Abstract
The mechanisms by which the fetal-type β-globin-like genes HBG1 and HBG2 are silenced in adult erythroid precursor cells remain a fundamental question in human biology and have therapeutic relevance to sickle cell disease and β-thalassemia. Here, we identify via a CRISPR–Cas9 genetic screen two members of the NFI transcription factor family—NFIA and NFIX—as HBG1/2 repressors. NFIA and NFIX are expressed at elevated levels in adult erythroid cells compared with fetal cells, and function cooperatively to repress HBG1/2 in cultured cells and in human-to-mouse xenotransplants. Genomic profiling, genome editing and DNA binding assays demonstrate that the potent concerted activity of NFIA and NFIX is explained in part by their ability to stimulate the expression of BCL11A, a known silencer of the HBG1/2 genes, and in part by directly repressing the HBG1/2 genes. Thus, NFI factors emerge as versatile regulators of the fetal-to-adult switch in β-globin production.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All RNA-seq, CUT&RUN and ATAC-seq data generated in this study have been deposited in the NIH Gene Expression Omnibus (GEO) with the accession code GSE180871. Gene abundances in fetal and adult erythroid cells were obtained from GSE90878 (ref. 24). Gene sets used for GSEA analysis are provided in this article. HUDEP2 CRISPR screen results were obtained from Supplementary Table 1 of a published study9. All unprocessed immunoblot scanned images and uncropped gel images are included in the Source data provided with this paper.
Code availability
All codes processing sequencing data were obtained from previous established pipelines and are provided in this article.
Change history
01 June 2022
A Correction to this paper has been published: https://doi.org/10.1038/s41588-022-01112-0
References
Orkin, S. H. Molecular medicine: found in translation. Med (N. Y.) 2, 122–136 (2021).
Menzel, S. et al. A QTL influencing F cell production maps to a gene encoding a zinc-finger protein on chromosome 2p15. Nat. Genet. 39, 1197–1199 (2007).
Uda, M. et al. Genome-wide association study shows BCL11A associated with persistent fetal hemoglobin and amelioration of the phenotype of β-thalassemia. Proc. Natl Acad. Sci. USA 105, 1620–1625 (2008).
Sankaran, V. G. et al. Human fetal hemoglobin expression is regulated by the developmental stage-specific repressor BCL11A. Science 322, 1839–1842 (2008).
Masuda, T. et al. Gene regulation: transcription factors LRF and BCL11A independently repress expression of fetal hemoglobin. Science 351, 285–289 (2016).
Liu, N. et al. Direct promoter repression by BCL11A controls the fetal to adult hemoglobin switch. Cell 173, 430–442.e17 (2018).
Martyn, G. E. et al. Natural regulatory mutations elevate the fetal globin gene via disruption of BCL11A or ZBTB7A binding. Nat. Genet. 50, 498–503 (2018).
Xu, J. et al. Corepressor-dependent silencing of fetal hemoglobin expression by BCL11A. Proc. Natl Acad. Sci. USA 110, 6518–6523 (2013).
Huang, P. et al. The HRI-regulated transcription factor ATF4 activates BCL11A transcription to silence fetal hemoglobin expression. Blood 135, 2121–2132 (2020).
Lan, X. et al. ZNF410 uniquely activates the NuRD component CHD4 to silence fetal hemoglobin expression. Mol. Cell 81, 239–254.e8 (2021).
Vinjamur, D. S. et al. ZNF410 represses fetal globin by singular control of CHD4. Nat. Genet. 53, 719–728 (2021).
Liu, N. et al. Transcription factor competition at the γ-globin promoters controls hemoglobin switching. Nat. Genet. 53, 511–520 (2021).
Sher, F. et al. Rational targeting of a NuRD subcomplex guided by comprehensive in situ mutagenesis. Nat. Genet. 51, 1149–1159 (2019).
