Abstract
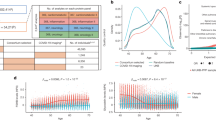
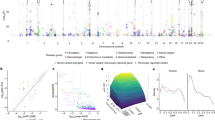
The human proteome is a major source of therapeutic targets. Recent genetic association analyses of the plasma proteome enable systematic evaluation of the causal consequences of variation in plasma protein levels. Here we estimated the effects of 1,002 proteins on 225 phenotypes using two-sample Mendelian randomization (MR) and colocalization. Of 413 associations supported by evidence from MR, 130 (31.5%) were not supported by results of colocalization analyses, suggesting that genetic confounding due to linkage disequilibrium is widespread in naïve phenome-wide association studies of proteins. Combining MR and colocalization evidence in cis-only analyses, we identified 111 putatively causal effects between 65 proteins and 52 disease-related phenotypes (https://www.epigraphdb.org/pqtl/). Evaluation of data from historic drug development programs showed that target-indication pairs with MR and colocalization support were more likely to be approved, evidencing the value of this approach in identifying and prioritizing potential therapeutic targets.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The data (GWAS summary statistics) used in the analyses described here are freely accessible in the MR-Base platform (https://www.mrbase.org/). All our analysis results for 989 proteins against 225 human phenotypes are freely available to browse, query and download in EpiGraphDB (https://www.epigraphdb.org/pqtl/). An application programming interface and R package documented on the website enable users to programmatically access data from the database.
Code availability
The code used in the MR and colocalization analyses described here are freely accessible via our GitHub repository (https://github.com/MRCIEU/epigraphdb-pqtl/). The MR analysis was conducted using TwoSampleMR R package (https://github.com/MRCIEU/TwoSampleMR/). We implemented the colocalization analysis using the coloc R package (created by C. Wallace and colleagues), which can be downloaded at https://cran.r-project.org/web/packages/coloc/index.html/.
References
Plenge, R. M., Scolnick, E. M. & Altshuler, D. Validating therapeutic targets through human genetics. Nat. Rev. Drug Discov. 12, 581–594 (2013).
Hay, M., Thomas, D. W., Craighead, J. L., Economides, C. & Rosenthal, J. Clinical development success rates for investigational drugs. Nat. Biotechnol. 32, 40–51 (2014).
Arrowsmith, J. & Miller, P. Phase II and Phase III attrition rates 2011–2012. Nat. Rev. Drug Discov. 12, 569 (2013).
Harrison, R. K. Phase II and phase III failures: 2013–2015. Nat. Rev. Drug Discov. 15, 817–818 (2016).
Cummings, J. L., Morstorf, T. & Zhong, K. Alzheimer’s disease drug-development pipeline: few candidates, frequent failures. Alzheimers Res. Ther. 6, 37 (2014).
Nelson, M. R. et al. The support of human genetic evidence for approved drug indications. Nat. Genet. 47, 856–860 (2015).
Zhu, Z. et al. Integration of summary data from GWAS and eQTL studies predicts complex trait gene targets. Nat. Genet. 48, 481–487 (2016).
Richardson, T. G. et al. Systematic Mendelian randomization framework elucidates hundreds of CpG sites which may mediate the influence of genetic variants on disease. Hum. Mol. Genet. 27, 3293–3304 (2018).
Sun, B. B. et al. Genomic atlas of the human plasma proteome. Nature 558, 73–79 (2018).
Chong, M. et al. Novel drug targets for ischemic stroke identified through Mendelian randomization analysis of the blood proteome. Circulation 140, 819–830 (2019).
Santos, R. et al. A comprehensive map of molecular drug targets. Nat. Rev. Drug Discov. 16, 19–34 (2017).
Imming, P., Sinning, C. & Meyer, A. Drugs, their targets and the nature and number of drug targets. Nat. Rev. Drug Discov. 5, 821–834 (2006).
Suhre, K. et al. Connecting genetic risk to disease end points through the human blood plasma proteome. Nat. Commun. 8, 14357 (2017).
Folkersen, L. et al. Mapping of 79 loci for 83 plasma protein biomarkers in cardiovascular disease. PLoS Genet. 13, e1006706 (2017).
Yao, C. et al. Genome-wide mapping of plasma proteins QTLs identifies putatively causal genes and pathways for cardiovascular disease. Nat. Commun. 9, 3268 (2018).
Emilsson, V. et al. Co-regulatory networks of human serum proteins link genetics to disease. Science 361, 769–773 (2018).
