Abstract
Knowledge of genome-wide genealogies for thousands of individuals would simplify most evolutionary analyses for humans and other species, but has remained computationally infeasible. We have developed a method, Relate, scaling to >10,000 sequences while simultaneously estimating branch lengths, mutational ages and variable historical population sizes, as well as allowing for data errors. Application to 1,000 Genomes Project haplotypes produces joint genealogical histories for 26 human populations. Highly diverged lineages are present in all groups, but most frequent in Africa. Outside Africa, these mainly reflect ancient introgression from groups related to Neanderthals and Denisovans, while African signals instead reflect unknown events unique to that continent. Our approach allows more powerful inferences of natural selection than has previously been possible. We identify multiple regions under strong positive selection, and multi-allelic traits including hair color, body mass index and blood pressure, showing strong evidence of directional selection, varying among human groups.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Relate-estimated coalescence rates, allele ages and selection P values for the 1,000GP can be downloaded from https://zenodo.org/record/3234689. Datasets used in the current study were obtained from the following URLs: 1,000GP phased dataset, https://mathgen.stats.ox.ac.uk/impute/1000GP_Phase3.html (13 January 2017); Genomic mask, ftp://ftp.1000genomes.ebi.ac.uk/vol1/ftp/release/20130502/supporting/accessible_genome_masks/ (20 July 2017); Human ancestral genome, ftp://ftp.1000genomes.ebi.ac.uk/vol1/ftp/phase1/analysis_results/supporting/ancestral_alignments/ (20 July 2017); Altai Neanderthal, http://cdna.eva.mpg.de/neandertal/Vindija/VCF/Altai/ (17 February 2018); Vindija Neanderthal, http://cdna.eva.mpg.de/neandertal/Vindija/VCF/Vindija33.19/ (1 May 2018); Denisovan, http://cdna.eva.mpg.de/neandertal/altai/Denisovan/ (2 March 2018); GWAS catalog, https://www.ebi.ac.uk/gwas/api/search/downloads/full (9 November 2017); PGC GWAS study, https://www.med.unc.edu/pgc/results-and-downloads (23 November 2018); HaploReg, http://archive.broadinstitute.org/mammals/haploreg/data/haploreg_v4.0_20151021.vcf.gz (21 October 2017); GTEx eQTL https://storage.googleapis.com/gtex_analysis_v7/single_tissue_eqtl_data/GTEx_Analysis_v7_eQTL.tar.gz (13 January 2019); UK Biobank GWAS summary statistics, http://www.nealelab.is/uk-biobank (4 October 2018); PopHumanScan, https://pophumanscan.uab.cat (13 January 2019).
Code availability
The software Relate can be downloaded from https://myersgroup.github.io/relate under an Academic Use Licence. External software used in the current study were downloaded from the following URLs: ARGweaver, https://github.com/mdrasmus/argweaver (24 January 2017); RENT+, https://github.com/SajadMirzaei/RentPlus (2 October 2017); msprime, https://github.com/tskit-dev/msprime (22 July 2017); msmc, https://github.com/stschiff/msmc2 (14 October 2017); SMC++, https://github.com/popgenmethods/smcpp (14 October 2017); simuPOP, http://simupop.sourceforge.net/ (27 June 2018); mbs, http://www.sendou.soken.ac.jp/esb/innan/InnanLab/ (27 June 2018); SDS, https://github.com/yairf/SDS (27 June 2018), selscan, https://github.com/szpiech/selscan (31 July 2018); hapbin, https://github.com/evotools/hapbin (11 December 2018).
References
Griffiths, R. C. & Marjoram, P. Ancestral inference from samples of DNA sequences with recombination. J. Comput. Biol. 3, 479–502 (1996).
Rasmussen, M. D., Hubisz, M. J., Gronau, I. & Siepel, A. Genome-wide inference of ancestral recombination graphs. PLoS Genet. 10, e1004342 (2014).
Kingman, J. F. C. On the genealogy of large populations. J. Appl. Probab. 19, 27–43 (1982).
Hudson, R. R. Properties of a neutral allele model with intragenic recombination. Theor. Popul. Biol. 23, 183–201 (1983).
