaba-Zhejiang University Joint Research Center of Future Digital Healthcare.
Author information
Authors and Affiliations
Contributions
G.G., X.H. and J. Wang conceived the study. G.G. and X.H. supervised the study. Jiaqi Li designed the model. X.H., R.W., M.J., X.H., H.C., Xinru Wang, Xueyi Wang, Y.L., D.J. and T.Z. performed all the experiments. Jiaqi Li, J. Wang, P.Z., Y.M., Z.S., L.F., L.M., W.E., Y.F., H.W., D.L., H.W., Jingyu Li, Q.G. and C.Y. performed all computational analyses. H.L., J.M. and K.L. helped with the cell-type annotation. J. Wu., S.H. and J.C. guided model design and parameter optimization. G.G., J. Wang., Jiaqi Li and P.Z. wrote the initial draft of the manuscript. All authors participated in discussion of results and manuscript editing.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Genetics thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Zebrafish cell landscape was constructed using Microwell-seq.
(a-b) t-SNE visualization of 635,228 single cells from whole bodies across three stages of zebrafish, colored by stage (a) and cell lineage (b). (c) Heatmap showing the scaled average expression levels of zebrafish cell type-specific marker genes (left), and relative gene expression of representative cell type-specific markers for each cell type overlaid on t-SNE plots (right). (d) Heatmap showing the correspondence between zebrafish cell landscape in this study (row) and tissue-specific zebrafish dataset from Jiang et al., 2021 (column). Blue refers to a mean AUROC greater than 0.9. (e-f) t-SNE showing 24 subclusters from zebrafish C1 (neuron) (e), and violin plot showing the scale normalized expression levels of representative markers for each subcluster (f).
Extended Data Fig. 2 Drosophila cell landscape was constructed using Microwell-seq.
(a-b) t-SNE visualization of 276,706 single cells from whole bodies across two stages of Drosophila, colored by stage (a) and cell lineage (b). (c) Heatmap showing the scaled average expression levels of Drosophila cell type-specific marker genes (left), and relative gene expression of representative cell type-specific markers for each cluster overlaid on t-SNE plots (right). (d) Heatmap showing the correspondence between Drosophila cell landscape in this study (row) and tissue-specific fly cell atlas construed by Li et al., 2021 (column). Blue refers to a mean AUROC greater than 0.9. (e-f) t-SNE showing 21 subclusters from Drosophila C1 (central nerve cell) (e), violin plot showing the scale normalized expression levels of representative markers for each subcluster (f).
Extended Data Fig. 3 Earthworm cell landscape was constructed using Microwell-seq.
(a) t-SNE visualization of 95,020 single cells from whole bodies of earthworm, colored by cell type (left) and cell lineage (right). (b) Heatmap showing the scaled average expression levels of earthworm cell type-specific marker genes (left), and relative gene expression of representative cell type-specific markers for each cluster overlaid on t-SNE plots (right). (c) The hierarchical clustering tree (right) showing the similarity among earthworm 62 cells, and the histogram plot (left) showing the subtypes of each cell type. The similarity refers to the AUROC score from MetaNeighbor analysis. (d-e) t-SNE showing 11 subclusters from earthworm C24 (neuron) (d), violin plot showing the scale normalized expression levels of representative markers for each subcluster (e).
Extended Data Fig. 4 Characteristics of immune-related structure cells in zebrafish, Drosophila, and earthworm.
(a) Ridge plots showing the scale normalized expression levels of known zebrafish epithelial markers and immune-related markers in zebrafish epithelial cells. (b) Ridge plots showing the the scale normalized expression levels of known Drosophila MT markers and immune-related markers in Drosophila MT cells (C34 and C69). (c) Ridge plots showing the scale normalized expression levels of two earthworm immune-related markers, evm.TU.Chr04.2499 and evm.TU.ctg2984.2 in earthworm coelomocytes (earthworm C0, C5, C6, C16, C19, C30, C32, C35, and C37). (d-e) t-SNE visualization of 11 subtypes of MT cells from Drosophila C34 and C69, and violin plot showing the scale normalized expression levels of representative markers for each subtype (e).
Extended Data Fig. 5 Cross-species analysis among eight species.
(a) Sankey diagrams showing homologous cell-type pairs between human and mouse obtained from SAMap analyses based on different datasets. (b) The Venn diagram showing the number of overlapping homologous cell-type pairs obtained from SAMap analyses based on different datasets. (c-e) The information of homologous cell types among human, mouse, zebrafish, Ciona, Drosophila, earthworm, C. elegans, and planarian, including the number of homologous cell-type pairs (c), aligned score (d), and number of enriched gene pairs (e). Boxplots represent the median, 25th percentile, and 75th percentile, and whiskers correspond to 1.5 times the interquartile range. N of cell types: human: 434, mouse: 432, zebrafish: 378, Ciona: 291, Drosophila: 288, earthworm: 110, C. elegans: 321, and planarians: 173.