Evans, D. M. & Davey Smith, G. Mendelian randomization: new applications in the coming age of hypothesis-free causality. Annu. Rev. Genomics Hum. Genet. 16, 327–350 (2015).
Millwood, I. Y. et al. Association of CETP gene variants with risk for vascular and nonvascular diseases among Chinese adults. JAMA Cardiol. 3, 34–43 (2018).
Interleukin-6 Receptor Mendelian Randomisation Analysis (IL6R MR) Consortium. The interleukin-6 receptor as a target for prevention of coronary heart disease: a mendelian randomisation analysis. Lancet 379, 1214–1224 (2012).
Swerdlow, D. I. et al. Selecting instruments for Mendelian randomization in the wake of genome-wide association studies. Int. J. Epidemiol. 45, 1600–1616 (2016).
Hemani, G. et al. The MR-Base platform supports systematic causal inference across the human phenome. eLife 7, e34408 (2018).
Timpson, N. J. et al. C-reactive protein levels and body mass index: elucidating direction of causation through reciprocal Mendelian randomization. Int. J. Obes. 35, 300–308 (2011).
Hemani, G., Tilling, K. & Davey Smith, G. Orienting the causal relationship between imprecisely measured traits using GWAS summary data. PLoS Genet. 13, e1007081 (2017).
Hemani, G. et al. Automating Mendelian randomization through machine learning to construct a putative causal map of the human phenome. Preprint at bioRxiv https://doi.org/10.1101/173682 (2017).
Bowden, J. et al. A framework for the investigation of pleiotropy in two-sample summary data Mendelian randomization. Stat. Med. 36, 1783–1802 (2017).
Giambartolomei, C. et al. Bayesian test for colocalisation between pairs of genetic association studies using summary statistics. PLoS Genet. 10, e1004383 (2014).
Yang, J. et al. Conditional and joint multiple-SNP analysis of GWAS summary statistics identifies additional variants influencing complex traits. Nat. Genet. 44, 369–375 (2012).
Burgess, S., Dudbridge, F. & Thompson, S. G. Combining information on multiple instrumental variables in Mendelian randomization: comparison of allele score and summarized data methods. Stat. Med. 35, 1880–1906 (2016).
de Lange, K. M. et al. Genome-wide association study implicates immune activation of multiple integrin genes in inflammatory bowel disease. Nat. Genet. 49, 256–261 (2017).
Solomon, T. et al. Identification of common and rare genetic variation associated with plasma protein levels using whole-exome sequencing and mass spectrometry. Circ. Genom. Precis. Med. 11, e002170 (2018).
Taylor, F. B. Jr, Peer, G. T., Lockhart, M. S., Ferrell, G. & Esmon, C. T. Endothelial cell protein C receptor plays an important role in protein C activation in vivo. Blood 97, 1685–1688 (2001).
Hashizume, M. et al. Tocilizumab, a humanized anti-IL-6R antibody, as an emerging therapeutic option for rheumatoid arthritis: molecular and cellular mechanistic insights. Int. Rev. Immunol. 34, 265–279 (2015).
Ridker, P. M. et al. Modulation of the interleukin-6 signalling pathway and incidence rates of atherosclerotic events and all-cause mortality: analyses from the Canakinumab Anti-Inflammatory Thrombosis Outcomes Study (CANTOS). Eur. Heart J. 39, 3499–3507 (2018).
Ferreira, R. C. et al. Functional IL6R 358Ala allele impairs classical IL-6 receptor signaling and influences risk of diverse inflammatory diseases. PLoS Genet. 9, e1003444 (2013).
Stacey, D. et al. Elucidating mechanisms of genetic cross-disease associations: an integrative approach implicates protein C as a causal pathway in arterial and venous diseases. Preprint at medRxiv https://doi.org/10.1101/2020.03.16.20036822 (2020).
Sanseau, P. et al. Use of genome-wide association studies for drug repositioning. Nat. Biotechnol. 30, 317–320 (2012).
Finan, C. et al. The druggable genome and support for target identification and validation in drug development. Sci. Transl. Med. 9, eaag1166 (2017).
Holmes, M. V., Ala-Korpela, M. & Smith, G. D. Mendelian randomization in cardiometabolic disease: challenges in evaluating causality. Nat. Rev. Cardiol. 14, 577–590 (2017).
Bowden, J., Davey Smith, G. & Burgess, S. Mendelian randomization with invalid instruments: effect estimation and bias detection through Egger regression. Int. J. Epidemiol. 44, 512–525 (2015).
