Abstract
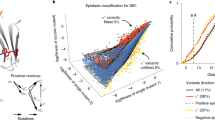
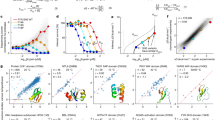
We describe an experimental method of three-dimensional (3D) structure determination that exploits the increasing ease of high-throughput mutational scans. Inspired by the success of using natural, evolutionary sequence covariation to compute protein and RNA folds, we explored whether ‘laboratory’, synthetic sequence variation might also yield 3D structures. We analyzed five large-scale mutational scans and discovered that the pairs of residues with the largest positive epistasis in the experiments are sufficient to determine the 3D fold. We show that the strongest epistatic pairings from genetic screens of three proteins, a ribozyme and a protein interaction reveal 3D contacts within and between macromolecules. Using these experimental epistatic pairs, we compute ab initio folds for a GB1 domain (within 1.8 Å of the crystal structure) and a WW domain (2.1 Å). We propose strategies that reduce the number of mutants needed for contact prediction, suggesting that genomics-based techniques can efficiently predict 3D structure.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The main data analyzed in this study are publicly available from the original publications (refs. 13,18,36,38,42,43). All other data supporting the findings of this study are available within the article and its Supplementary Information files, and from the GitHub repository (https://github.com/debbiemarkslab/3D_from_DMS_Extended_Data).
Code availability
The code used in this study (along with folded models) is available at https://github.com/debbiemarkslab/3D_from_DMS_Extended_Data, and utilities for folding and ranking are available from the EVcouplings GitHub repository (https://github.com/debbiemarkslab/EVcouplings).
References
Hopf, T. A. et al. Three-dimensional structures of membrane proteins from genomic sequencing. Cell 149, 1607–1621 (2012).
Marks, D. S. et al. Protein 3D structure computed from evolutionary sequence variation. PLoS ONE 6, e28766 (2011).
Hopf, T. A. et al. Sequence co-evolution gives 3D contacts and structures of protein complexes. eLife 3, e03430 (2014).
Weinreb, C. et al. 3D RNA and functional interactions from evolutionary couplings. Cell 165, 963–975 (2016).
Toth-Petroczy, A. et al. Structured states of disordered proteins from genomic sequences. Cell 167, 158–170.e12 (2016).
Morcos, F. et al. Direct-coupling analysis of residue coevolution captures native contacts across many protein families. Proc. Natl Acad. Sci. USA 108, E1293–E1301 (2011).
Kosciolek, T. & Jones, D. T. De novo structure prediction of globular proteins aided by sequence variation-derived contacts. PLoS ONE 9, e92197 (2014).
Ovchinnikov, S. et al. Large-scale determination of previously unsolved protein structures using evolutionary information. eLife 4, e09248 (2015).
Finn, R. D. et al. The Pfam protein families database: towards a more sustainable future. Nucleic Acids Res. 44, D279–D285 (2016).
Romero, P. A., Tran, T. M. & Abate, A. R. Dissecting enzyme function with microfluidic-based deep mutational scanning. Proc. Natl Acad. Sci. USA 112, 7159–7164 (2015).
Roscoe, B. P. & Bolon, D. N. Systematic exploration of ubiquitin sequence, E1 activation efficiency, and experimental fitness in yeast. J. Mol. Biol. 426, 2854–2870 (2014).
Roscoe, B. P., Thayer, K. M., Zeldovich, K. B., Fushman, D. & Bolon, D. N. Analyses of the effects of all ubiquitin point mutants on yeast growth rate. J. Mol. Biol. 425, 1363–1377 (2013).
Melamed, D., Young, D. L., Gamble, C. E., Miller, C. R. & Fields, S. Deep mutational scanning of an RRM domain of the Saccharomyces cerevisiae poly(A)-binding protein. RNA 19, 1537–1551 (2013).
Stiffler, M. A., Hekstra, D. R. & Ranganathan, R. Evolvability as a function of purifying selection in TEM-1 β-lactamase. Cell 160, 882–892 (2015).
