Abstract
Subcutaneous panniculitis-like T cell lymphoma (SPTCL), a non-Hodgkin lymphoma, can be associated with hemophagocytic lymphohistiocytosis (HLH), a life-threatening immune activation that adversely affects survival1,2. T cell immunoglobulin mucin 3 (TIM-3) is a modulator of immune responses expressed on subgroups of T and innate immune cells. We identify in ~60% of SPTCL cases germline, loss-of-function, missense variants altering highly conserved residues of TIM-3, c.245A>G (p.Tyr82Cys) and c.291A>G (p.Ile97Met), each with specific geographic distribution. The variant encoding p.Tyr82Cys TIM-3 occurs on a potential founder chromosome in patients with East Asian and Polynesian ancestry, while p.Ile97Met TIM-3 occurs in patients with European ancestry. Both variants induce protein misfolding and abrogate TIM-3’s plasma membrane expression, leading to persistent immune activation and increased production of inflammatory cytokines, including tumor necrosis factor-α and interleukin-1β, promoting HLH and SPTCL. Our findings highlight HLH–SPTCL as a new genetic entity and identify mutations causing TIM-3 alterations as a causative genetic defect in SPTCL. While HLH–SPTCL patients with mutant TIM-3 benefit from immunomodulation, therapeutic repression of the TIM-3 checkpoint may have adverse consequences.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Raw exome sequence data have been deposited at the European Genome-phenome Archive (EGA), which is hosted by the European Bioinformatics Institute (EMBL-EBI) and the Centre for Genomic Regulation (CRG), under accession number EGAS00001002765.
Change history
14 November 2018
In the version of this article originally published, the main-text sentence “In three patients of European ancestry, we identified the germline variant encoding p.Ile97Met in TIM-3, which was homozygous in two (P12 and P13) and heterozygous in one (P15) in the germline but with no TIM-3 plasma membrane expression in the tumor” misstated the identifiers of the two homozygous individuals, which should have been P13 and P14. The error has been corrected in the HTML, PDF and print versions of the paper.
References
Willemze, R. et al. Subcutaneous panniculitis-like T-cell lymphoma: definition, classification, and prognostic factors: an EORTC Cutaneous Lymphoma Group Study of 83 cases. Blood 111, 838–845 (2008).
Swerdlow, S. H. et al. The 2016 revision of the World Health Organization classification of lymphoid neoplasms. Blood 127, 2375–2390 (2016).
Gonzalez, C. L., Medeiros, L. J., Braziel, R. M. & Jaffe, E. S. T-cell lymphoma involving subcutaneous tissue. A clinicopathologic entity commonly associated with hemophagocytic syndrome. Am. J. Surg. Pathol. 15, 17–27 (1991).
Huppmann, A. R., Xi, L., Raffeld, M., Pittaluga, S. & Jaffe, E. S. Subcutaneous panniculitis-like T-cell lymphoma in the pediatric age group: a lymphoma of low malignant potential. Pediatr. Blood Cancer 60, 1165–1170 (2013).
Willemze, R. Cutaneous lymphomas with a panniculitic presentation. Semin. Diagn. Pathol. 34, 36–43 (2017).
Pincus, L. B. et al. Subcutaneous panniculitis-like T-cell lymphoma with overlapping clinicopathologic features of lupus erythematosus: coexistence of 2 entities? Am. J. Dermatopathol. 31, 520–526 (2009).
Oschlies, I. et al. Subcutaneous panniculitis-like T-cell lymphoma in children: a detailed clinicopathological description of 11 multifocal cases with a high frequency of haemophagocytic syndrome. Br. J. Dermatol. 172, 793–797 (2015).
Michonneau, D. et al. Subcutaneous panniculitis-like T-cell lymphoma: immunosuppressive drugs induce better response than polychemotherapy. Acta Dermato-Venereol. 97, 358–364 (2017).
Gau, J. P. et al. Subcutaneous panniculitis-like T cell lymphoma: familial aggregation while different response to chemotherapy. Int. J. Hematol. 89, 63–65 (2009).
Berg, K. D. et al. Transmission of a T-cell lymphoma by allogeneic bone marrow transplantation. N. Engl. J. Med. 345, 1458–1463 (2001).
Monney, L. et al. Th1-specific cell surface protein Tim-3 regulates macrophage activation and severity of an autoimmune disease. Nature 415, 536–541 (2002).