McVean, G. A. T. & Cardin, N. J. Approximating the coalescent with recombination. Philos. Trans. R. Soc. Lond. B 360, 1387–1393 (2005).
Hein, J. Reconstructing evolution of sequences subject to recombination using parsimony. Math. Biosci. 98, 185–200 (1990).
Song, Y. S. & Hein, J. Constructing minimal ancestral recombination graphs. J. Comput. Biol. 12, 147–169 (2005).
Kececioglu, J. & Gusfield, D. Reconstructing a history of recombinations from a set of sequences. Discret. Appl. Math. 88, 239–260 (1998).
Wang, L., Zhang, K. & Zhang, L. Perfect phylogenetic networks with recombination. J. Comput. Biol. 8, 69–78 (2001).
Wu, Y. New methods for inference of local tree topologies with recombinant SNP sequences in populations. IEEE/ACM Trans. Comput. Biol. Bioinforma. 8, 182–193 (2011).
Mirzaei, S. & Wu, Y. RENT+: an improved method for inferring local genealogical trees from haplotypes with recombination. Bioinformatics 33, 1021–1030 (2017).
Menozzi, P., Piazza, A. & Cavalli-Sforza, L. Synthetic maps of human gene frequencies in Europeans. Science 201, 786–792 (1978).
Pritchard, J. K., Stephens, M. & Donnelly, P. Inference of population structure using multilocus genotype data. Genetics 155, 945–959 (2000).
Novembre, J. et al. Genes mirror geography within Europe. Nature 456, 98–101 (2008).
Li, H. & Durbin, R. Inference of human population history from individual whole-genome sequences. Nature 475, 493–496 (2011).
Henderson, D., Zhu, S. & Lunter, G. Demographic inference using particle filters for continuous Markov jump processes. Preprint at bioRxiv https://doi.org/10.1101/382218 (2018).
Falush, D., Stephens, M. & Pritchard, J. K. Inference of population structure using multilocus genotype data: linked loci and correlated allele frequencies. Genetics 164, 1567–1587 (2003).
Reich, D. D. E. et al. Linkage disequilibrium in the human genome. Nature 411, 199–204 (2001).
Schiffels, S. & Durbin, R. Inferring human population size and separation history from multiple genome sequences. Nat. Genet. 46, 919–925 (2014).
Terhorst, J., Kamm, J. A. & Song, Y. S. Robust and scalable inference of population history froth hundreds of unphased whole genomes. Nat. Genet. 49, 303–309 (2017).
Green, R. E. et al. A draft sequence of the Neandertal genome. Science 328, 710–722 (2010).
Sudmant, P. H. et al. An integrated map of structural variation in 2,504 human genomes. Nature 526, 75–81 (2015).
The 1000 Genomes Project Consortium. A global reference for human genetic variation. Nature 526, 68–74 (2015).
Harris, K. Evidence for recent, population-specific evolution of the human mutation rate. Proc. Natl Acad. Sci. USA 112, 3439–3444 (2015).
Voight, B. F., Kudaravalli, S., Wen, X. & Pritchard, J. K. A map of recent positive selection in the human genome. PLoS Biol. 4, e72 (2006).
Li, N. & Stephens, M. Modeling linkage disequilibrium and identifying recombination hotspots using single-nucleotide polymorphism data. Genetics 165, 2213–2233 (2003).
Kelleher, J., Etheridge, A. M. & McVean, G. Efficient coalescent simulation and genealogical analysis for large sample sizes. PLoS Comput. Biol. 12, e1004842 (2016).
Bae, C. J., Douka, K. & Petraglia, M. D. On the origin of modern humans: Asian perspectives. Science 358, eaai9067 (2017).
Liu, X. & Fu, Y.-X. Exploring population size changes using SNP frequency spectra. Nat. Genet. 47, 555–559 (2015).
Chheda, H. et al. Whole genome view of the consequences of a population bottleneck using 2926 genome sequences from Finland and United Kingdom. Eur. J. Hum. Genet. 25, 477–484 (2017).