McLaughlin, R. N. Jr, Poelwijk, F. J., Raman, A., Gosal, W. S. & Ranganathan, R. The spatial architecture of protein function and adaptation. Nature 491, 138–142 (2012).
Kitzman, J. O., Starita, L. M., Lo, R. S., Fields, S. & Shendure, J. Massively parallel single-amino-acid mutagenesis. Nat. Methods 12, 203–206 (2015).
Melnikov, A., Rogov, P., Wang, L., Gnirke, A. & Mikkelsen, T. S. Comprehensive mutational scanning of a kinase in vivo reveals substrate-dependent fitness landscapes. Nucleic Acids Res. 42, e112 (2014).
Araya, C. L. et al. A fundamental protein property, thermodynamic stability, revealed solely from large-scale measurements of protein function. Proc. Natl Acad. Sci. USA 109, 16858–16863 (2012).
Firnberg, E., Labonte, J. W., Gray, J. J. & Ostermeier, M. A comprehensive, high-resolution map of a gene’s fitness landscape. Mol. Biol. Evol. 31, 1581–1592 (2014).
Starita, L. M. et al. Massively parallel functional analysis of BRCA1 RING domain variants. Genetics 200, 413–422 (2015).
Rockah-Shmuel, L., Toth-Petroczy, A. & Tawfik, D. S. Systematic mapping of protein mutational space by prolonged drift reveals the deleterious effects of seemingly neutral mutations. PLoS Comput. Biol. 11, e1004421 (2015).
Jacquier, H. et al. Capturing the mutational landscape of the β-lactamase TEM-1. Proc. Natl Acad. Sci. USA 110, 13067–13072 (2013).
Qi, H. et al. A quantitative high-resolution genetic profile rapidly identifies sequence determinants of hepatitis C viral fitness and drug sensitivity. PLoS Pathog. 10, e1004064 (2014).
Wu, N. C. et al. Functional constraint profiling of a viral protein reveals discordance of evolutionary conservation and functionality. PLoS Genet. 11, e1005310 (2015).
Mishra, P., Flynn, J. M., Starr, T. N. & Bolon, D. N. Systematic mutant analyses elucidate general and client-specific aspects of Hsp90 function. Cell Rep. 15, 588–598 (2016).
Doud, M. B. & Bloom, J. D. Accurate measurement of the effects of all amino-acid mutations to influenza hemagglutinin. Viruses 8, E155 (2016).
Deng, Z. et al. Deep sequencing of systematic combinatorial libraries reveals β-lactamase sequence constraints at high resolution. J. Mol. Biol. 424, 150–167 (2012).
Starita, L. M. et al. Activity-enhancing mutations in an E3 ubiquitin ligase identified by high-throughput mutagenesis. Proc. Natl Acad. Sci. USA 110, E1263–E1272 (2013).
Aakre, C. D. et al. Evolving new protein–protein interaction specificity through promiscuous intermediates. Cell 163, 594–606 (2015).
Julien, P., Minana, B., Baeza-Centurion, P., Valcarcel, J. & Lehner, B. The complete local genotype–phenotype landscape for the alternative splicing of a human exon. Nat. Commun. 7, 11558 (2016).
Li, C., Qian, W., Maclean, C. J. & Zhang, J. The fitness landscape of a tRNA gene. Science 352, 837–840 (2016).
Mavor, D. et al. Determination of ubiquitin fitness landscapes under different chemical stresses in a classroom setting. eLife 5, e15802 (2016).
Fowler, D. M. & Fields, S. Deep mutational scanning: a new style of protein science. Nat. Methods 11, 801–807 (2014).
Gasperini, M., Starita, L. & Shendure, J. The power of multiplexed functional analysis of genetic variants. Nat. Protoc. 11, 1782–1787 (2016).
Starita, L. M. et al. Variant interpretation: functional assays to the rescue. Am. J. Hum. Genet. 101, 315–325 (2017).
Kobori, S. & Yokobayashi, Y. High-throughput mutational analysis of a twister ribozyme. Angew. Chem. Int. Ed. Engl. 55, 10354–10357 (2016).
