Abstract
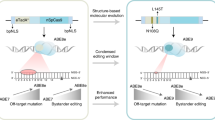
Adenine base editors (ABEs) are precise gene-editing agents that convert A:T pairs into G:C through a deoxyinosine intermediate. Existing ABEs function most effectively when the target A is in a TA context. Here we evolve the Escherichia coli transfer RNA-specific adenosine deaminase (TadA) to generate TadA8r, which extends potent deoxyadenosine deamination to RA (R = A or G) and is faster in processing GA than TadA8.20 and TadA8e, the two most active TadA variants reported so far. ABE8r, comprising TadA8r and a Streptococcus pyogenes Cas9 nickase, expands the editing window at the protospacer adjacent motif-distal end and outperforms ABE7.10, ABE8.20 and ABE8e in correcting disease-associated G:C-to-A:T transitions in the human genome, with a controlled off-target profile. We show ABE8r-mediated editing of clinically relevant sites that are poorly accessed by existing editors, including sites in PCSK9, whose disruption reduces low-density lipoprotein cholesterol, and ABCA4-p.Gly1961Glu, the most frequent mutation in Stargardt disease.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All next-generation sequencing data have been deposited to the NCBI’s Gene Expression Omnibus and can be accessed through accession no. GSE243181 (ref. 65). Amplicon sequencing data have been deposited to the NCBI Sequence Read Archive under BioProject no. PRJNA925224.
References
Landrum, M. J. et al. ClinVar: public archive of relationships among sequence variation and human phenotype. Nucleic Acids Res. 42, D980–D985 (2014).
Landrum, M. J. et al. ClinVar: public archive of interpretations of clinically relevant variants. Nucleic Acids Res. 44, D862–D868 (2016).
Gaudelli, N. M. et al. Programmable base editing of A•T to G•C in genomic DNA without DNA cleavage. Nature 551, 464–471 (2017).
Rees, H. A. & Liu, D. R. Base editing: precision chemistry on the genome and transcriptome of living cells. Nat. Rev. Genet. 19, 770–788 (2018).
Zeng, Y. et al. Correction of the Marfan syndrome pathogenic FBN1 mutation by base editing in human cells and heterozygous embryos. Mol. Ther. 26, 2631–2637 (2018).
Ryu, S. M. et al. Adenine base editing in mouse embryos and an adult mouse model of Duchenne muscular dystrophy. Nat. Biotechnol. 36, 536–539 (2018).
Liu, Z. et al. Highly efficient RNA-guided base editing in rabbit. Nat. Commun. 9, 2717 (2018).
Song, C. Q. et al. Adenine base editing in an adult mouse model of tyrosinaemia. Nat. Biomed. Eng. 4, 125–130 (2020).
Li, C. et al. Expanded base editing in rice and wheat using a Cas9-adenosine deaminase fusion. Genome Biol. 19, 59 (2018).
Hua, K., Tao, X., Yuan, F., Wang, D. & Zhu, J. K. Precise A•T to G•C base editing in the rice genome. Mol. Plant 11, 627–630 (2018).
Yan, F. et al. Highly efficient A.T to G.C base editing by Cas9n-guided tRNA adenosine deaminase in rice. Mol. Plant 11, 631–634 (2018).
Koblan, L. W. et al. In vivo base editing rescues Hutchinson–Gilford progeria syndrome in mice. Nature 589, 608–614 (2021).
Newby, G. A. et al. Base editing of haematopoietic stem cells rescues sickle cell disease in mice. Nature 595, 295–302 (2021).
Arbab, M. et al. Base editing rescue of spinal muscular atrophy in cells and in mice. Science 380, eadg6518 (2023).
Musunuru, K. et al. In vivo CRISPR base editing of PCSK9 durably lowers cholesterol in primates. Nature 593, 429–434 (2021).
Rothgangl, T. et al. In vivo adenine base editing of PCSK9 in macaques reduces LDL cholesterol levels. Nat. Biotechnol. 39, 949–957 (2021).
Zhang, W. et al. Multiplex precise base editing in cynomolgus monkeys. Nat. Commun. 11, 2325 (2020).
Lee, R. G. et al. Efficacy and safety of an investigational single-course CRISPR base editing therapy targeting PCSK9 in non-human primate and mouse models. Circulation 147, 242–253 (2023).
Wolf, J., Gerber, A. P. & Keller, W. TadA, an essential tRNA-specific adenosine deaminase from Escherichia coli. EMBO J. 21, 3841–3851 (2002).
Koblan, L. W. et al. Improving cytidine and adenine base editors by expression optimization and ancestral reconstruction. Nat. Biotechnol. 36, 843–846 (2018).
Arbab, M. et al. Determinants of base editing outcomes from target library analysis and machine learning. Cell 182, 463–480 (2020).