Duret, L. & Galtier, N. Biased gene conversion and the evolution of mammalian genomic landscapes. Annu. Rev. Genom. Hum. Genet. 10, 285–311 (2009).
Meyer, M. et al. A high-coverage genome sequence from an archaic Denisovan individual. Science 338, 222–226 (2012).
Prüfer, K. et al. The complete genome sequence of a Neanderthal from the Altai Mountains. Nature 505, 43–49 (2014).
Sankararaman, S., Patterson, N., Li, H., Pääbo, S. & Reich, D. The date of interbreeding between Neandertals and modern humans. PLoS Genet. 8, e1002947 (2012).
Fu, Q. et al. An early modern human from Romania with a recent Neanderthal ancestor. Nature 524, 216–219 (2015).
Hammer, M. F., Woerner, A. E., Mendez, F. L., Watkins, J. C. & Wall, J. D. Genetic evidence for archaic admixture in Africa. Proc. Natl Acad. Sci. USA 108, 15123–15128 (2011).
Ragsdale, A. P. & Gravel, S. Models of archaic admixture and recent history from two-locus statistics. PLoS Genet. https://doi.org/10.1371/journal.pgen.1008204 (2019).
Mathieson, I. et al. Genome-wide patterns of selection in 230 ancient Eurasians. Nature 528, 499–503 (2015).
Edge, M. & Coop, G. Reconstructing the history of polygenic scores using coalescent trees. Genetics 211, 235–262 (2019).
Simons, Y. B., Bullaughey, K., Hudson, R. R. & Sella, G. A population genetic interpretation of GWAS findings for human quantitative traits. PLoS Biol. 16, e2002985 (2018).
Enattah, N. S. et al. Identification of a variant associated with adult-type hypolactasia. Nat. Genet. 30, 233–237 (2002).
Hardouin, E. et al. Positive Selection in East Asians for an EDAR Allele that Enhances NF-κB Activation. PLoS ONE 3, e2209 (2008).
Miretti, M. M. et al. A high-resolution linkage-disequilibrium map of the human major histocompatibility complex and first generation of tag single-nucleotide polymorphisms. Am. J. Hum. Genet. 76, 634–646 (2005).
Sadier, A., Viriot, L., Pantalacci, S. & Laudet, V. The ectodysplasin pathway: from diseases to adaptations. Trends Genet. 30, 24–31 (2014).
Pritchard, J. K., Pickrell, J. K. & Coop, G. The genetics of human adaptation: hard sweeps, soft sweeps, and polygenic adaptation. Curr. Biol. 20, R208–R215 (2010).
Zhang, G., Muglia, L. J., Chakraborty, R., Akey, J. M. & Williams, S. M. Signatures of natural selection on genetic variants affecting complex human traits. Appl. Transl. Genomics 2, 78–94 (2013).
Berg, J. J. & Coop, G. A population genetic signal of polygenic adaptation. PLoS Genet. 10, e1004412 (2014).
Field, Y. et al. Detection of human adaptation during the past 2000 years. Science 354, 760–764 (2016).
Sohail, M. et al. Signals of polygenic adaptation on height have been overestimated due to uncorrected population structure in genome-wide association studies. eLife 8, e39702 (2019).
Berg, J. J. et al. Reduced signal for polygenic adaptation of height in UK Biobank. eLife 8, e39725 (2019).
Maruyama, T. The age of an allele in a finite population. Genet. Res. 23, 137 (1974).
Kiezun, A. et al. Deleterious alleles in the human genome are on average younger than neutral alleles of the same frequency. PLoS Genet. 9, e1003301 (2013).
Bycroft, C. et al. The UK Biobank resource with deep phenotyping and genomic data. Nature 562, 203–209 (2018).
Casto, A. M. & Feldman, M. W. Genome-wide association study SNPs in the human genome diversity project populations: does selection affect unlinked SNPs with shared trait associations? PLoS Genet. 7, e1001266 (2011).
Wilde, S. et al. Direct evidence for positive selection of skin, hair, and eye pigmentation in Europeans during the last 5,000 y. Proc. Natl Acad. Sci. USA 111, 4832–4837 (2014).