Starr, T. N., Picton, L. K. & Thornton, J. W. Alternative evolutionary histories in the sequence space of an ancient protein. Nature 549, 409–413 (2017).
Sarkisyan, K. S. et al. Local fitness landscape of the green fluorescent protein. Nature 533, 397–401 (2016).
Chen, J. & Stites, W. E. Energetics of side chain packing in staphylococcal nuclease assessed by systematic double mutant cycles. Biochemistry 40, 14004–14011 (2001).
Ackermann, E. J., Ang, E. T., Kanter, J. R., Tsigelny, I. & Taylor, P. Identification of pairwise interactions in the α-neurotoxin–nicotinic acetylcholine receptor complex through double mutant cycles. J. Biol. Chem. 273, 10958–10964 (1998).
Horovitz, A. Double-mutant cycles: a powerful tool for analyzing protein structure and function. Fold. Des. 1, R121–R126 (1996).
Olson, C. A., Wu, N. C. & Sun, R. A comprehensive biophysical description of pairwise epistasis throughout an entire protein domain. Curr. Biol. 24, 2643–2651 (2014).
Diss, G. & Lehner, B. The genetic landscape of a physical interaction. eLife 7, e32472 (2018).
Adkar, B. V. et al. Protein model discrimination using mutational sensitivity derived from deep sequencing. Structure 20, 371–381 (2012).
Sahoo, A., Khare, S., Devanarayanan, S., Jain, P. C. & Varadarajan, R. Residue proximity information and protein model discrimination using saturation-suppressor mutagenesis. eLife 4, e09532 (2015).
Melamed, D., Young, D. L., Miller, C. R. & Fields, S. Combining natural sequence variation with high throughput mutational data to reveal protein interaction sites. PLoS Genet. 11, e1004918 (2015).
Salinas, V. H. & Ranganathan, R. Coevolution-based inference of amino acid interactions underlying protein function. eLife 7, e34300 (2018).
Kamisetty, H., Ovchinnikov, S. & Baker, D. Assessing the utility of coevolution-based residue–residue contact predictions in a sequence- and structure-rich era. Proc. Natl Acad. Sci. USA 110, 15674–15679 (2013).
Gronenborn, A. M. et al. A novel, highly stable fold of the immunoglobulin binding domain of streptococcal protein G. Science 253, 657–661 (1991).
Gallagher, T., Alexander, P., Bryan, P. & Gilliland, G. L. Two crystal structures of the B1 immunoglobulin-binding domain of streptococcal protein G and comparison with NMR. Biochemistry 33, 4721–4729 (1994).
Tomlinson, J. H., Craven, C. J., Williamson, M. P. & Pandya, M. J. Dimerization of protein G B1 domain at low pH: a conformational switch caused by loss of a single hydrogen bond. Proteins 78, 1652–1661 (2010).
Bouvignies, G., Meier, S., Grzesiek, S. & Blackledge, M. Ultrahigh-resolution backbone structure of perdeuterated protein GB1 using residual dipolar couplings from two alignment media. Angew. Chem. Int. Ed. Engl. 45, 8166–8169 (2006).
Bouvignies, G., Markwick, P., Bruschweiler, R. & Blackledge, M. Simultaneous determination of protein backbone structure and dynamics from residual dipolar couplings. J. Am. Chem. Soc. 128, 15100–15101 (2006).
Li, F., Grishaev, A., Ying, J. & Bax, A. Side chain conformational distributions of a small protein derived from model-free analysis of a large set of residual dipolar couplings. J. Am. Chem. Soc. 137, 14798–14811 (2015).
Wylie, B. J. et al. Ultrahigh resolution protein structures using NMR chemical shift tensors. Proc. Natl Acad. Sci. USA 108, 16974–16979 (2011).
Lian, L. Y., Derrick, J. P., Sutcliffe, M. J., Yang, J. C. & Roberts, G. C. Determination of the solution structures of domains II and III of protein G from Streptococcus by 1H nuclear magnetic resonance. J. Mol. Biol. 228, 1219–1234 (1992).