Zhao, Q. Y. et al. Statistical inference in two-sample summary-data Mendelian randomization using robust adjusted profile score. Ann. Statist. 48, 1742–1769 (2020).
Evans, D. M. et al. Mining the human phenome using allelic scores that index biological intermediates. PLoS Genet. 9, e1003919 (2013).
Timpson, N. J. One size fits all: are there standard rules for the use of genetic instruments in Mendelian randomization? Int. J. Epidemiol. 45, 1617–1618 (2016).
Hemani, G., Bowden, J. & Davey Smith, G. Evaluating the potential role of pleiotropy in Mendelian randomization studies. Hum. Mol. Genet. 27, R195–R208 (2018).
Wu, Y. et al. Colocalization of GWAS and eQTL signals at loci with multiple signals identifies additional candidate genes for body fat distribution. Hum. Mol. Genet. 28, 4161–4172 (2019).
Wainberg, M. et al. Opportunities and challenges for transcriptome-wide association studies. Nat. Genet. 51, 592–599 (2019).
Uhlén, M. et al. Tissue-based map of the human proteome. Science 347, 1260419 (2015).
GTEx Consortium. Genetic effects on gene expression across human tissues. Nature 550, 204–213 (2017).
Kim-Hellmuth, S. et al. Genetic regulatory effects modified by immune activation contribute to autoimmune disease associations. Nat. Commun. 8, 266 (2017).
Boyd, A. et al. Cohort Profile: the ‘children of the 90 s’—the index offspring of the Avon Longitudinal Study of Parents and Children. Int. J. Epidemiol. 42, 111–127 (2013).
Fraser, A. et al. Cohort profile: the Avon Longitudinal Study of Parents And Children: ALSPAC mothers cohort. Int. J. Epidemiol. 42, 97–110 (2013).
McLaren, W. et al. The ensembl variant effect predictor. Genome Biol. 17, 122 (2016).
Nyholt, D. R. A simple correction for multiple testing for single-nucleotide polymorphisms in linkage disequilibrium with each other. Am. J. Hum. Genet. 74, 765–769 (2004).
Cichonska, A. et al. MetaCCA: summary statistics-based multivariate meta-analysis of genome-wide association studies using canonical correlation analysis. Bioinformatics 32, 1981–1989 (2016).
Zheng, J. et al. PhenoSpD: an integrated toolkit for phenotypic correlation estimation and multiple testing correction using GWAS summary statistics. Gigascience 7, giy090 (2018).
Lawlor, D. A., Harbord, R. M., Sterne, J. A. C., Timpson, N. & Davey Smith, G. Mendelian randomization: using genes as instruments for making causal inferences in epidemiology. Stat. Med. 27, 1133–1163 (2008).
Burgess, S., Zuber, V., Valdes-Marquez, E., Sun, B. B. & Hopewell, J. C. Mendelian randomization with fine-mapped genetic data: choosing from large numbers of correlated instrumental variables. Genet. Epidemiol. 41, 714–725 (2017).
Yavorska, O. O. & Burgess, S. MendelianRandomization: an R package for performing Mendelian randomization analyses using summarized data. Int. J. Epidemiol. 46, 1734–1739 (2017).
Haycock, P. C. et al. Best (but oft-forgotten) practices: the design, analysis and interpretation of Mendelian randomization studies. Am. J. Clin. Nutr. 103, 965–978 (2016).
Di Angelantonio, E. et al. Efficiency and safety of varying the frequency of whole blood donation (INTERVAL): a randomized trial of 45,000 donors. Lancet 390, 2360–2371 (2017).