Zhou, D., Liu, K., Sun, C. W., Pawlik, K. M. & Townes, T. M. KLF1 regulates BCL11A expression and γ- to β-globin gene switching. Nat. Genet. 42, 742–744 (2010).
Tanabe, O. et al. An embryonic/fetal β-type globin gene repressor contains a nuclear receptor TR2/TR4 heterodimer. EMBO J. 21, 3434–3442 (2002).
Rupon, J. W., Wang, S. Z., Gaensler, K., Lloyd, J. & Ginder, G. D. Methyl binding domain protein 2 mediates γ-globin gene silencing in adult human βYAC transgenic mice. Proc. Natl Acad. Sci. USA 103, 6617–6622 (2006).
Krivega, I. & Dean, A. Chromatin looping as a target for altering erythroid gene expression. Ann. N. Y. Acad. Sci. 1368, 31–39 (2016).
Renneville, A. et al. EHMT1 and EHMT2 inhibition induces fetal hemoglobin expression. Blood 126, 1930–1939 (2015).
Macari, E. R. & Lowrey, C. H. Induction of human fetal hemoglobin via the NRF2 antioxidant response signaling pathway. Blood 117, 5987–5997 (2011).
Doerfler, P. A. et al. Activation of γ-globin gene expression by GATA1 and NF-Y in hereditary persistence of fetal hemoglobin. Nat. Genet. 53, 1177–1186 (2021).
Kurita, R., Suda, N., Sudo, K., Miharada, K. & Hiroyama, T. Establishment of immortalized human erythroid progenitor cell lines able to produce enucleated red blood cells. PLoS ONE 8, e59890 (2013).
Gronostajski, R. M. Roles of the NFI/CTF gene family in transcription and development. Gene 249, 31–45 (2000).
Danjou, F. et al. Genome-wide association analyses based on whole-genome sequencing in Sardinia provide insights into regulation of hemoglobin levels. Nat. Genet. 47, 1264–1271 (2015).
Lessard, S., Beaudoin, M., Orkin, S. H., Bauer, D. E. & Lettre, G. 14q32 and let-7 microRNAs regulate transcriptional networks in fetal and adult human erythroblasts. Hum. Mol. Genet. 27, 1411–1420 (2018).
Huang, P. et al. Comparative analysis of three-dimensional chromosomal architecture identifies a novel fetal hemoglobin regulatory element. Genes Dev. 31, 1704–1713 (2017).
Gier, R. A. et al. High-performance CRISPR-Cas12a genome editing for combinatorial genetic screening. Nat. Commun. 11, 3455 (2020).
Bauer, D. E. et al. A erythroid enhancer of BCL11A subject to genetic variation. Science 342, 253–257 (2013).
Grevet, J. D. et al. Domain-focused CRISPR screen identifies HRI as a fetal hemoglobin regulator in human erythroid cells. Science 361, 285–290 (2018).
Starnes, L. M. et al. NFI-A directs the fate of hematopoietic progenitors to the erythroid or granulocytic lineage and controls β-globin and G-CSF receptor expression. Blood 114, 1753–1763 (2009).
McIntosh, B. E. et al. Nonirradiated NOD,B6.SCID Il2rγ−/− KitW41/W41 (NBSGW) mice support multilineage engraftment of human hematopoietic cells. Stem Cell Rep. 4, 171–180 (2015).
Métais, J. Y. et al. Genome editing of HBG1 and HBG2 to induce fetal hemoglobin. Blood Adv. 3, 3379–3392 (2019).
Holmfeldt, P. et al. Nfix is a novel regulator of murine hematopoietic stem and progenitor cell survival. Blood 122, 2987–2996 (2013).
Hall, T. et al. Nfix promotes survival of immature hematopoietic cells via regulation of c-Mpl. Stem Cells 36, 943–950 (2018).
Tangprasittipap, A. et al. Comparison of gene expression profiles between human erythroid cells derived from fetal liver and adult peripheral blood. PeerJ 2018, e5527 (2018).