Sabatos, C. A. et al. Interaction of Tim-3 and Tim-3 ligand regulates T helper type 1 responses and induction of peripheral tolerance. Nat. Immunol. 4, 1102–1110 (2003).
Sanchez-Fueyo, A. et al. Tim-3 inhibits T helper type 1-mediated auto- and alloimmune responses and promotes immunological tolerance. Nat. Immunol. 4, 1093–1101 (2003).
Cao, E. et al. T cell immunoglobulin mucin-3 crystal structure reveals a galectin-9-independent ligand-binding surface. Immunity 26, 311–321 (2007).
Hudjashov, G. et al. Investigating the origins of eastern Polynesians using genome-wide data from the Leeward Society Isles. Sci. Rep. 8, 1823 (2018).
Anderson, A. C., Joller, N. & Kuchroo, V. K. Lag-3, Tim-3, and TIGIT: co-inhibitory receptors with specialized functions in immune regulation. Immunity 44, 989–1004 (2016).
Das, M., Zhu, C. & Kuchroo, V. K. Tim-3 and its role in regulating anti-tumor immunity. Immunol. Rev. 276, 97–111 (2017).
Kayser, M. et al. Genome-wide analysis indicates more Asian than Melanesian ancestry of Polynesians. Am. J. Hum. Genet. 82, 194–198 (2008).
Schymkowitz, J. et al. The FoldX web server: an online force field. Nucleic Acids Res. 33, W382–W388 (2005).
Pachlopnik Schmid, J. et al. Inherited defects in lymphocyte cytotoxic activity. Immunol. Rev. 235, 10–23 (2010).
Takada, H. et al. Increased serum levels of interferon-γ-inducible protein 10 and monokine induced by gamma interferon in patients with haemophagocytic lymphohistiocytosis. Clin. Exp. Immunol. 133, 448–453 (2003).
Wang, W. et al. Negative regulation of Nod-like receptor protein 3 inflammasome activation by T cell Ig mucin-3 protects against peritonitis. Immunology 153, 71–83 (2018).
Sepulveda, F. E. & de Saint Basile, G. Hemophagocytic syndrome: primary forms and predisposing conditions. Curr. Opin. Immunol. 49, 20–26 (2017).
Gautron, A.-S., Dominguez-Villar, M., de Marcken, M. & Hafler, D. A. Enhanced suppressor function of TIM-3+FoxP3+ regulatory T cells. Eur. J. Immunol. 44, 2703–2711 (2014).
Greil, R., Hutterer, E., Hartmann, T. N. & Pleyer, L. Reactivation of dormant anti-tumor immunity—a clinical perspective of therapeutic immune checkpoint modulation. Cell Commun. Signal. 15, 5 (2017).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Li, H. & Durbin, R. Fast and accurate long-read alignment with Burrows-Wheeler transform. Bioinformatics 26, 589–595 (2010).
McKenna, A. et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
Tan, A., Abecasis, G. R. & Kang, H. M. Unified representation of genetic variants. Bioinformatics 31, 2202–2204 (2015).
Cingolani, P. et al. A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strain w 1118; iso-2; iso-3. Fly 6, 80–92 (2012).
Paila, U., Chapman, B. A., Kirchner, R. & Quinlan, A. R. GEMINI: integrative exploration of genetic variation and genome annotations. PLoS Comput. Biol. 9, e1003153 (2013).
1000 Genomes Project Consortium. A map of human genome variation from population-scale sequencing. Nature 467, 1061–1073 (2010).
Hamel, N. et al. On the origin and diffusion of BRCA1 c.5266dupC (5382insC) in European populations. Eur. J. Hum. Genet. 19, 300–306 (2011).
Cote, M. et al. Munc18-2 deficiency causes familial hemophagocytic lymphohistiocytosis type 5 and impairs cytotoxic granule exocytosis in patient NK cells. J. Clin. Investig. 119, 3765–3773 (2009).