Song, M. et al. Sequence-specific prediction of the efficiencies of adenine and cytosine base editors. Nat. Biotechnol. 38, 1037–1043 (2020).
Kim, N. et al. Deep learning models to predict the editing efficiencies and outcomes of diverse base editors. Nat. Biotechnol. https://doi.org/10.1038/s41587-023-01792-x (2023).
Gaudelli, N. M. et al. Directed evolution of adenine base editors with increased activity and therapeutic application. Nat. Biotechnol. 38, 892–900 (2020).
Richter, M. F. et al. Phage-assisted evolution of an adenine base editor with improved Cas domain compatibility and activity. Nat. Biotechnol. 38, 883–891 (2020).
Li, J. et al. Structure-guided engineering of adenine base editor with minimized RNA off-targeting activity. Nat. Commun. 12, 2287 (2021).
Fang, G. et al. Genome-wide mapping of methylated adenine residues in pathogenic Escherichia coli using single-molecule real-time sequencing. Nat. Biotechnol. 30, 1232–1239 (2012).
Marinus, M. G. & Lobner-Olesen, A. DNA methylation. EcoSal Plus https://doi.org/10.1128/ecosalplus.ESP-0003-2013 (2014).
Losey, H. C., Ruthenburg, A. J. & Verdine, G. L. Crystal structure of Staphylococcus aureus tRNA adenosine deaminase TadA in complex with RNA. Nat. Struct. Mol. Biol. 13, 153–159 (2006).
Cadwell, R. C. & Joyce, G. F. Randomization of genes by PCR mutagenesis. PCR Methods Appl. 2, 28–33 (1992).
Grunewald, J. et al. CRISPR DNA base editors with reduced RNA off-target and self-editing activities. Nat. Biotechnol. 37, 1041–1048 (2019).
Thuronyi, B. W. et al. Continuous evolution of base editors with expanded target compatibility and improved activity. Nat. Biotechnol. 37, 1070–1079 (2019).
Kleinstiver, B. P. et al. Engineered CRISPR-Cas9 nucleases with altered PAM specificities. Nature 523, 481–485 (2015).
Nishimasu, H. et al. Engineered CRISPR-Cas9 nuclease with expanded targeting space. Science 361, 1259–1262 (2018).
Miller, S. M. et al. Continuous evolution of SpCas9 variants compatible with non-G PAMs. Nat. Biotechnol. 38, 471–481 (2020).
Walton, R. T., Christie, K. A., Whittaker, M. N. & Kleinstiver, B. P. Unconstrained genome targeting with near-PAMless engineered CRISPR-Cas9 variants. Science 368, 290–296 (2020).
Wang, X. et al. Cas12a base editors induce efficient and specific editing with low DNA damage response. Cell Rep. 31, 107723 (2020).
Ran, F. A. et al. In vivo genome editing using Staphylococcus aureus Cas9. Nature 520, 186–191 (2015).
Kleinstiver, B. P. et al. Broadening the targeting range of Staphylococcus aureus CRISPR-Cas9 by modifying PAM recognition. Nat. Biotechnol. 33, 1293–1298 (2015).
Zetsche, B. et al. Cpf1 is a single RNA-guided endonuclease of a class 2 CRISPR-Cas system. Cell 163, 759–771 (2015).
Kleinstiver, B. P. et al. Engineered CRISPR-Cas12a variants with increased activities and improved targeting ranges for gene, epigenetic and base editing. Nat. Biotechnol. 37, 276–282 (2019).
Tsai, S. Q. et al. GUIDE-seq enables genome-wide profiling of off-target cleavage by CRISPR-Cas nucleases. Nat. Biotechnol. 33, 187–197 (2015).
Liang, P. et al. Genome-wide profiling of adenine base editor specificity by EndoV-seq. Nat. Commun. 10, 67 (2019).
Jin, S. et al. Cytosine, but not adenine, base editors induce genome-wide off-target mutations in rice. Science 364, 292–295 (2019).
Zuo, E. et al. Cytosine base editor generates substantial off-target single-nucleotide variants in mouse embryos. Science 364, 289–292 (2019).
Doman, J. L., Raguram, A., Newby, G. A. & Liu, D. R. Evaluation and minimization of Cas9-independent off-target DNA editing by cytosine base editors. Nat. Biotechnol. 38, 620–628 (2020).
Yu, Y. et al. Cytosine base editors with minimized unguided DNA and RNA off-target events and high on-target activity. Nat. Commun. 11, 2052 (2020).
Wang, L. et al. Eliminating base-editor-induced genome-wide and transcriptome-wide off-target mutations. Nat. Cell Biol. 23, 552–563 (2021).