Skene, P. J. & Henikoff, S. An efficient targeted nuclease strategy for high-resolution mapping of DNA binding sites. eLife 6, e21856 (2017).
Zhu, Q., Liu, N., Orkin, S. & Yuan, G.-C. CUT&RUNTools: a flexible pipeline for CUT&RUN processing and footprint analysis. Genome Biol. 20, 192 (2019).
Piper, M., Gronostajski, R. & Messina, G. Nuclear factor one X in development and disease. Trends Cell Biol. 29, 20–30 (2019).
Klemm, S. L., Shipony, Z. & Greenleaf, W. J. Chromatin accessibility and the regulatory epigenome. Nat. Rev. Genet. 20, 207–220 (2019).
Buenrostro, J., Wu, B., Chang, H. & Greenleaf, W. ATAC-seq: a method for assaying chromatin accessability genome-wide. Curr. Protoc. Mol. Biol. 109, 21.29.1–21.29.9 (2015).
Davis, C. A. et al. The Encyclopedia of DNA Elements (ENCODE): data portal update. Nucleic Acids Res. 46, D794–D801 (2018).
Venkataraman, A. et al. A toolbox of immunoprecipitation-grade monoclonal antibodies to human transcription factors. Nat. Methods 15, 330–338 (2018).
Shah, M., Funnell, A. P. W., Quinlan, K. G. R. & Crossley, M. Hit and run transcriptional repressors are difficult to catch in the act. BioEssays 41, e1900041 (2019).
Xu, J. et al. Correction of sickle cell disease in adult mice by interference with fetal hemoglobin silencing. Science 334, 993–996 (2011).
Jawaid, K., Wahlberg, K., Thein, S. L. & Best, S. Binding patterns of BCL11A in the globin and GATA1 loci and characterization of the BCL11A fetal hemoglobin locus. Blood Cells Mol. Dis. 45, 140–146 (2010).
Xu, J. et al. Transcriptional silencing of γ-globin by BCL11A involves long-range interactions and cooperation with SOX6. Genes Dev. 24, 783–789 (2010).
Khandros, E. et al. Understanding heterogeneity of fetal hemoglobin induction through comparative analysis of F and A erythroblasts. Blood 135, 1957–1968 (2020).
Meers, M. P., Bryson, T. D., Henikoff, J. G. & Henikoff, S. Improved CUT&RUN chromatin profiling tools. eLife 8, e46314 (2019).
Huang, J. et al. Dynamic control of enhancer repertoires drives lineage and stage-specific transcription during hematopoiesis. Dev. Cell 36, 9–23 (2016).
Hiraike, Y. et al. NFIA differentially controls adipogenic and myogenic gene program through distinct pathways to ensure brown and beige adipocyte differentiation. PLoS Genet. 16, e1009044 (2020).
Messina, G. et al. Nfix regulates fetal-specific transcription in developing skeletal muscle. Cell 140, 554–566 (2010).
Taglietti, V. et al. Nfix induces a switch in Sox6 transcriptional activity to regulate MyHC-I expression in fetal muscle. Cell Rep. 17, 2354–2366 (2016).
Yi, Z. et al. Sox6 directly silences epsilon globin expression in definitive erythropoiesis. PLoS Genet. 2, 129–139 (2006).
Quevedo, M. et al. Mediator complex interaction partners organize the transcriptional network that defines neural stem cells. Nat. Commun. 10, 2669 (2019).
Lan, X. et al. The E3 ligase adaptor molecule SPOP regulates fetal hemoglobin levels in adult erythroid cells. Blood Adv. 3, 1586–1597 (2019).
Peslak, S. A. et al. HRI depletion cooperates with pharmacologic inducers to elevate fetal hemoglobin and reduce sickle cell formation. Blood Adv. 4, 4560–4572 (2020).
