Abstract
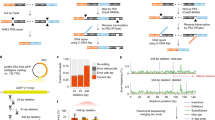
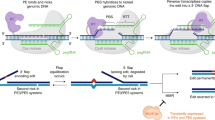
The targeted deletion, replacement, integration or inversion of genomic sequences could be used to study or treat human genetic diseases, but existing methods typically require double-strand DNA breaks (DSBs) that lead to undesired consequences, including uncontrolled indel mixtures and chromosomal abnormalities. Here we describe twin prime editing (twinPE), a DSB-independent method that uses a prime editor protein and two prime editing guide RNAs (pegRNAs) for the programmable replacement or excision of DNA sequences at endogenous human genomic sites. The two pegRNAs template the synthesis of complementary DNA flaps on opposing strands of genomic DNA, which replace the endogenous DNA sequence between the prime-editor-induced nick sites. When combined with a site-specific serine recombinase, twinPE enabled targeted integration of gene-sized DNA plasmids (>5,000 bp) and targeted sequence inversions of 40 kb in human cells. TwinPE expands the capabilities of precision gene editing and might synergize with other tools for the correction or complementation of large or complex human pathogenic alleles.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Source data are provided with this paper. High-throughput sequencing data have been deposited in the NCBI Sequence Read Archive database under accession PRJNA770428. Additional data used in the study can be accessed on Figshare (https://doi.org/10.6084/m9.figshare.c.5674708).
References
Auton, A. et al. A global reference for human genetic variation. Nature 526, 68–74 (2015).
Landrum, M. J. et al. ClinVar: public archive of relationships among sequence variation and human phenotype. Nucleic Acids Res. 42, D980–D985 (2014).
Weischenfeldt, J., Symmons, O., Spitz, F. & Korbel, J. O. Phenotypic impact of genomic structural variation: insights from and for human disease. Nat. Rev. Genet. 14, 125–138 (2013).
Cox, D. B., Platt, R. J. & Zhang, F. Therapeutic genome editing: prospects and challenges. Nat. Med. 21, 121–131 (2015).
Doudna, J. A. The promise and challenge of therapeutic genome editing. Nature 578, 229–236 (2020).
Anzalone, A. V., Koblan, L. W. & Liu, D. R. Genome editing with CRISPR–Cas nucleases, base editors, transposases and prime editors. Nat. Biotechnol. 38, 824–844 (2020).
Jinek, M. et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816–821 (2012).
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013).
Mali, P. et al. RNA-guided human genome engineering via Cas9. Science 339, 823–826 (2013).
Komor, A. C., Kim, Y. B., Packer, M. S., Zuris, J. A. & Liu, D. R. Programmable editing of a target base in genomic DNA without double-stranded DNA cleavage. Nature 533, 420–424 (2016).
Gaudelli, N. M. et al. Programmable base editing of A•T to G•C in genomic DNA without DNA cleavage. Nature 551, 464–471 (2017).
Anzalone, A. V. et al. Search-and-replace genome editing without double-strand breaks or donor DNA. Nature 576, 149–157 (2019).
Canver, M. C. et al. Characterization of genomic deletion efficiency mediated by clustered regularly interspaced short palindromic repeats (CRISPR)/Cas9 nuclease system in mammalian cells. J. Biol. Chem. 289, 21312–21324 (2014).
Suzuki, K. et al. In vivo genome editing via CRISPR/Cas9 mediated homology-independent targeted integration. Nature 540, 144–149 (2016).
Wang, B. et al. Highly efficient CRISPR/HDR-mediated knock-in for mouse embryonic stem cells and zygotes. Biotechniques 59, 201–202, 204, 206–208 (2015).
Pawelczak, K. S., Gavande, N. S., VanderVere-Carozza, P. S. & Turchi, J. J. Modulating DNA repair pathways to improve precision genome engineering. ACS Chem. Biol. 13, 389–396 (2018).
Branzei, D. & Foiani, M. Regulation of DNA repair throughout the cell cycle. Nat. Rev. Mol. Cell Biol. 9, 297–308 (2008).
Heyer, W. D., Ehmsen, K. T. & Liu, J. Regulation of homologous recombination in eukaryotes. Annu. Rev. Genet. 44, 113–139 (2010).
Gasperini, M. et al. CRISPR/Cas9-mediated scanning for regulatory elements required for HPRT1 expression via thousands of large, programmed genomic deletions. Am. J. Hum. Genet. 101, 192–205 (2017).
Kosicki, M., Tomberg, K. & Bradley, A. Repair of double-strand breaks induced by CRISPR–Cas9 leads to large deletions and complex rearrangements. Nat. Biotechnol. 36, 765–771 (2018).
Alanis-Lobato, G. et al. Frequent loss of heterozygosity in CRISPR–Cas9-edited early human embryos. Proc. Natl Acad. Sci. USA 118, e2004832117 (2021).
Song, Y. et al. Large-fragment deletions induced by Cas9 cleavage while not in the BEs system. Mol. Ther. Nucleic Acids 21, 523–526 (2020).