Acknowledgements
This work was funded in part by the Foundation of Stars. N.J. is a member of the Penny Cole Laboratory and the recipient of a Chercheur Boursier, Chaire de Recherche Award from the Fonds de la Recherche du Québec en Santé. D.-A.K.-Q. is a Herman Clinical Fellow. We thank N. Brousse and the Groupe Français d’Etude des Lymphomes Cutanés for their valuable contributions. We also thank S. Lade from the Peter MacCallum Cancer Centre and the Victorian Comprehensive Cancer Centre, Melbourne; P. McKelvie from the Department of Pathology at St. Vincent’s Hospital Melbourne; D. MacGregor from the Department of Pathology at the Royal Children’s Hospital (RCH) Melbourne; L. Dalla-Pozza from the Cancer Center for Children at The Children’s Hospital at Westmead, Sydney; C. Picard from the Study Center of Immunodeficiencies (CEDI) at Hôpital-Necker Enfants Malades, Paris; and G. Ménasché from INSERM U1163 at Institut Imagine, Paris. The Children’s Cancer Centre Tissue Bank at RCH Melbourne runs thanks to the generous support of Cancer in Kids @ RCH, Leukaemia Auxiliary at RCH, the Murdoch Children’s Research Institute, and the RCH Foundation. The Tumour Bank at the Children’s Hospital at Westmead is generously supported by the Kids Cancer Project. W.D.F. is funded by the Canadian Institute for Health Research (FDN-148390). The G.d.S.B. lab (INSERM 1163 at Institut Imagine) is Equipe labélisée Fondation pour la Recherche Médicale (FRM: DEQ20150734354) and is supported by l’Agence Nationale de la Recherche (ANR-12-BSV1-0020-01 and the Investissements d’Avenir program) and by the Fondation Imagine. A.F. is supported by Collège de France. T.G. and J.P. are recipients of Canadian Institutes of Health Research and Fonds de Recherche du Québec postdoctoral fellowships, respectively. A.B. is a recipient of a Fonds de Recherche du Québec doctoral studentship. E.T.V. was granted funding by Fundação de Amparo a Pesquisa do Estado de São Paulo (FAPESP, grant no. 2015/20142-0/Brazil), as part of a research fellowship abroad program to participate in this project. We thank M. Fu and S.-B. Feng of the Molecular Imaging facility at the Research Institute of the McGill University Health Centre for their assistance with the confocal microscope.
Author information
Authors and Affiliations
Contributions
T.G., D.-A.K.-Q., J.P., E.T.V., A.G., S.K., F.S., L.G.M., N.H., M.T., A.B., S.D., D.D., D.S., and F.G. performed experiments and analyzed data. H.N., J.M., W.D.F., S.G., and R.D. analyzed data. D.Moshous, J.C., S.A., C.B.-F., P.N., B.B.-M., D.Mitchell, C.T., M.Battistella, V.-H.N., R.C., J.-S.D., C.M., H.M.P., M.Besnard, S.B., P.G.E., S.F., and A.F. contributed materials and analyzed data. F.E.S., B.N., D.Michonneau, G.d.S.B., and N.J. conceived and designed the experiments and analyzed data. All authors contributed to the written manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1
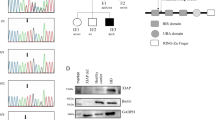
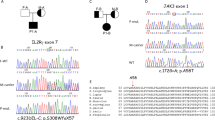
Sequencing chromatograms showing HAVCR2 mutations from homozygous affected individuals and heterozygous healthy relatives compared to wild-type HAVCR2 from a healthy control.
Supplementary Figure 2 Frequencies of the Y82C and I97M variants in different populations according to their ethnic origin.
(a), Distributions of the Y82C and I97M variants across all gnomAD populations. (b), Frequency of the Y82C variant in the Tahiti population from Polynesia. The chi-squared test was used to calculate Hardy–Weinberg equilibrium (HWE) P values.
Supplementary Figure 3 Expression of TIM-3 mutants in SPTCL patients and stably transfected HEK-293 cells.
(a), Flow cytometry of TIM-3 expression on monocytes (CD14+) from a Y82C TIM-3 heterozygous individual shows an intermediate level of expression as compared to TIM-3 expression on patient and control cells. One representative experiment of the four heterozygous individuals analyzed is provided. (b), Complete image of the western blot shown in Fig. 3c.
Supplementary Figure 4 Y82C and I97M lead to TIM-3 protein misfolding and impaired surface expression.