Bulik-Sullivan, B. K. et al. LD score regression distinguishes confounding from polygenicity in genome-wide association studies. Nat. Genet. 47, 291–295 (2015).
Turchin, M. C. et al. Evidence of widespread selection on standing variation in Europe at height-associated SNPs. Nat. Genet. 44, 1015–1019 (2012).
Robinson, M. R. et al. Population genetic differentiation of height and body mass index across Europe. Nat. Genet. 47, 1357–1362 (2015).
Novick, D., Montgomery, W., Treuer, T., Moneta, M. V. & Haro, J. M. Sex differences in the course of schizophrenia across diverse regions of the world. Neuropsychiatr. Dis. Treat. 12, 2927–2939 (2016).
Crespi, B., Summers, K. & Dorus, S. Adaptive evolution of genes underlying schizophrenia. Proc. R. Soc. B 274, 2801–2810 (2007).
Young, J. H. et al. Differential susceptibility to hypertension is due to selection during the out-of-Africa expansion. PLoS Genet. 1, e82 (2005).
Hinch, A. G. et al. The landscape of recombination in African Americans. Nature 476, 170–175 (2011).
Fledel-Alon, A. et al. Variation in human recombination rates and its genetic determinants. PLoS ONE 6, e20321 (2011).
Kelleher, J., Wong, Y., Albers, P., Wohns, A. W. & McVean, G. Inferring the ancestry of everyone. Preprint at bioRxiv https://doi.org/10.1101/458067 (2018).
Griffiths, R. C. & Tavaré, S. The age of a mutation in a general coalescent tree. Stoch. Model. 14, 273–295 (1998).
Peng, B. & Kimmel, M. simuPOP: a forward-time population genetics simulation environment. Bioinformatics 21, 3686–3687 (2005).
Teshima, K. M. & Innan, H. mbs: modifying Hudson’s ms software to generate samples of DNA sequences with a biallelic site under selection. BMC Bioinformatics 10, 166 (2009).
Ruiz-Linares, A. et al. Admixture in Latin America: geographic structure, phenotypic diversity and self-perception of ancestry based on 7,342 individuals. PLoS Genet. 10, e1004572 (2014).
Ward, L. D. & Kellis, M. HaploReg: a resource for exploring chromatin states, conservation, and regulatory motif alterations within sets of genetically linked variants. Nucleic Acids Res. 40, D930–D934 (2011).
MacArthur, J. et al. The new NHGRI-EBI Catalog of published genome-wide association studies (GWAS Catalog). Nucleic Acids Res. 45, D896–D901 (2016).
Ruderfer, D. M. et al. Genomic dissection of bipolar disorder and schizophrenia, including 28 subphenotypes. Cell 173, 1705–1715.e16 (2018).
Acknowledgements
We thank N. Barton, D. Falush, M. Przeworski, G. Sella, J. Terhorst, P. Palamara, G. Lunter, J. Marchini, S. Hu, C. B. Cole, T. Aid and C. E. West for helpful comments, ideas and suggestions. L.S. acknowledges the support provided through the Engineering and Physical Sciences Research Council (grant number EP/G03706X/1). M.F. acknowledges the support provided through the Natural Sciences and Engineering Research Council of Canada (PGS D) and the Clarendon Scholarship. S.R.M. acknowledges the support provided by the Wellcome Trust Investigator Award (grant number 098387/Z/12/Z and 212284/Z/18/Z). For computation we used the Oxford Biomedical Research Computing facility, a joint development between the Wellcome Centre for Human Genetics and the Big Data Institute supported by Health Data Research UK and the NIHR Oxford Biomedical Research Centre. Financial support was provided by the Wellcome Trust Core Award grant number 203141/Z/16/Z. The views expressed are those of the authors and not necessarily those of the NHS, the NIHR or the Department of Health.
Author information
Authors and Affiliations
Contributions
S.R.M. designed the study. L.S. and S.R.M. developed Relate with contributions by M.F. in the development of the algorithm for estimating coalescence rates. L.S. and S.R.M. performed the analysis, S.S. provided supplementary data and L.S. and S.R.M. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
S.R.M. is a director of GENSCI limited. The remaining authors declare no competing financial interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Schematics of the tree builder, branch length estimator, and modified Li-and-Stephens HMM.