Bloh, K. et al. Deconvolution of Complex DNA Repair (DECODR): establishing a novel deconvolution algorithm for comprehensive analysis of CRISPR-edited Sanger sequencing data. CRISPR J. 4, 120–131 (2021).
Pinello, L. et al. Analyzing CRISPR genome-editing experiments with CRISPResso. Nat. Biotechnol. 34, 695–697 (2016).
Qin, K. et al. SIRT6-mediated transcriptional suppression of Txnip is critical for pancreatic beta cell function and survival in mice. Diabetologia 61, 906–918 (2018).
Slomiany, B. A., Kelly, M. M. & Kurtz, D. T. Extraction of nuclear proteins with increased DNA binding activity. Biotechniques 28, 938–942 (2000).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Li, B. & Dewey, C. N. RSEM: accurate transcript quantification from RNA-seq data with or without a reference genome. BMC Bioinformatics 12, 323 (2011).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Wu, T. et al. clusterProfiler 4.0: a universal enrichment tool for interpreting omics data. Innovation 2, 100141 (2021).
Yu, F., Sankaran, V. G. & Yuan, G.-C. CUT&RUNTools 2.0: a pipeline for single-cell and bulk-level CUT&RUN and CUT&Tag data analysis. Bioinformatics 38, 252–254 (2021).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Danecek, P. et al. Twelve years of SAMtools and BCFtools. Gigascience 10, giab008 (2021).
Ramírez, F. et al. deepTools2: a next generation web server for deep-sequencing data analysis. Nucleic Acids Res. 44, W160–W165 (2016).
Zhang, Y. et al. Model-based Analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2009).
Acknowledgements
We thank the staff of CHOP Flow Cytometry Core Facility for assistance with cell sorting, and members of the Blobel laboratory for helpful discussions. HUDEP2 cells were a gift from R. Kurita and Y. Nakamura (RIKEN BioResource Center). The Fred Hutchinson Cancer Research Center Cooperative Center of Excellence in Hematology was supported by NIDDK Grant no. DK106829. This work was supported by NIH grants from the National Heart, Lung, and Blood Institute (grant no. HL119479) and research funding from Pfizer (G.A.B.); the National Institute of Diabetes and Digestive and Kidney Diseases (grant no. R24DK106766) (G.A.B., M.J.W. and R.C.H.); grant no. P01HL053749 (to M.J.W.), grant no. R01HL156647 (to M.J.W.), the Assisi Foundation of Memphis (to M.J.W.); the Doris Duke Charitable Foundation grant no. 2017093 (to M.J.W.); grant no. R01HL147879 (to S.T.C.); a K08 training grant (no. K08-DK129716) and the Doris Duke Charitable Foundation Physician Scientist Fellowship grant no. 2020062 (S.A.P.); a T32 training grant (no. HL007150-42) and an American Society of Hematology Research Training Award for Fellows (E.K.); NIDDK grant no. F32DK118822 and Cooley’s Anemia Foundation (to P.A.D.); and the St. Jude Children’s Research Hospital Collaborative Research Consortium on Novel Gene Therapies for Sickle Cell Disease. We thank the DiGaetano family for their generous support.
Author information
Authors and Affiliations
Contributions
P.H. and G.A.B. conceived the project. K.Q., P.H., R.F., C.A.K., S.A.P., E.K., M.S., X.L., T.M., B.G. and O.A. conducted experiments. P.A.D. generated the HUDEP2Δεγδβ/GγΑγ cell line. S.T.C. and J.S. provided reagents. R.C.H., M.J.W. and G.A.B. acquired funding and supervised the project. K.Q., P.H. and G.A.B. designed the experiments and performed data analyses. K.Q. and G.A.B. wrote the manuscript with input from all authors. All authors read and approved the contents of this manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Genetics thanks John Strouboulis and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Co-depletion of NFIA and NFIX reactivates γ-globin in HUDEP2 cells.