Jin, S. et al. Rationally designed APOBEC3B cytosine base editors with improved specificity. Mol. Cell 79, 728–740 e726 (2020).
Rees, H. A., Wilson, C., Doman, J. L. & Liu, D. R. Analysis and minimization of cellular RNA editing by DNA adenine base editors. Sci. Adv. 5, eaax5717 (2019).
Grunewald, J. et al. Transcriptome-wide off-target RNA editing induced by CRISPR-guided DNA base editors. Nature 569, 433–437 (2019).
Zhou, C. et al. Off-target RNA mutation induced by DNA base editing and its elimination by mutagenesis. Nature 571, 275–278 (2019).
Sanchez-Rivera, F. J. et al. Base editing sensor libraries for high-throughput engineering and functional analysis of cancer-associated single nucleotide variants. Nat. Biotechnol. 40, 862–873 (2022).
Rivera, A. et al. A comprehensive survey of sequence variation in the ABCA4 (ABCR) gene in Stargardt disease and age-related macular degeneration. Am. J. Hum. Genet. 67, 800–813 (2000).
Fujinami, K. et al. Detailed genetic characteristics of an international large cohort of patients with Stargardt disease: ProgStar study report 8. Br. J. Ophthalmol. 103, 390–397 (2019).
Muller, A. et al. High-efficiency base editing for Stargardt disease in mice, non-human primates, and human retina tissue. Preprint at bioRxiv https://doi.org/10.1101/2023.04.17.535579 (2023).
Anzalone, A. V. et al. Search-and-replace genome editing without double-strand breaks or donor DNA. Nature 576, 149–157 (2019).
Xiao, Y. L. et al. Transcriptome-wide profiling and quantification of N6-methyladenosine by enzyme-assisted adenosine deamination. Nat. Biotechnol. 41, 993–1003 (2023).
Lapinaite, A. et al. DNA capture by a CRISPR-Cas9-guided adenine base editor. Science 369, 566–571 (2020).
Ranzau, B. L., Rallapalli, K. L., Evanoff, M., Paesani, F. & Komor, A. C. The wild-type tRNA adenosine deaminase enzyme TadA is capable of sequence-specific DNA base editing. ChemBioChem 24, e202200788 (2023).
Kohli, R. M. et al. A portable hot spot recognition loop transfers sequence preferences from APOBEC family members to activation-induced cytidine deaminase. J. Biol. Chem. 284, 22898–22904 (2009).
Carpenter, M. A., Rajagurubandara, E., Wijesinghe, P. & Bhagwat, A. S. Determinants of sequence-specificity within human AID and APOBEC3G. DNA Repair 9, 579–587 (2010).
Wang, M., Rada, C. & Neuberger, M. S. Altering the spectrum of immunoglobulin V gene somatic hypermutation by modifying the active site of AID. J. Exp. Med. 207, 141–153 (2010).
Clement, K. et al. CRISPResso2 provides accurate and rapid genome editing sequence analysis. Nat. Biotechnol. 37, 224–226 (2019).
Xiao, Y. L., Wu, Y. & Tang, W. Directed evolution of an adenine base editor with increased activity and context compatibility. Gene Expression Omnibus https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE243181 (2023).
Acknowledgements
We thank H. Yan for optimizing the transfection workflow and K. M. Watters for scientific editing of the paper. This work was completed in part with computing resources provided by the University of Chicago Research Computing Center. We thank the Single Cell Immunophenotyping Core Facility at the University of Chicago for sequencing support. W.T. is supported by the Searle Scholars Program (grant no. SSP-2021-113), the Cancer Research Foundation Young Investigator Program, the American Cancer Society (grant no. RSG-22-043-01-ET) and the David & Lucile Packard Foundation (grant no. 2022-74685).
Author information
Authors and Affiliations
Contributions
Y.-L.X. and W.T. conceived and designed the study. Y.-L.X. carried out directed evolution, purified and characterized TadA8r, constructed the paired sgRNA-target library and evaluated TadA8r in human cells. Y.W. assisted with deaminase purification and characterization. Y.W. analyzed transcriptome-wide off-target effects for all ABEs and editing data generated using the paired sgRNA-target library. W.T. supervised the study. Y.-L.X., Y.W. and W.T. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
A patent has been filed for TadA8r and its applications in gene editing by the University of Chicago.
Peer review
Peer review information
Nature Biotechnology thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Notes 1–4, Tables 1–10 and Figs. 1–39.
Supplementary Table 1
Evaluated ABE targets of clinical relevance_full table.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Xiao, YL., Wu, Y. & Tang, W. An adenine base editor variant expands context compatibility. Nat Biotechnol (2024). https://doi.org/10.1038/s41587-023-01994-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41587-023-01994-3