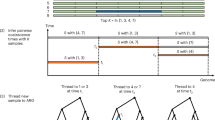
a, Schematic of the hierarchical clustering algorithm for estimating tree topology. In the case of no recombination, the algorithm obtains matrix d containing the number of derived mutations as input. Row i of this matrix determines the order in which haplotype i coalesced with other haplotypes. Using Eq. (1) in Supplementary Note: Method Details, the algorithm finds the pair that coalesces with each other before coalescing with any other sequence. In the example shown here, we can coalesce haplotypes 0 and 2 or haplotypes 3 and 4. We choose to coalesce haplotypes 1 and 2 first because the symmetrised distance is smaller for this pair, however this choice does not affect tree topology in this case. The resulting tree topology is consistent with the gene tree describing the data. In contrast, when the hierarchical clustering algorithm is applied to the symmetrised matrix (d(i,j) + d(j,i))i,j = 1,…,N, haplotypes 2 and 3 are coalesced first and the constructed tree topology is wrong. This is equivalent to applying the UPGMA algorithm to the symmetrised matrix of derived mutations (Sokal R., Michener C. University of Kansas Science Bulletin 38, 1409–1438, 1958). b, Schematic of possible proposal moves in the MCMC algorithm for estimating branch lengths. We propose either a change in the order of coalescence events or a change in the time while k ancestors remain. c, Schematic of the modified Li-and-Stephens hidden Markov model (HMM) applied to haplotype k, which has alleles 0, 1, 1 at loci ℓ - 1, ℓ, ℓ +1. The emission and transition probabilities shown correspond to the path indicated by the red solid line. At SNP ℓ - 1, the reference haplotype is h1 which has allele 1. Because the allele of haplotype k is 0, the allele of the MRCA with h1 is also 0 assuming that every mutation is unique in history. Therefore, the emission probability equals 1-p, where p is the probability of a mutation. At SNP ℓ, the reference haplotype has changed to h2. The alleles of haplotype k and h2 are 1. Therefore, the MRCA has allele 1 and the emission probability is given by 1-p. At SNP ℓ + 1, haplotype k has allele 1. The allele of the reference haplotype h2 is 0 and so is that of the MRCA, such that the emission probability equals p. Using this HMM, we calculate the likelihood Pm(Hℓ = j | D(k)). This is the likelihood of copying from reference haplotype j at SNP ℓ, conditional on observing D(k). We notice that Pm(Hℓ = j | D(k)) is obtained as the sum of all possible paths when Hℓ = j is fixed (indicated by the dashed lines).
Supplementary Figure 2 Mapping rule for mutations and sensitivity of the modified Li-and-Stephens HMM to parameter choice.
a, Schematic illustrating which parts of Relate use heuristic approaches. b, Heatmap showing the necessary and sufficient overlap between the set of descendants of a branch (Ct) and the set of carriers of the derived allele (Cd), given |Ct| and |Cd|, with N=100, as determined by Eqs. (4) and (5). Colors show |Ct ∩ Cd|/min{|Ct|, |Cd|}, where white indicates that a mutation can never be mapped for the corresponding combination of |Ct| and |Cd|. c,d, Number of non-mapping SNPs for different values of p (horizontal axis) and R (vertical axis) for N=500 (c) and N=1000 (d) (Supplementary Note Method details, Section 3.3 for definition of parameters). The subsets of haplotypes are chosen uniformly at random from all haplotypes. We calculated the mean over 50 randomly chosen subregions of length 1200 SNPs on chromosome 20. In our implementation, we fixed p = 0.025 and R = 2500.
Supplementary Figure 3 Performance of Relate on simulated data.