a, Diagram depicting the structure of NFI gene products. b, Schematic of multiplex AsCas12a sgRNAs. c, Editing efficiency of NFIA, NFIX, NFIC, and BCL11A +58 intronic enhancer in indicated sgRNA infected HUDEP2 cells by AsCas12a (control, n = 4; sgNFIA, n = 4; sgNFIX, n = 4, sgNFIC, n = 2; sgNFIA&X, n = 3; sgNFIA&X&C, n = 4; sgBCL11A +58, n = 2). n represents biological replicates generated from different experiments. Genomic DNAs were obtained from indicated 5 days post-infected HUDEP2 cells. d, Representative gating strategy and HbF staining results in control and indicated Cas12a sgRNA-infected HUDEP2 cells. The first gate selects the live cells in the population. The second gate distinguishes the HbF+ and HbF– population. Experiments were performed twice with similar results. e–h, HUDEP2 cells expressing CRISPR-Cas9 were infected with pLRG2.1 or pLRCherry2.1 lentivirus carrying control sgRNA or sgRNAs targeting indicated NFI genes or the BCL11A +58 enhancer and analyzed at the end of 5 days differentiation. e, Representative immunoblots of NFIA, NFIX, and γ-globin. β-Actin was used as loading control. Experiments were performed three times with similar results. f–h, RT-qPCR quantification of HBG1/2, HBB, and the ratio of HBG/(HBG + HBB) mRNA. Data were normalized to GAPDH (n = 3) and expressed as means ± SEM. *p < 0.05, **p < 0.01. p values were calculated by comparing indicated samples to control using parametric paired two-tailed Student’s t test. f, sgNFIA #1, p = 0.0422; sgNFIA #2, p = 0.0161; sgNFIX #1, p = 0.0644; sgNFIX #2, p = 0.4230; sgNFIA&NFIX #1, p = 0.0154; sgNFIA&NFIX #2, p = 0.0150; sgBCL11A +58, p = 0.0288. g, sgNFIA #1, p = 0.0033; sgNFIA #2, p = 0.0052; sgNFIX #1, p = 0.0005; sgNFIX #2, p = 0.0265; sgNFIA&NFIX #1, p = 0.0007; sgNFIA&NFIX #2, p = 0.0024; sgBCL11A +58, p = 0.0009.
Extended Data Fig. 2 Co-depletion of NFIA and NFIX reactivates γ-globin in primary erythroblasts derived from healthy donors.
a, Three-phase in vitro culture system for human CD34+ HSPCs to differentiate into mature red cells. Cas9 and indicated NFI sgRNA RNPs were transfected into CD34+ HSPCs on day 4 of culture by electroporation. Erythroblasts were harvested and analyzed by RT-qPCR, immunoblot, HbF staining, and HPLC on day 13 or 15 of culture. b, Representative immunoblots of NFIA, NFIX, and γ-globin from untargeted primary cells at indicated days of differentiation. Experiments were performed on two donors with similar results. c, Representative HbF staining result from indicated gene edited primary erythroblasts at day 15 of differentiation. d, Representative CD71 and CD235a flow cytometry of primary erythroblasts at day 15 of differentiation. e, Representative Wright-Giemsa stains of primary erythroblasts at day 15 of differentiation. Experiments in c–e were performed on three donors/biological replicates with similar results. Scale bar, 25 μm.
Extended Data Fig. 3 Co-depletion of NFIA and NFIX reactivates γ-globin in primary erythroblasts derived from SCD patients.
a, Procedure for depleting NFIA and/or NFIX in SCD CD34+ HSPCs in primary culture (n = 1). Isolated CD34+ HSPCs were transfected by indicated RNPs on day 4 of culture. On day 13, a subset of cells was harvested for RT-qPCR analysis. On day 15, a subset of cells was harvested for immunoblots and HbF flow cytometry analysis. On day 21, the remaining cells were collected for low O2 sickling assay. b, Immunoblots of NFIA, NFIX, γ-globin, and β-globin (n = 1 donor). c, The ratio of HBG/(HBG + HBB) as determined by RT-qPCR (n = 1 donor). d, HbF+ fraction (%) as determined by HbF flow cytometry (n = 1 donor). e, Representative micrographs of sickle cells, and quantification of the percentage of sickle cells (n = 1 donor). Scale bar, 25 μm.
