Abstract
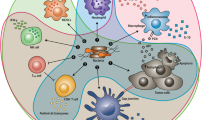
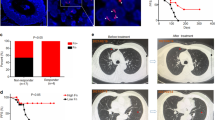
Increasing evidence implicates the tumor microbiota as a factor that can influence cancer progression. In patients with colorectal cancer (CRC), we found that pre-resection antibiotics targeting anaerobic bacteria substantially improved disease-free survival by 25.5%. For mouse studies, we designed an antibiotic silver-tinidazole complex encapsulated in liposomes (LipoAgTNZ) to eliminate tumor-associated bacteria in the primary tumor and liver metastases without causing gut microbiome dysbiosis. Mouse CRC models colonized by tumor-promoting bacteria (Fusobacterium nucleatum spp.) or probiotics (Escherichia coli Nissle spp.) responded to LipoAgTNZ therapy, which enabled more than 70% long-term survival in two F. nucleatum-infected CRC models. The antibiotic treatment generated microbial neoantigens that elicited anti-tumor CD8+ T cells. Heterologous and homologous bacterial epitopes contributed to the immunogenicity, priming T cells to recognize both infected and uninfected tumors. Our strategy targets tumor-associated bacteria to elicit anti-tumoral immunity, paving the way for microbiome–immunotherapy interventions.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The 16S rRNA data are available through the National Center for Biotechnology Information (NCBI) Sequence Read Archive (accession numbers SRR23197060–SRR23197067, BioProject PRJNA926798). The whole-exome sequencing data are available through the Sequence Read Archive (BioProject PRJNA926643). The genome of Fusobacterium nucleatum subsp. nucleatum ATCC 25586 (GCA_000007325.1) is available on KEGG (https://www.genome.jp/kegg-bin/show_organism?org=fnu). Mus musculus genome (GRCm39 reference annotation release 109) is available at the NCBI (https://www.ncbi.nlm.nih.gov/genome/annotation_euk/Mus_musculus/109/). Fusobacterium nucleatum reference proteome (UP000002521_190304) is available at the European Bioinformatics Institute (https://www.ebi.ac.uk/reference_proteomes/). All other data supporting the findings of this study are available from the corresponding authors upon reasonable request. Source data are provided with this paper.
Code availability
The distance map for FISH images analysis is available on Zenodo (https://doi.org/10.5281/zenodo.8200515)78.
References
Bodinier, M. et al. Efficient detection and immunomagnetic sorting of specific T cells using multimers of MHC class I and peptide with reduced CD8 binding. Nat. Med. 6, 707–710 (2000).
Alard, E. et al. Advances in anti-cancer immunotherapy: CAR-T cell, checkpoint inhibitors, dendritic cell vaccines, and oncolytic viruses, and emerging cellular and molecular targets. Cancers (Basel) 12, 1826 (2020).
Le, D. T. et al. PD-1 blockade in tumors with mismatch-repair deficiency. N. Engl. J. Med. 372, 2509–2520 (2015).
Nejman, D. et al. The human tumor microbiome is composed of tumor type-specific intracellular bacteria. Science 368, 973–980 (2020).
Fluckiger, A. et al. Cross-reactivity between tumor MHC class I-restricted antigens and an enterococcal bacteriophage. Science 369, 936–942 (2020).
Kalaora, S. et al. Identification of bacteria-derived HLA-bound peptides in melanoma. Nature 592, 138–143 (2021).
Yang, Y. et al. Focused specificity of intestinal TH17 cells towards commensal bacterial antigens. Nature 510, 152–156 (2014).
Zheng, D.-W. et al. Phage-guided modulation of the gut microbiota of mouse models of colorectal cancer augments their responses to chemotherapy. Nat. Biomed. Eng. 3, 717–728 (2019).
Goodman, A. L. & Gordon, J. I. Our unindicted coconspirators: human metabolism from a microbial perspective. Cell Metab. 12, 111–116 (2010).
Kostic, A. D. et al. Genomic analysis identifies association of Fusobacterium with colorectal carcinoma. Genome Res. 22, 292–298 (2012).
Sepich-Poore, G. D. et al. The microbiome and human cancer. Science 371, eabc4552 (2021).
Derosa, L. et al. Microbiota-centered interventions: the next breakthrough in immuno-oncology? Cancer Discov. 11, 2396–2412 (2021).