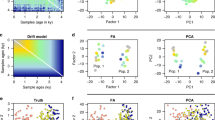
a, Estimated times to most recent common ancestors (TMRCAs) between pairs of haplotypes compared to the truth for Relate, ARGweaver, and Rent+. b, Estimated ages of mutations plotted against the true age for Relate, ARGweaver, and Rent+. We determine the age of a mutation by placing it at the midpoint of the branch onto which it maps. In a and b, we simulate N=200 haplotypes with 2Ne = 40,000. c, TMRCAs between pairs of haplotypes compared to the truth for a simulated data set with N=200 haplotypes and a population bottleneck resembling that of Europeans, where branch lengths are estimated using a constant population size of 2Ne = 30,000. d, Estimated TMRCAs compared to the truth for the same example as in c, where branch lengths and population size history are jointly inferred. e, Robinson-Foulds distance and f, pairwise TMRCA distance averaged over 2.4Mb for Relate, ARGweaver, and RENT+. We estimate genealogies for N=50 haplotypes at different number of errors. In addition, we show the accuracy of the genealogy corresponding to N=50 haplotypes, embedded in an estimated genealogy for N = 1000 haplotypes (see Supplementary Note: Simulations, Section 2.1 for details). g, Robustness of Relate with respect to randomly introduced flipped mutations. We show the fraction of SNPs mapping to a unique branch, fraction of correctly flipped SNPs, and fraction of correctly unflipped SNPs for Relate. We exclude SNPs at frequency 1, which always map to the tree. We simulate 2.5Mb for N=200 haplotypes with 2Ne = 30,000. h, Population size estimates for simulations with a discrete bottleneck, an increasing trend, and a decreasing trend in populations size. Estimates using Relate are shown by the blue solid line. We apply SMC++ to the same data set and we also apply MSMC2 with 2 and 8 haplotypes. In the inset, we show the mutation rate over time estimated by Relate. For each scenario, we simulate 200Mb for N=200 haplotypes. In all simulations, the mutation rate is set to 1.25 × 10–8 and recombination rates are taken from the 1000 Genomes Project map for chromosome 1.
Supplementary Figure 4 Accuracy under perturbations from infinite-sites, constant mutation rate, or perfect phase.
a, Expected heterozygosity (π) calculated for 20,000 randomly chosen 100kb windows. Circles show the mean and bars indicate the 2.5th and 97.5th percentiles. b, Derived allele frequencies, and c, LD decay patterns. For a, b, and c, we used ten 1000 Genomes Project individuals, and simulated 20 haplotypes using the demographic histories estimated by Relate. Each statistic is calculated using chromosome 1 (see Supplementary Note: Simulations, Section 3 for details). d, Ratio of estimated and true age of a mutation, estimated as the mean of the lower and upper ends of the branch onto which the mutation maps, as a function of DAF. Circles show the mean ratio and bars indicate the 2.5th and 97.5th percentiles. Base-line simulation assumes infinite-sites and a constant mutation rate of 1.25 × 10-8. We introduce perturbations, such as a variable mutation rate to a subset of sites, hypermutable base-pair positions emulating CpG dinucleotides, and inferred phase (see Supplementary Note Simulations, Section 4 for details). e, Accuracy of Relate-estimated population sizes on the same simulations as in (a). f, Normalised mutation rate for null mutations with a constant mutation rate of 1.25 × 10-8 and a mutation category with an activity period in [10,50) kYBP during which the mutation rate doubled (dashed lines). g, Fraction of not mapping mutations as a function of DAF for the simulation with CpG-like mutations, categorised by whether the CpG-like site mutated once or more than once.
Supplementary Figure 5 Properties of the genealogy constructed for the 1000 Genomes Project data set.
a, Number of trees built versus the recombination distance for all 22 chromosomes. b, Mean number of SNPs that map to a unique branch versus the recombination distance in that bin. Every point represents a subregion of 105 SNPs. c, Fraction of SNPs that could not be mapped to a unique branch for SNPs excluding singletons (left) and SNPs with derived allele frequencies larger than 10 (right). d, Fraction of SNPs that could not be mapped to a unique branch for all 96 possible triplet mutations, excluding singletons. e, Fraction of SNPs that were flipped for all 96 possible triplet mutations, excluding singletons. In d and e, CpG transitions are indicated in red. The 95% confidence intervals of the means are indicated by black brackets. Each triplet mutation category comprises at least 46,000 mutations. f, Fraction of non-mapping SNPs by derived allele frequency of the mutation in the sample. For each frequency, we divide the number of non-mapping mutations of that frequency by the number of mutations of that frequency. g, Fraction of flipped SNPs by derived allele frequency of the mutation after flipping. For each frequency, we divide the number of flipped SNPs of that frequency (after flipping) by the number of SNPs of that frequency.