Extended Data Fig. 4 Co-depletion of NFIA and NFIX strongly reactivates γ-globin in human-to-mouse xenotransplants.
a, Percentage of human CD45+ cells from the bone marrow of the transplanted NBSGW mice (n = 5 mice/biological replicates). b, Composition of myeloid, B cells, and HSPCs from CD45+ sorted population and percentage of erythroid cells from hCD45- cells (n = 5 mice/ biological replicates). c–e, Indel analysis of NFIA, NFIX, and BCL11A +58 enhancer in input HSPCs, total engrafted cells, B cells, T cells, and HSPCs isolated from indicated recipient animals 16 weeks post-transplantation (n = 5 mice/ biological replicates). Data are expressed as means ± SEM.
Extended Data Fig. 5 NFIA and NFIX support an adult-type erythroblast transcription program.
a, Clustering analysis of 430 NFIA and NFIX co-depletion affected genes in HUDEP2 cells. A total of four distinct expression patterns are identified. Boxplot showing expression levels of genes in Group 1 (n = 234) and Group 2 (n = 174) display a correlation with the dosage of total NFI proteins. Y-axes represents z-scores of gene abundance. Lower whisker, smallest observation greater than or equal to lower hinge - 1.5 x Interquartile range (IQR); lower hinge, 25% quantile; median, 50% quantile; upper hinge, 75% quantile; upper whisker, largest observation less than or equal to upper hinge + 1.5 x IQR. b–d, Scatter plot showing the comparison of adult globin genes (HBA1/2, HBB, and HBD) and red cell differentiation-related genes (GATA1, ALAS2, and BAND3) in representative control, NFIA and NFIX single or co-depleted (sgNFIA&NFIX) HUDEP2 cells.
Extended Data Fig. 6 Characterization of NFIA and NFIX genomic occupancy profiles by CUT&RUN.
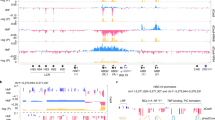
a, The statistics of NFIA and NFIX peaks identified from HUDEP2 and primary erythroblast CUT&RUN experiments. b, Genomic features of NFIA and NFIX CUT&RUN peaks. c, Distribution of NFIA and NFIX-binding loci relative to gene transcription start site. d, Genomic occupancies of NFIA and NFIX at β-globin locus in HUDEP2 and primary erythroblasts. NFIA KO (sgNFIA), NFIX KO (sgNFIX), and IgG were used as controls. e, CUT&RUN signals at the BCL11A locus with three indicated NFIA antibodies in primary cells at day 10 of differentiation.
Extended Data Fig. 7 NFI factors support BCL11A gene expression in adult erythroid cells.
a, Venn diagrams showing statistics of ATAC-seq peaks (chromatin open regions) in control and NFIA and NFIX co-depleted (sgNFIA&NFIX) replicate samples. b, ATAC-seq tracks at the β-globin locus in control and NFIA and NFIX co-depleted (sgNFIA&NFIX) HUDEP2 cells.