Extended Data Fig. 6 Screening conserved regulators underlying cell lineages.
(a) The bar chart showing the percentage of cell lineage-specific TFs in humans, mice, zebrafish, Ciona, Drosophila, earthworm, C. elegans, and planarians across different conservative levels based on homologous genes (left) obtained from SAMap and 1-to-1 orthologous genes (right). The group number refers to the conservative level. TFs were divided into eight different conservative levels (Level 1–8) based on the conversion of homologous TFs among eight species. Level 1 means that TFs have no homologous genes in other species, and Level 8 means that TFs have homologous genes in all other seven species. (b) The bar chart showing the conserved neuron-related TFs between human and other species. The blue bar refers to the number of homologous TF pairs between human and other species, the yellow bar refers to the number of human neuron-related TFs involved in homologous gene pairs, and the grey bar refers to the number of other species neuron-related TFs involved in homologous gene pairs. (c) Conserved neuron-related regulators across eight species showing a comprehensive homologous link from lower to higher organisms. (d) Sankey diagram showing homologous relationships among vertebrates’ immune-related TFs. (e) Sankey diagram showing homologous relationships of immune-related TFs between Drosophila and human.
Extended Data Fig. 7 The performance of model training for eight species.
(a) The AUROC values of major cell lineages for humans, mice, zebrafish, Ciona, Drosophila, earthworm, C. elegans, and planarians. (b) Boxplots of AUPR values for eight species. Boxplots represent the median, 25th percentile, and 75th percentile, and whiskers correspond to 1.5 times the interquartile range. N of cells: humans: 134,557, mice: 179,344, zebrafish: 241,233, Ciona: 12,489, Drosophila: 77,337, earthworm: 29,609, C. elegans: 30,515, and planarians: 50,562.
Extended Data Fig. 8 Benchmark of prediction performance.
(a) The AUROC values of Nvwa, Basset, DeepSEA, Beluga, Basenji, SVM, random labels and random features on human (n = 134,557) and Drosophila (n = 77,337) specific datasets. (b) The AUROC of multiple genome training for zebrafish (n = 241,233) and C. elegans (n = 30,515). Boxplots represent the median, 25th percentile, and 75th percentile, and whiskers correspond to 1.5 times the interquartile range. (c) The heatmaps showing the correlation between observed and Nvwa-predicted cell type-specific transcription for eight species.
Extended Data Fig. 9 Overview of sequence patterns recognized by Nvwa.
(a) Mean saliency scores show transcriptional start site and important information-rich region recognized by Nvwa. (b) Comparison of the first-layer convolution filters derived from feature map-based approaches and gradient-based TF-MoDISco on Drosophila-specific model. (c) Examples of known TFBS compared with the PWMs of Nvwa first-layer in humans, mice, zebrafish, Ciona, Drosophila, C. elegans, and planarians.
Extended Data Fig. 10 Overview of sequence motifs and their influence recognized by Nvwa.
(a) Volcano plot of Nvwa first-layer filters for humans, mice, zebrafish, Ciona, Drosophila, earthworm, C. elegans, and planarians. The x-axis represents the information contents (IC) of a Filter, the y-axis represents the overall influence on of a Filter, Filters with high influence are tagged as up, and Filters with low influence are tagged as down. Those reductant Filters are tagged as triangle and non-reductant Filters are tagged as dots, the size of elements represents the reproducibility in each independent cross-validation run. (b) Barplot of the Nvwa and single-cell ATAC cell type specific motifs for mouse. Neuronal cells (C12, n=29 and C5, n=169 for Nvwa and sci-ATAC data respectively) and endothelial cells (C50, n=31 and C22, n=136 for Nvwa and sci-ATAC data respectively) were shown. Hit indicates the same motifs, NotHit indicates the different motifs identified by TomTom. X-axis indicates the percentage. (c) Barplot of the Nvwa and single -cell ATAC cell type specific transcription factor for Drosophila. Neuronal cells (C1, n=293, CB, n=639 and OL, n=484 for Nvwa and Flybrain (GSE163697) data respectively) were shown. Hit indicates the same TFs, and NotHit indicates the different TFs annotated by TomTom. X-axis indicates the percentage.
Supplementary information
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, J., Wang, J., Zhang, P. et al. Deep learning of cross-species single-cell landscapes identifies conserved regulatory programs underlying cell types. Nat Genet 54, 1711–1720 (2022). https://doi.org/10.1038/s41588-022-01197-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-022-01197-7
This article is cited by
-
Discovery of an unconventional lamprey lymphocyte lineage highlights divergent features in vertebrate adaptive immune system evolution
Nature Communications (2024)
-
Delineating mouse β-cell identity during lifetime and in diabetes with a single cell atlas
Nature Metabolism (2023)
-
Inflammation and aging: signaling pathways and intervention therapies
Signal Transduction and Targeted Therapy (2023)
-
Convergent differentiation of multiciliated cells
Scientific Reports (2023)