Song, M., Chan, A. T. & Sun, J. Influence of the gut microbiome, diet, and environment on risk of colorectal cancer. Gastroenterology 158, 322–340 (2020).
Poore, G. D. et al. Microbiome analyses of blood and tissues suggest cancer diagnostic approach. Nature 579, 567–574 (2020).
Rousseau, B. et al. Impact of antibiotics (ATB) on the recurrence of resected colorectal cancer (CRC): results of EVADER-1 a nation-wide pharmacoepidemiologic study. J. Clin. Oncol. 38, 4106 (2020).
Sargent, D. J. et al. A pooled analysis of adjuvant chemotherapy for resected colon cancer in elderly patients. N. Engl. J. Med. 345, 1091–1097 (2001).
Castellarin, M. et al. Fusobacterium nucleatum infection is prevalent in human colorectal carcinoma. Genome Res. 22, 299–306 (2012).
Bullman, S. et al. Analysis of Fusobacterium persistence and antibiotic response in colorectal cancer. Science 358, 1443–1448 (2017).
Zhou, Z., Chen, J., Yao, H. & Hu, H. Fusobacterium and colorectal cancer. Front. Oncol. 8, 371 (2018).
Brennan, C. A. & Garrett, W. S. Fusobacterium nucleatum—symbiont, opportunist and oncobacterium. Nat. Rev. Microbiol. 17, 156–166 (2019).
Sun, Y. et al. Adaptive-guided-coupling-probability level set for retinal layer segmentation. IEEE J. Biomed. Health Inform. 24, 3236–3247 (2020).
Canale, F. P. et al. Metabolic modulation of tumours with engineered bacteria for immunotherapy. Nature 598, 662–666 (2021).
Wilson, M. R. et al. The human gut bacterial genotoxin colibactin alkylates DNA. Science 363, eaar7785 (2019).
Riquelme, E. et al. Tumor microbiome diversity and composition influence pancreatic cancer outcomes. Cell 178, 795–8062 (2019).
Ahn, J. et al. Human gut microbiome and risk for colorectal cancer. J. Natl Cancer Inst. 105, 1907–1911 (2013).
Pinato, D. J. et al. Association of prior antibiotic treatment with survival and response to immune checkpoint inhibitor therapy in patients with cancer. JAMA Oncol. 5, 1774–1778 (2019).
Arane, K. & Goldman, R. D. Fusobacterium infections in children. Can. Fam. Physician 62, 813–814 (2016).
Leitsch, D. et al. Pyruvate:ferredoxin oxidoreductase and thioredoxin reductase are involved in 5-nitroimidazole activation while flavin metabolism is linked to 5-nitroimidazole resistance in Giardia lamblia. J. Antimicrob. Chemother. 66, 1756–1765 (2011).
Upcroft, P. & Upcroft, J. A. Drug targets and mechanisms of resistance in the anaerobic protozoa. Clin. Microbiol. Rev. 14, 150–164 (2001).
Dingsdag, S. A. & Hunter, N. Metronidazole: an update on metabolism, structure–cytotoxicity and resistance mechanisms. J. Antimicrob. Chemother. 73, 265–279 (2018).
Qiu, K., Durham, P. G. & Anselmo, A. C. Inorganic nanoparticles and the microbiome. Nano Res. 11, 4936–4954 (2018).
Kim, D. et al. Selective killing of pathogenic bacteria by antimicrobial silver nanoparticle–cell wall binding domain conjugates. ACS Appl. Mater. Interfaces 10, 13317–13324 (2018).
Prabhu, S. & Poulose, E. K. Silver nanoparticles: mechanism of antimicrobial action, synthesis, medical applications, and toxicity effects. Int. Nano Lett. 2, 32 (2012).
Waszczykowska, A., Żyro, D., Ochocki, J. & Jurowski, P. Clinical application and efficacy of silver drug in ophthalmology: a literature review and new formulation of EYE drops with drug silver (I) complex of metronidazole with improved dosage form. Biomedicines 9, 210 (2021).
Starek, M., Dąbrowska, M., Chebda, J., Żyro, D. & Ochocki, J. Stability of metronidazole and its complexes with silver(I) salts under various stress conditions. Molecules 26, 3582 (2021).