Acknowledgements
We are extremely grateful to all the families who took part in the ALSPAC study, the midwives for their help in recruiting them and the whole ALSPAC team, including interviewers, computer and laboratory technicians, clerical workers, research scientists, volunteers, managers, receptionists and nurses. We acknowledge J. Bowden for statistical support and advice relating to MR-Egger regression. This publication is the work of the authors, and J. Zheng will serve as guarantor for the contents of this paper. J.Z. is funded by a Vice-Chancellor’s Fellowship from the University of Bristol. This research was also funded by the UK Medical Research Council Integrative Epidemiology Unit (MC_UU_00011/1 and MC_UU_00011/4), GlaxoSmithKline, Biogen and the Cancer Research Integrative Cancer Epidemiology Programme (C18281/A19169). The UK Medical Research Council and Wellcome (grant no. 102215/2/13/2) and the University of Bristol provided core support for ALSPAC. A comprehensive list of grant funding is available on the ALSPAC website (https://www.bristol.ac.uk/alspac/external/documents/grant-acknowledgements.pdf/). T.R.G. holds a Turing Fellowship with the Alan Turing Institute. G.H. is funded by the Wellcome Trust and the Royal Society (208806/Z/17/Z). M.V.H. is supported by a British Heart Foundation Intermediate Clinical Research Fellowship (FS/18/23/33512) and the National Institute for Health Research (NIHR) Oxford Biomedical Research Centre. This work has been supported by the NIHR Biomedical Research Centre at University Hospitals Bristol and Weston NHS Foundation Trust and the University of Bristol (G.D.S. and T.R.G.). The views expressed are those of the authors and not necessarily those of the NIHR or the Department of Health and Social Care. This work was supported by the Elizabeth Blackwell Institute for Health Research University of Bristol and the Medical Research Council Proximity to Discovery award. P.E. is supported by Cancer Research UK (CRUK; C18281/A19169). S.L. is funded by the Bau Tsu Zung Bau Kwan Yeun Hing Research and Clinical Fellowship (200008682.920006.20006.400.01) from the University of Hong Kong. J.D. is funded by a NIHR Senior Investigator award. J.D. sits on the International Cardiovascular and Metabolic advisory board for Novartis (since 2010), the UK Biobank Steering Committee (since 2011), and is a member of the MRC International Advisory Group (ING) London (since 2013), the MRC High Throughput Science ‘Omics Panel’, London (since 2013), the Scientific Advisory Committee for Sanofi (since 2013), the International Cardiovascular and Metabolism Research and Development Portfolio Committee for Novartis and the AstraZeneca Genomics advisory board (since 2018). P.C.H. is supported by CRUK Population Research Postdoctoral Fellowship C52724/A20138.
Participants in the INTERVAL randomized controlled trial were recruited with the active collaboration of NHS Blood and Transplant England (https://ww.nhsbt.nhs.uk/), which has supported fieldwork and other elements of the trial. DNA extraction and genotyping was co-funded by the NIHR, the NIHR BioResource (https://bioresource.nihr.ac.uk/) and the NIHR Cambridge Biomedical Research Centre at the Cambridge University Hospitals NHS Foundation Trust. The academic coordinating centre for INTERVAL was supported by core funding from the NIHR Blood and Transplant Research Unit in Donor Health and Genomics (NIHR BTRU-2014–10024), UK Medical Research Council (MR/L003120/1), British Heart Foundation (SP/09/002; RG/13/13/30194; RG/18/13/33946) and the NIHR Cambridge Biomedical Research Centre at the Cambridge University Hospitals NHS Foundation Trust. A complete list of the investigators and contributors to the INTERVAL trial is provided in Di Angelantonio et al.59. The academic coordinating centre thank blood donor center staff and blood donors for participating in the INTERVAL trial.
We gratefully acknowledge all studies and databases that have made their GWAS summary data available for this study: arcOGEN (Arthritis Research UK Osteoarthritis Genetics), BCAC (the Breast Cancer Association Consortium), C4D (Coronary Artery Disease Genetics Consortium), CARDIoGRAM (Coronary ARtery DIsease Genome-wide Replication and Meta-analysis), CKDGen (Chronic Kidney Disease Genetics consortium), DIAGRAM (DIAbetes Genetics Replication And Meta-analysis), EAGLE (EArly Genetics and Lifecourse Epidemiology Consortium), EAGLE Eczema (EArly Genetics and Lifecourse Epidemiology Eczema Consortium), EGG (Early Growth Genetics Consortium), ENIGMA (Enhancing Neuro Imaging Genetics through Meta-Analysis), GCAN (Genetic Consortium for Anorexia Nervosa), GEFOS (GEnetic Factors for OSteoporosis Consortium), GIANT (Genetic Investigation of ANthropometric Traits), GIS (Genetics of Iron Status consortium), GLGC (Global Lipids Genetics Consortium), GliomaScan (cohort-based GWAS of glioma), GPC (Genetics of Personality Consortium), GUGC (Global Urate and Gout consortium), HaemGen (hematological and platelet traits genetics consortium), IGAP (International Genomics of Alzheimer’s Project), IIBDGC (International Inflammatory Bowel Disease Genetics Consortium), ILCCO (International Lung Cancer Consortium), IMSGC (International Multiple Sclerosis Genetic Consortium), ISGC (International Stroke Genetics Consortium), MAGIC (Meta-Analyses of Glucose and Insulin-related traits Consortium), MDACC (MD Anderson Cancer Center), MESA (Multi-Ethnic Study of Atherosclerosis), Neale’s lab (a team of researchers from Benjamin Neale’s group, who made the UK Biobank GWAS summary statistics publically available), OCAC (Ovarian Cancer Association Consortium), IPSCSG (the International PSC study group), NHGRI-EBI GWAS catalog (National Human Genome Research Institute and European Bioinformatics Institute Catalog of published GWAS), PanScan (Pancreatic Cancer Cohort Consortium), PGC (Psychiatric Genomics Consortium), Project MinE consortium, ReproGen (Reproductive ageing Genetics consortium), SSGAC (Social Science Genetics Association Consortium), TAG (Tobacco and Genetics Consortium) and the UK Biobank.