(a), Plot showing the impaired surface expression of the two TIM-3 mutants on the same HEK-293T cells as in Fig. 3d, using flow cytometry analysis, by APC-conjugated TIM-3 antibody or isotype control antibody. (b), Cell lysates from wild-type and Y82C or I97M TIM-3-transfected HEK-293 cells were put on a 7.5% native gel and immunoblotted against α-FLAG. Misfolded aggregates are highlighted with arrows. (c), The same lysates from (b) were submitted to an N-deglycosylation assay using the PNGase F enzyme and following the manufacturer’s denaturation protocol. These lysates as well as control lysates (without PNGase F) were immunoblotted with α-FLAG and α-β-actin. Experiments in (b) and (c) were repeated three times, and representative gels are shown. (d), Analysis of the wild-type and mutant (Y82C and I97M) bands before and after PNGase F treatment. MW, molecular weight.
Supplementary Figure 5 Cytotoxic lymphocytes from P4 display similar degranulation activity to control lymphocytes.
One representative experiment over four patients analyzed (P1–P4) is shown.
Supplementary Figure 6 Increased secretion of inflammatory cytokines by TIM-3-deficient leukocytes.
(a), Cytokine production by T lymphoblasts from Y82C TIM-3 patients as compared to controls. TNF-α, IL-2 and IFN-γ were quantified by CBA (*P < 0.05). Data are means +/− s.d. from three different patients (P3, P4, P11). Data represent the results of different independent experiments performed with different biological samples (cell cultures from cells obtained at different time points) obtained from different controls and three different TIM-3 Y82C patients. An unpaired t test was used to determine statistical significance between control (n = 15) and Y82C (n = 10) samples. b,c, Increased TNF-α production (b) and IL-1β production (c) by macrophages from patients (P3 and P11) with Y82C TIM-3, as compared to wild-type TIM-3 controls, in response to TLR4 (LPS) and NLRP3 (nigericin) activation (as done in Fig. 4). Graphs represent means +/− s.e.m. from two independent experiments in duplicate (*P < 0.05).
Supplementary Figure 7 FOXP3+CD4+ Treg cells are significantly decreased in panniculitis from TIM-3-mutant (Y82C, I97M) SPTCL as compared to TIM-3-wild-type SPTCL.
Immunofluorescence using anti-CD4 and anti-FOXP3 antibodies was performed on consecutive slides from a wild-type (WT) SPTCL sample for TIM-3 (left) or the same slide for Y82C (middle) and I97M (right) TIM-3-mutant SPTCL samples. Cells positive for FOXP3 and CD4 staining were counted using Fiji software, and graphs represent means +/− s.e.m. for CD4+ and FOXP3+CD4+ cells from five different fields (lower left). A significantly decreased number of CD4+ and FOXP3+CD4+ cells in TIM-3-mutant SPTCL was observed when comparing TIM-3-mutant to TIM-3-wild-type SPTCL (P < 0.001, one-way ANOVA). Representative images mounted using Photoshop are shown.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 and Supplementary Note
Supplementary Table 1
Clinical information on the 27 SPTCL patients analyzed in this study
Supplementary Table 2
Summary of whole-exome sequencing metrics for 17 SPTCL patients
Supplementary Table 3
List of the variants identified in exome datasets of 17 SPTCL cases, including 7 tumor–normal pairs, 7 tumors only and 3 germline-only samples
Rights and permissions
About this article
Cite this article
Gayden, T., Sepulveda, F.E., Khuong-Quang, DA. et al. Germline HAVCR2 mutations altering TIM-3 characterize subcutaneous panniculitis-like T cell lymphomas with hemophagocytic lymphohistiocytic syndrome. Nat Genet 50, 1650–1657 (2018). https://doi.org/10.1038/s41588-018-0251-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-018-0251-4
This article is cited by
-
Lymphoma-associated hemophagocytic lymphohistiocytosis (LA-HLH): a scoping review unveils clinical and diagnostic patterns of a lymphoma subgroup with poor prognosis
Leukemia (2024)
-
Diagnostic and prognostic molecular pathology of lymphoid malignancies
Virchows Archiv (2024)
-
Germline HAVCR2/TIM-3 Checkpoint Inhibitor Receptor Deficiency in Recurrent Autoinflammatory Myocarditis
Journal of Clinical Immunology (2024)
-
Post-transplant Inflammatory Bowel Disease Associated with Donor-Derived TIM-3 Deficiency
Journal of Clinical Immunology (2024)
-
Subcutaneous panniculitis-like T-cell lymphoma associated with hemophagocytic lymphohistiocytosis: a systematic review of 63 patients reported in the literature
Clinical and Experimental Medicine (2023)