Radko, L., Stypuła-Trębas, S., Posyniak, A., Żyro, D. & Ochocki, J. Silver(I) complexes of the pharmaceutical agents metronidazole and 4-hydroxymethylpyridine: comparison of cytotoxic profile for potential clinical application. Molecules (Basel) 24, 1949 (2019).
Vander Heiden, M. G., Cantley, L. C. & Thompson, C. B. Understanding the Warburg effect: the metabolic requirements of cell proliferation. Science 324, 1029–1033 (2009).
Yu, J. et al. Remote loading paclitaxel–doxorubicin prodrug into liposomes for cancer combination therapy. Acta Pharm. Sin. B 10, 1730–1740 (2020).
Guo, G., Guo, T. & Chang, X. Effect of CYP3A4 genetic polymorphisms on pharmacokinetics of tinidazole. Journal of Chinese Pharmaceutical Sciences 29, 272–279 (2020).
Sivanesan, S. et al. Pharmacokinetics of the individual major components of polymyxin B and colistin in rats. J. Nat. Prod. 80, 225–229 (2017).
Rubinstein, M. R. et al. Fusobacterium nucleatum promotes colorectal carcinogenesis by modulating E-cadherin/β-catenin signaling via its FadA adhesin. Cell Host Microbe 14, 195–206 (2013).
Geng, F., Zhang, Y., Lu, Z., Zhang, S. & Pan, Y. Fusobacterium nucleatum caused DNA damage and promoted cell proliferation by the Ku70/p53 pathway in oral cancer cells. DNA Cell Biol. 39, 144–151 (2020).
Zhang, S. et al. Fusobacterium nucleatum promotes epithelial-mesenchymal transiton through regulation of the lncRNA MIR4435-2HG/miR-296-5p/Akt2/SNAI1 signaling pathway. FEBS J. 287, 4032–4047 (2020).
Park, E. M. et al. Targeting the gut and tumor microbiota in cancer. Nat. Med. 28, 690–703 (2022).
Smith, M. et al. Gut microbiome correlates of response and toxicity following anti-CD19 CAR T cell therapy. Nat. Med. 28, 713–723 (2022).
Lobionda, S., Sittipo, P., Kwon, H. Y. & Lee, Y. K. The role of gut microbiota in intestinal inflammation with respect to diet and extrinsic stressors. Microorganisms 7, 271 (2019).
Nie, K. et al. Roseburia intestinalis: a beneficial gut organism from the discoveries in genus and species. Front. Cell. Infect. Microbiol. 11, 757718 (2021).
Okano, K., Shimoda, T. & Matsumura, Y. Clinicopathologic and immunohistochemical study of early colorectal cancer with liver metastases. J. Gastroenterol. 34, 334–340 (1999).
Goodwin, T. J., Zhou, Y., Musetti, S. N., Liu, R. & Huang, L. Local and transient gene expression primes the liver to resist cancer metastasis. Sci. Transl. Med. 8, 364ra153 (2016).
Han, Y. K. et al. Hypoxia induces immunogenic cell death of cancer cells by enhancing the exposure of cell surface calreticulin in an endoplasmic reticulum stress-dependent manner. Oncol Lett. 18, 6269–6274 (2019).
Cao, J. et al. Heat shock protein 70 attenuates hypoxiainduced apoptosis of pulmonary microvascular endothelial cells isolated from neonatal rats. Mol. Med. Rep. 24, 690 (2021).
He, Y., Xiang, Z. & Mobley, H. L. Vaxign: the first web-based vaccine design program for reverse vaccinology and applications for vaccine development. J. Biomed. Biotechnol. 2010, 297505 (2010).
Ong, E. et al. Vaxign2: the second generation of the first Web-based vaccine design program using reverse vaccinology and machine learning. Nucleic Acids Res. 49, W671–W678 (2021).
Ali Mohammed, M. M., Pettersen, V. K., Nerland, A. H., Wiker, H. G. & Bakken, V. Label-free quantitative proteomic analysis of the oral bacteria Fusobacterium nucleatum and Porphyromonas gingivalis to identify protein features relevant in biofilm formation. Anaerobe 72, 102449 (2021).