Brunet, E. & Jasin, M. Induction of chromosomal translocations with CRISPR–Cas9 and other nucleases: understanding the repair mechanisms that give rise to translocations. Adv. Exp. Med. Biol. 1044, 15–25 (2018).
Nahmad, A. D. et al. Frequent aneuploidy in primary human T cells following CRISPR–Cas9 cleavage. Preprint at https://www.biorxiv.org/content/10.1101/2021.08.20.457092v1.abstract (2021).
Leibowitz, M. L. et al. Chromothripsis as an on-target consequence of CRISPR–Cas9 genome editing. Nat. Genet. 53, 895–905 (2021).
Haapaniemi, E., Botla, S., Persson, J., Schmierer, B. & Taipale, J. CRISPR–Cas9 genome editing induces a p53-mediated DNA damage response. Nat. Med. 24, 927–930 (2018).
Ihry, R. J. et al. p53 inhibits CRISPR–Cas9 engineering in human pluripotent stem cells. Nat. Med. 24, 939–946 (2018).
Enache, O. M. et al. Cas9 activates the p53 pathway and selects for p53-inactivating mutations. Nat. Genet. 52, 662–668 (2020).
Merrick, C. A., Zhao, J. & Rosser, S. J. Serine integrases: advancing synthetic biology. ACS Synth. Biol. 7, 299–310 (2018).
Karpinski, J. et al. Directed evolution of a recombinase that excises the provirus of most HIV-1 primary isolates with high specificity. Nat. Biotechnol. 34, 401–409 (2016).
Chaikind, B., Bessen, J. L., Thompson, D. B., Hu, J. H. & Liu, D. R. A programmable Cas9-serine recombinase fusion protein that operates on DNA sequences in mammalian cells. Nucleic Acids Res. 44, 9758–9770 (2016).
Gaj, T. et al. Enhancing the specificity of recombinase-mediated genome engineering through dimer interface redesign. J. Am. Chem. Soc. 136, 5047–5056 (2014).
Kim, A. I. et al. Mycobacteriophage Bxb1 integrates into the Mycobacterium smegmatis groEL1 gene. Mol. Microbiol. 50, 463–473 (2003).
Choi, J. et al. Precise genomic deletions using paired prime editing. Nat. Biotechnol. https://doi.org/10.1038/s41587-020 (2021).
Lin, Q. et al. High-efficiency prime editing with optimized, paired pegRNAs in plants. Nat. Biotechnol. https://doi.org/10.1038/s41587-021-00868-w (2021).
Scriver, C. R. The PAH gene, phenylketonuria, and a paradigm shift. Hum. Mutat. 28, 831–845 (2007).
Nelson, J. W. et al. Engineered pegRNAs improve prime editing efficiency. Nat. Biotechnol. https://doi.org/10.1038/s41587-021-01039-7 (2021).
Flanigan, K. M. et al. Mutational spectrum of DMD mutations in dystrophinopathy patients: application of modern diagnostic techniques to a large cohort. Hum. Mutat. 30, 1657–1666 (2009).
Aartsma-Rus, A. et al. Development of exon skipping therapies for duchenne muscular dystrophy: a critical review and a perspective on the outstanding issues. Nucleic Acid Ther. 27, 251–259 (2017).
Kim, D. Y., Moon, S. B., Ko, J.-H., Kim, Y.-S. & Kim, D. Unbiased investigation of specificities of prime editing systems in human cells. Nucleic Acids Res. 48, 10576–10589 (2020).
Jin, S. et al. Genome-wide specificity of prime editors in plants. Nat. Biotechnol. 39, 1292–1299 (2021).
Duportet, X. et al. A platform for rapid prototyping of synthetic gene networks in mammalian cells. Nucleic Acids Res. 42, 13440–13451 (2014).
Jusiak, B. et al. Comparison of integrases identifies Bxb1-GA mutant as the most efficient site-specific integrase system in mammalian cells. ACS Synth. Biol. 8, 16–24 (2019).
Sharma, R. et al. In vivo genome editing of the albumin locus as a platform for protein replacement therapy. Blood 126, 1777–1784 (2015).
Nathwani, A. C. et al. Long-term safety and efficacy of factor IX gene therapy in hemophilia B. N. Engl. J. Med. 371, 1994–2004 (2014).
Bessen, J. L. et al. High-resolution specificity profiling and off-target prediction for site-specific DNA recombinases. Nat. Commun. 10, 1937 (2019).
Bondeson, M. L. et al. Inversion of the IDS gene resulting from recombination with IDS-related sequences is a common cause of the Hunter syndrome. Hum. Mol. Genet. 4, 615–621 (1995).
Chen, X. et al. In trans paired nicking triggers seamless genome editing without double-stranded DNA cutting. Nat. Commun. 8, 657 (2017).
Park, C. Y. et al. Targeted inversion and reversion of the blood coagulation factor 8 gene in human iPS cells using TALENs. Proc. Natl Acad. Sci. USA 111, 9253–9258 (2014).