Derrick, J. P. & Wigley, D. B. The third IgG-binding domain from streptococcal protein G. An analysis by X-ray crystallography of the structure alone and in a complex with Fab. J. Mol. Biol. 243, 906–918 (1994).
Alexander, P. A., He, Y., Chen, Y., Orban, J. & Bryan, P. N. A minimal sequence code for switching protein structure and function. Proc. Natl Acad. Sci. USA 106, 21149–21154 (2009).
He, Y., Chen, Y., Alexander, P., Bryan, P. N. & Orban, J. NMR structures of two designed proteins with high sequence identity but different fold and function. Proc. Natl Acad. Sci. USA 105, 14412–14417 (2008).
He, Y., Chen, Y., Alexander, P. A., Bryan, P. N. & Orban, J. Mutational tipping points for switching protein folds and functions. Structure 20, 283–291 (2012).
Ferguson, N. et al. Using flexible loop mimetics to extend Φ-value analysis to secondary structure interactions. Proc. Natl Acad. Sci. USA 98, 13008–13013 (2001).
Pires, J. R. et al. Solution structures of the YAP65 WW domain and the variant L30 K in complex with the peptides GTPPPPYTVG, N-(n-octyl)-GPPPY and PLPPY and the application of peptide libraries reveal a minimal binding epitope. J. Mol. Biol. 314, 1147–1156 (2001).
Martinez-Rodriguez, S., Bacarizo, J., Luque, I. & Camara-Artigas, A. Crystal structure of the first WW domain of human YAP2 isoform. J. Struct. Biol. 191, 381–387 (2015).
Aragon, E. et al. Structural basis for the versatile interactions of Smad7 with regulator WW domains in TGF-β pathways. Structure 20, 1726–1736 (2012).
Aragon, E. et al. A Smad action turnover switch operated by WW domain readers of a phosphoserine code. Genes Dev. 25, 1275–1288 (2011).
Deo, R. C., Bonanno, J. B., Sonenberg, N. & Burley, S. K. Recognition of polyadenylate RNA by the poly(A)-binding protein. Cell 98, 835–845 (1999).
Safaee, N. et al. Interdomain allostery promotes assembly of the poly(A) mRNA complex with PABP and eIF4G. Mol. Cell 48, 375–386 (2012).
Ovchinnikov, S., Kamisetty, H. & Baker, D. Robust and accurate prediction of residue–residue interactions across protein interfaces using evolutionary information. eLife 3, e02030 (2014).
Glover, J. N. & Harrison, S. C. Crystal structure of the heterodimeric bZIP transcription factor c-Fos–c-Jun bound to DNA. Nature 373, 257–261 (1995).
Roth, A. et al. A widespread self-cleaving ribozyme class is revealed by bioinformatics. Nat. Chem. Biol. 10, 56–60 (2014).
Liu, Y., Wilson, T. J., McPhee, S. A. & Lilley, D. M. Crystal structure and mechanistic investigation of the twister ribozyme. Nat. Chem. Biol. 10, 739–744 (2014).
Ren, A. et al. In-line alignment and Mg2+ coordination at the cleavage site of the env22 twister ribozyme. Nat. Commun. 5, 5534 (2014).
Miao, Z. & Westhof, E. RNA structure: advances and assessment of 3D structure prediction. Ann. Rev. Biophys. 46, 483–503 (2017).
Brunger, A. T. Version 1.2 of the Crystallography and NMR System. Nat. Protoc. 2, 2728–2733 (2007).
Bradley, P., Misura, K. M. & Baker, D. Toward high-resolution de novo structure prediction for small proteins. Science 309, 1868–1871 (2005).
Ekeberg, M., Lövkvist, C., Lan, Y., Weigt, M. & Aurell, E. Improved contact prediction in proteins: using psuedolikelihoods to infer Potts models. Phys. Rev. E 87, 012707 (2013).
Marks, D. S., Hopf, T. A. & Sander, C. Protein structure prediction from sequence variation. Nat. Biotechnology 30, 1072–1080 (2012).