Supplementary Figure 6 Historical effective population sizes and evidence of introgression, mutation rate trends for 96 triplet mutations.
a, Historical effective population sizes for all 26 populations in the 1000 Genomes Project dataset. For each population, we first extract the genealogy corresponding to that population. We then estimate the population size using this genealogy. b, Number of mutations on branches with an upper end older than 1M YBP and lower end younger than 30,000 YBP, categorised by whether the mutation is additionally found only in Neanderthals, only in Denisovans, both, or neither. For each category, we also distinguish whether the mutation is unique to the population of interest or shared with other populations in AFR, EUR, SAS, or EAS. c, Number of mutations binned by age of upper and lower coalescence event, relative to the expected number of mutations when randomising topology while fixing ages of coalescence events for four simulated data sets (Methods). We simulated 3000Mb with population size histories of YRI, CHB, GBR, and BEB. d, Normalised mutation rate of triplet mutations for all 96 possible categories. Analogous to Fig. 4a.
Supplementary Figure 7 Power simulations for selection test.
a, Ratio of estimated and true lower-end ages of the branches onto which a mutation with present-day DAF of 0.5 maps. This mutation has a selection coefficient of 0, 0.001, or 0.01 and is positioned at 10Mb of a 20Mb simulated genomic region with Relate-estimated population size histories for GBR and YRI. We simulated 100 realisations of N=200 haplotypes. Circles indicate the mean ratios. b, P-values for selection evidence in simulations calculated using true trees (horizontal axis) and estimated trees (vertical axis) for the same simulation scenario as in a. We plot p-values pR of the Relate Selection Test for 500 loci under no selection (circles) and 200 loci under weak selection (triangles). c, d, Power simulations with N=1000 haplotypes and present-day derived allele frequencies of 0.3, 0.5, and 0.7. We assume a population size history estimated for YRI (c), and GBR (d), respectively. The significance threshold is 0.05. We show power estimates using the p-values for trees estimated by Relate, as well as those for the true trees. In both cases, we estimate power using raw p-values of our test statistic (top row) and empirical p-values given the distribution of raw p-values in the neutral case (bottom row). For iHS, SDS, and trSDS, power is estimated by standardising raw scores by the frequency specific mean and standard deviation under the null. In the top row, we assume a standard normal distribution of the standardised score and in the bottom row, we calculate empirical p-values by determining a critical score corresponding to the 0.05 significance level in the neutral case.
Supplementary Figure 8 Histograms of p-values for evidence of selection of traits.
Histograms of p-values for evidence of selection of traits (Methods). We aggregated both effect directions of 84 considered traits, as well as populations in each of the four considered geographic regions (AFR, EAS, EUR, SAS).
Supplementary information
Supplementary Information
Supplementary Figs. 1–8, Tables 1–3 and Note
Rights and permissions
About this article
Cite this article
Speidel, L., Forest, M., Shi, S. et al. A method for genome-wide genealogy estimation for thousands of samples. Nat Genet 51, 1321–1329 (2019). https://doi.org/10.1038/s41588-019-0484-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-019-0484-x
This article is cited by
-
kalis: a modern implementation of the Li & Stephens model for local ancestry inference in R
BMC Bioinformatics (2024)
-
Climate change from an ectotherm perspective: evolutionary consequences and demographic change in amphibian and reptilian populations
Biodiversity and Conservation (2024)
-
The selection landscape and genetic legacy of ancient Eurasians
Nature (2024)
-
Impact of population structure in the estimation of recent historical effective population size by the software GONE
Genetics Selection Evolution (2023)
-
The history and organization of the Workshop on Population and Speciation Genomics
Evolution: Education and Outreach (2023)