Extended Data Fig. 8 NFI factors directly silence HBG1/2 genes.
a, NFIA and NFIC ChIP-seq signals at the β-globin locus in K562 cells. NFIA ChIP-seq data were downloaded from GEO (GSM2574788) and NFIC ChIP-seq data were downloaded from ENCODE (ENCSR796ITY). b–c, Footprint analysis of NFIA CUT&RUN data at NFI binding sites that were protected from pAG-MNase cut in CUT&RUN assays. The location of site 1 is approximately 450 bp upstream of HBG1/2 TSS (transcription start site), whereas site 2 is in HBG1/2 first intron, 200 bp downstream of HBG1/2 TSS. Data were aggregates from three replicates of NFIA CUT&RUN-sequencing results of HUDEP2 clone 6 cells. d, Footprint analysis of NFIA CUT&RUN data at the HBG1/2 TSS-proximal region. Boxes denote three putative NFI binding sites that were not protected from the pAG-MNase cut in CUT&RUN assays. The BCL11A and NFYA binding sites are underlined3. Data were aggregates from three replicates of NFIA HUDEP2 clone 6 CUT&RUN. e–g, Electrophoretic mobility shift assay (EMSA) examining the binding of NFIA (e) and NFIX (f-g) with probes containing identified NFI binding sites sequence. In all experiments, the interaction of NFIA/NFIX and IRDye 700 labeled probes produced a gel shift which was competed with unlabeled cold probes in 50 (50x) or 200 (200x) times excess molar concentration, including a random sequence (random), wild type (WT), or two mutant sequences (Left mut or Right mut) containing either left or right half of the NFI full motif (TGGA or GCCT for site 1 and TGGC or TCCA for site 2). The identity of the NFIA/X-DNA complex was verified with an anti-AM tag antibody that recognized the recombinant NFIA or NFIX protein and generated a ‘supershift’. Left mut: probes contain four mutated sequences at the left half of the NFI motif sequences; Right mut: probes contain right mutated sequences at the left half of the NFI motif sequences.
Extended Data Fig. 9 Two putative NFI motifs are required for NFI binding in HBG1/2 genes.
a, HUDEP2 clone 6 and K562 cells expressing CRISPR-Cas9 were infected with sgRNAs targeting NFI binding site 1 (NFI –451) and site 2 (NFI +202). NFIA CUT&RUN tracks at the HBG1/2 gene locus in control and NFI binding site 1 (NFI –451) and site 2 (NFI +202) edited HUDEP2 clone 6 and K562 cells. b, HUDEP2Δεγδβ/GγΑγ cells expressing CRISPR-Cas9 were infected with sgRNAs targeting NFI binding site 1 (NFI –451) and site 2 (NFI +202). Genomic DNA of control and edited cells were isolated and subject to PCR and sanger sequencing analysis (n = 2 biological replicates, generated from independent experiments). The arrows denote the sgRNAs target sites.
Supplementary information
Supplementary Information
Supplementary Tables 6–11.
Supplementar Tables
Supplementary Tables 1–5.
Source data
Source Data Fig. 1
Unprocessed western blots.
Source Data Fig. 2
Unprocessed western blots.
Source Data Fig. 5
Unprocessed western blots.
Source Data Fig. 6
Unprocessed EMSA gels.
Source Data Extended Data Fig. 1
Unprocessed western blots.
Source Data Extended Data Fig. 2
Unprocessed western blots.
Source Data Extended Data Fig. 3
Unprocessed western blots.
Source Data Extended Data Fig. 8
Unprocessed EMSA gels.
Rights and permissions
About this article
Cite this article
Qin, K., Huang, P., Feng, R. et al. Dual function NFI factors control fetal hemoglobin silencing in adult erythroid cells. Nat Genet 54, 874–884 (2022). https://doi.org/10.1038/s41588-022-01076-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-022-01076-1
This article is cited by
-
Efficient engineering of human and mouse primary cells using peptide-assisted genome editing
Nature Biotechnology (2024)
-
Erythroid lineage chromatin accessibility maps facilitate identification and validation of NFIX as a fetal hemoglobin repressor
Communications Biology (2023)
-
A regulatory network controlling ovarian granulosa cell death
Cell Death Discovery (2023)
-
CRISPR/Cas-based gene editing in therapeutic strategies for beta-thalassemia
Human Genetics (2023)
-
HIC2 controls developmental hemoglobin switching by repressing BCL11A transcription
Nature Genetics (2022)