Mohammed, M. M. A., Pettersen, V. K., Nerland, A. H., Wiker, H. G. & Bakken, V. Quantitative proteomic analysis of extracellular matrix extracted from mono- and dual-species biofilms of Fusobacterium nucleatum and Porphyromonas gingivalis. Anaerobe 44, 133–142 (2017).
Cameron, M. & Williams, H. E. Comparing compressed sequences for faster nucleotide BLAST searches. IEEE/ACM Trans. Comput. Biol. Bioinform. 4, 349–364 (2007).
Stevens, J., Wiesmüller, K.-H., Walden, P. & Joly, E. Peptide length preferences for rat and mouse MHC class I molecules using random peptide libraries. Eur. J. Immunol. 28, 1272–1279 (1998).
Gil-Cruz, C. et al. Microbiota-derived peptide mimics drive lethal inflammatory cardiomyopathy. Science 366, 881–886 (2019).
Sato, T. et al. Single Lgr5 stem cells build crypt-villus structures in vitro without a mesenchymal niche. Nature 459, 262–265 (2009).
LeBlanc, D. J., Lee, L. N. & Abu-Al-Jaibat, A. Molecular, genetic, and functional analysis of the basic replicon of pVA380-1, a plasmid of oral streptococcal origin. Plasmid 28, 130–145 (1992).
Kennedy, M. J. et al. Rapid blue-light-mediated induction of protein interactions in living cells. Nat. Methods 7, 973–975 (2010).
Krienke, C. et al. A noninflammatory mRNA vaccine for treatment of experimental autoimmune encephalomyelitis. Science 371, 145–153 (2021).
Hu, Z. et al. Personal neoantigen vaccines induce persistent memory T cell responses and epitope spreading in patients with melanoma. Nat. Med. 27, 515–525 (2021).
Munoz-Price, L. S., Frencken, J. F., Tarima, S. & Bonten, M. Handling time-dependent variables: antibiotics and antibiotic resistance. Clin. Infect. Dis. 62, 1558–1563 (2016).
Song, W. et al. Trapping of lipopolysaccharide to promote immunotherapy against colorectal cancer and attenuate liver metastasis. Adv. Mater. 30, e1805007 (2018).
Song, W. et al. Synergistic and low adverse effect cancer immunotherapy by immunogenic chemotherapy and locally expressed PD-L1 trap. Nat. Commun. 9, 2237 (2018).
Hu, M. et al. Relaxin gene delivery mitigates liver metastasis and synergizes with check point therapy. Nat. Commun. 10, 2993 (2019).
Gmur, R., Wyss, C., Xue, Y., Thurnheer, T. & Guggenheim, B. Gingival crevice microbiota from Chinese patients with gingivitis or necrotizing ulcerative gingivitis. Eur. J. Oral Sci. 112, 33–41 (2004).
Caporaso, J. G. et al. Global patterns of 16S rRNA diversity at a depth of millions of sequences per sample. Proc. Natl Acad. Sci. USA 108, 4516–4522 (2011).
Mohsen, A., Park, J., Chen, Y. A., Kawashima, H. & Mizuguchi, K. Impact of quality trimming on the efficiency of reads joining and diversity analysis of Illumina paired-end reads in the context of QIIME1 and QIIME2 microbiome analysis frameworks. BMC Bioinformatics 20, 581 (2019).
Caporaso, J. G. et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 7, 335–336 (2010).
Callahan, B. J. et al. DADA2: high-resolution sample inference from Illumina amplicon data. Nat. Methods 13, 581–583 (2016).
DeSantis, T. Z. et al. Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. Appl. Environ. Microbiol. 72, 5069–5072 (2006).
Quast, C. et al. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res. 41, D590–D596 (2013).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013).
Price, M. N., Dehal, P. S. & Arkin, A. P. FastTree 2—approximately maximum-likelihood trees for large alignments. PLoS ONE 5, e9490 (2010).
Segata, N. et al. Metagenomic biomarker discovery and explanation. Genome Biol. 12, R60 (2011).