Tang, Y. et al. Protein structure determination by combining sparse NMR data with evolutionary couplings. Nat. Methods 12, 751–754 (2015).
Meiler, J. & Baker, D. Rapid protein fold determination using unassigned NMR data. Proc. Natl Acad. Sci. USA 100, 15404–15409 (2003).
Sjodt, M. et al. Structure of the peptidoglycan polymerase RodA resolved by evolutionary coupling analysis. Nature 556, 118–121 (2018).
Cheng, C. C. et al. Consistent global structures of complex RNA states through multidimensional chemical mapping. eLife 4, e07600 (2015).
Das, R. et al. Structural inference of native and partially folded RNA by high-throughput contact mapping. Proc. Natl Acad. Sci. USA 105, 4144–4149 (2008).
Matreyek, K. A. et al. Multiplex assessment of protein variant abundance by massively parallel sequencing. Nat. Genet. 50, 874–882 (2018).
Schmiedel, J. & Lehner, B. Determining protein structures using deep mutagenesis. Nat. Genet. https://doi.org/10.1038/s41588-019-0431-x (2019).
Fowler, D. M. et al. High-resolution mapping of protein sequence–function relationships. Nat. Methods 7, 741–746 (2010).
Buchan, D. W., Minneci, F., Nugent, T. C., Bryson, K. & Jones, D. T. Scalable web services for the PSIPRED protein analysis workbench. Nucleic Acids Res. 41, W349–W357 (2013).
Jones, D. T. Protein secondary structure prediction based on position-specific scoring matrices. J. Mol. Biol. 292, 195–202 (1999).
Van Zundert, G. C. P. et al. The HADDOCK2.2 web server: user-friendly integrative modeling of biomolecular complexes. J. Mol. Biol. 428, 720–725 (2016).
Bonneau, R. et al. Rosetta in CASP4: progress in ab initio protein structure prediction. Proteins 5, 119–126 (2011).
Acknowledgements
The authors thank the Marks, Sander and Silver laboratories for discussion and support. The authors also thank S. Ovchinnikov for performing ab initio structure predictions with Rosetta for comparison. Partial financial support for C.S. was provided from the US NIH (RO1 GM106303). K.P.B. thanks the NIH (R01 R01GM120574) for financial support.
Author information
Authors and Affiliations
Contributions
N.K.R., K.P.B. and D.S.M. performed the main analyses. N.J.R., K.P.B. and D.S.M. wrote the manuscript. F.J.P., M.A.S., N.P.G. and C.S. helped edit the manuscript. D.S.M. conceived the project. D.S.M. and C.S. supervised the study.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–9
Supplementary Table 1
Epistasis and fitness values for all double mutants
Supplementary Table 2
Top epistasis pairs for each pair of positions, with distance in Xtal or NMR
Supplementary Table 3
Precision of top epistatic pairs versus contacts in experimental structures
Supplementary Table 4
Epistasis versus experimental secondary structure scores
Supplementary Table 5
Accuracy of ab initio models versus experimental structures
Supplementary Table 6
RMSD between experimental structures
Supplementary Table 7
Small-library subsampling 3D fold accuracy and epistatic pair precisions
Rights and permissions
About this article
Cite this article
Rollins, N.J., Brock, K.P., Poelwijk, F.J. et al. Inferring protein 3D structure from deep mutation scans. Nat Genet 51, 1170–1176 (2019). https://doi.org/10.1038/s41588-019-0432-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-019-0432-9
This article is cited by
-
Protein design using structure-based residue preferences
Nature Communications (2024)
-
Predicting the Functional Changes in Protein Mutations Through the Application of BiLSTM and the Self-Attention Mechanism
Annals of Data Science (2024)
-
SESNet: sequence-structure feature-integrated deep learning method for data-efficient protein engineering
Journal of Cheminformatics (2023)
-
An Atlas of Variant Effects to understand the genome at nucleotide resolution
Genome Biology (2023)
-
ACIDES: on-line monitoring of forward genetic screens for protein engineering
Nature Communications (2023)