J.Z. acknowledges his grandmother ChenZhu for all her support, may she rest in peace.
Author information
Authors and Affiliations
Contributions
J.Z., V.H. and D.B. performed the MR analysis. J.Z. and D.B. performed the colocalization analysis. J.Z. performed the conditional analysis. V.H., Y.L., B.E. and T.R.G. developed the database and web browser. J.Z., V.W. and M.R.H. performed the drug target prioritization and enrichment analysis. J.Z. and R.S. conducted the druggable genome analysis. J.Z. and P.E. conducted the pathway and protein–protein interaction analysis. M.R.H., A.G., T.G.R., B.E., H.M.M., J.Y., C.L., S.L. and J.R. conducted supporting analyses. J.R.S., B.B.S., J.D., H.R. and J.C.M. provided key data and supported the MR analysis. M.R.H., S.B., J.Z.L., K.E., L.M., M.V.H., D.W. and M.R.N. reviewed the paper and provided key comments. J.Z., V.H., D.B., V.W., P.C.H., A.S.B., G.D.S., G.H., R.A.S. and T.R.G. are the members of the Proteome MR writing group and wrote the manuscript. J.Z., T.R.G. and R.A.S. conceived and designed the study and oversaw all analyses.
Corresponding authors
Ethics declarations
Competing interests
A.G., L.M., M.R.H., D.W., M.R.N., R.S. and R.A.S. are employees and shareholders in GlaxoSmithKline. H.R., J.Z.L. and K.E. are employees and shareholders in Biogen. J.Z. and V.H. are employed on a grant funded by GlaxoSmithKline. D.B. is employed on a grant funded by Biogen. T.R.G., G.H. and G.D.S. receive funding from GlaxoSmithKline and Biogen for the work described here. A.S.B. has received grants from Merck, Novartis, Biogen, Pfizer and AstraZeneca. M.V.H. has collaborated with Boehringer Ingelheim in research and, in accordance with the policy of the Clinical Trial Service Unit and Epidemiological Studies Unit (University of Oxford), did not accept any personal payment. This work was supported by Health Data Research UK, which is funded by the UK Medical Research Council, Engineering and Physical Sciences Research Council, Economic and Social Research Council, Department of Health and Social Care (England), Chief Scientist Office of the Scottish Government Health and Social Care Directorates, Health and Social Care Research and Development Division (Welsh Government), Public Health Agency (Northern Ireland), the British Heart Foundation and the Wellcome Trust.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–10 and Supplementary Note
Supplementary Tables
Supplementary Tables 1–27
Supplementary Data 1
Data for bidirectional MR and Steiger filtering results.
Supplementary Data 2
Data for a detailed comparison within each protein group using Venn diagrams.
Rights and permissions
About this article
Cite this article
Zheng, J., Haberland, V., Baird, D. et al. Phenome-wide Mendelian randomization mapping the influence of the plasma proteome on complex diseases. Nat Genet 52, 1122–1131 (2020). https://doi.org/10.1038/s41588-020-0682-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-020-0682-6
This article is cited by
-
Insight into telomere regulation: road to discovery and intervention in plasma drug-protein targets
BMC Genomics (2024)
-
Integrative multi-omics analysis identifies genetically supported druggable targets and immune cell specificity for myasthenia gravis
Journal of Translational Medicine (2024)
-
Clinically relevant plasma proteome for adiposity depots: evidence from systematic mendelian randomization and colocalization analyses
Cardiovascular Diabetology (2024)
-
Systematic proteome-wide Mendelian randomization using the human plasma proteome to identify therapeutic targets for lung adenocarcinoma
Journal of Translational Medicine (2024)
-
Genetic determinants of plasma protein levels in the Estonian population
Scientific Reports (2024)