Sun, Y. et al. Killing tumor-associated bacteria with a liposomal antibiotic generates microbial neoantigens that induce anti-tumor immune responses. (2023). https://zenodo.org/record/8200515
Acknowledgements
K.-H.L. passed away before the submission of the manuscript. The paper is dedicated in memory of him. We thank the University of North Carolina’s Department of Chemistry Mass Spectrometry Core Laboratory, especially E. D. Weatherspoon, for assistance with mass spectrometry analysis. We thank the University of North Carolina’s Department of Microscopy Services Laboratory, especially V. J. Madden, for assistance with TEM imaging. We thank the University of North Carolina’s Animal Histopathology and Laboratory Medicine Core, especially L. Wang, for assistance with histology and toxicity analysis. We thank the University of North Carolina’s Cryo-EM Core, especially J. Peck, for assistance with cryo-EM imaging. We thank the University of North Carolina’s Nanomedicines Characterization Core Facility, especially M. Sokolsky, for assistance with ICP‒MS analysis. We thank the University of North Carolina’s Microbiome Core Facility for 16S rRNA gene sequencing, the Cancer Center Support Grant (P30 CA016086) and the Center for Gastrointestinal Biology and Disease (P30 DK34987). Figures 2a, 3a, 4a, 4o and 5d and Supplementary Figs. 4a, 6b, 10h and 12a were created with BioRender. The work was supported by NIH grant CA198999 (to L.H. and A.A.), the Fred Eshelman Distinguished Professorship (to L.H.), the Institut National du Cancer (InCa), the Nuovo-Soldati Foundation, Swim Across America and, in part, through NIH/NCI Cancer Center Support Grant P30 CA008748 (to B.R., M.B.F. and O.A.). M.B.F. is funded by NIH T32‐CA009512 and an ASCO Young Investigator Award.
Author information
Authors and Affiliations
Contributions
L.H., A.A. and M.W. conceived and designed the research. L.H., M.W., K.Q., A.V., A.A., W.S., J.G., J.A., J.N. and J.P.-Y.T. designed the experiments and analyzed the data. B.R., C.L.B.-B., I.K., P.-J.B., M.H., M.F., E.D., A.H. and F.R. generated the database and did methodology, statistical analyses, models and figures for clinical data. O.A. analyzed the whole-genome sequencing data. K.Q. established the culture and plating system for F. nucleatum and modeled the growth curve versus turbidity. G.H. did sample preparation and data analysis for 16S rRNA sequencing. H.S. aligned epitopes from bacteria proteome. Y.H. ran the topology analysis for bacteria genome. Y.-Y.C., K.-H.L. and M.W profiled the pharmacokinetics of the drugs. Y.Z., Y.L. and M.W. sampled the tissues and serum for pharmacokinetic and toxicity assays. M.M. did deconvolution and reconstruction of the images. Y.S. coded for the FISH quantification. J.G., X.Z., Y.Z. and M.W. performed the surgery and in vivo mouse experiments. X.Z. and M.W. did flow cytometry analysis. L.L., P.A. and M.W. purified and analyzed T cell epitopes. M.W. and Y.Z. prepared the frozen sections and immunofluorescence, FISH staining and qPCR assays. M.W., B.R. and L.H. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
L. Huang: Consultant with PDS Biotechnology and Stemirna Therapeutics. B. Rousseau: Advisory/Consultancy, Speaker Bureau/Expert testimony: Bayer; Advisory/Consultancy, Speaker Bureau/Expert testimony: Roche; Travel/Accommodation/Expenses: Servier; Travel/Accommodation/Expenses: Astellas; Speaker Bureau/Expert testimony: Gilead.
Peer review
Peer review information
Nature Biotechnology thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Methods, Supplementary Figs. 1–16 and Supplementary Tables 1–3
Supplementary Video
Reconstruction of image of F. nucleatum-infected CT26FL3(Luc/RFP) cells
Supplementary Data
Source Data Supplementary Figs. 4 and 6–14
Source data
Source Data Fig. 1
Statistical Source Data
Source Data Fig. 5
Unprocessed western blots
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wang, M., Rousseau, B., Qiu, K. et al. Killing tumor-associated bacteria with a liposomal antibiotic generates neoantigens that induce anti-tumor immune responses. Nat Biotechnol (2023). https://doi.org/10.1038/s41587-023-01957-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41587-023-01957-8
This article is cited by
-
Targeting the tumour’s little helpers
Nature Reviews Cancer (2023)
-
Targeting tumour-associated bacteria
Nature Reviews Drug Discovery (2023)
-
Antibiotic nanoparticles boost antitumor immunity
Nature Biotechnology (2023)