Li, J. et al. Efficient inversions and duplications of mammalian regulatory DNA elements and gene clusters by CRISPR/Cas9. J. Mol. Cell. Biol. 7, 284–298 (2015).
Clement, K. et al. CRISPResso2 provides accurate and rapid genome editing sequence analysis. Nat. Biotechnol. 37, 224–226 (2019).
Clement, K., Farouni, R., Bauer, D. E. & Pinello, L. AmpUMI: design and analysis of unique molecular identifiers for deep amplicon sequencing. Bioinformatics 34, i202–i210 (2018).
Shen, W., Le, S., Li, Y. & Hu, F. SeqKit: a cross-platform and ultrafast toolkit for FASTA/Q file manipulation. PLoS ONE 11, e0163962 (2016).
Levy, J. M. & Nicoll, R. A. Membrane-associated guanylate kinase dynamics reveal regional and developmental specificity of synapse stability. J. Physiol. 595, 1699–1709 (2017).
Koblan, L. W. et al. In vivo base editing rescues Hutchinson–Gilford progeria syndrome in mice. Nature 589, 608–614 (2021).
Gaudelli, N. M. et al. Directed evolution of adenine base editors with increased activity and therapeutic application. Nat. Biotechnol. 38, 892–900 (2020).
Acknowledgements
We thank E. Sontheimer’s group for sharing Huh7 cells and members of the Liu Laboratory for helpful discussions. This work was supported by the Merkin Institute of Transformative Technologies in Healthcare, National Institutes of Health grants U01 AI142756, RM1 HG009490 and R35 GM118062, the Bill and Melinda Gates Foundation, and the Howard Hughes Medical Institute. A.V.A. acknowledges a Jane Coffin Childs postdoctoral fellowship through the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Contributions
A.V.A., X.D.G., C.J.P., A.T.N. and J.M.L. designed experiments. A.V.A., X.D.G., C.J.P., A.T.N., L.W.K., A.R. and J.A.M.M. performed experiments and analyzed data. A.V.A., X.D.G., C.J.P. and D.R.L. wrote the manuscript. D.R.L. supervised the research.
Corresponding author
Ethics declarations
Competing interests
D.R.L. is a consultant and equity holder of Beam Therapeutics, Prime Medicine, Pairwise Plants and Chroma Medicine, companies that use genome editing or genome engineering technologies. A.V.A., C.J.P. and J.M.L. are currently employees at Prime Medicine. A.V.A., X.D.G., C.J.P., J.M.L. and D.R.L. have filed patent applications on twinPE and prime editing through the Broad Institute.
Additional information
Peer review information Nature Biotechnology thanks the anonymous reviewers for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Twin prime editing mediates sequence replacements at CCR5.
(a) Replacement of endogenous sequence within CCR5 region 1 with a 108-bp fragment of FKBP12 cDNA using twinPE (FKBP12 sequence oriented in the forward direction,) or PE3 (FKBP12 sequence oriented in the reverse direction). For PE3 editing, pegRNA RT templates were designed to encode 108 base pairs of FKBP12 cDNA sequence and one of three different target-site homology sequence lengths. For PE3 edits, each pegRNA was tested with three nicking sgRNAs. (b) Replacement of endogenous sequence within CCR5 region 2 with a 108-bp fragment of FKBP12 cDNA sequence using twinPE (FKBP12 sequence oriented in the forward direction) or PE3 (FKBP12 sequence oriented in the reverse direction). As in (a), PE3 edits were tested with pegRNAs containing RT templates that were designed to encode 108 base pairs of FKBP12 cDNA sequence and one of three different target-site homology sequence lengths. For PE3 edits, each pegRNA was tested with three nicking sgRNAs. Values and error bars reflect the mean and s.d. of three independent biological replicates. (c) Transfection of HEK293T cells with a pair of pegRNAs targeting CCR5 leads to replacement of 53 base pairs of endogenous sequence with 113 base pairs (attB–[27-bp spacer]–attP) or 103 base pairs (attB–[27-bp spacer]–attB) of exogenous sequence. Values and error bars reflect the mean and s.d. of three independent biological replicates.
Extended Data Fig. 2 Recoding of PAH exon sequences in HEK293T cells via twinPE.
Screen of pegRNA pairs targeting PAH for recoding of (a) exons 2, 4 and 5, (b) exon 7, and (c) exons 9, 10, 11, and 12. RT templates of pegRNAs encoded partially recoded exonic sequence to optimize orthogonality to the endogenous gene sequence. For each spacer pair, nine pegRNA combinations were tested using three PBS variants for each spacer in a three-by-three matrix, with RT templates encoding the recoded exonic sequence, which was held constant for given spacer pairs. Sequences of pegRNAs are listed in Supplementary Table 1. Sequences of recoded exonic sequences are listed in Supplementary Table 4. Values in (a), (b) and exon 9 in (c) reflect single biological replicates. Values for exons 10, 11 and 12 in (c) reflect the mean of three independent biological replicates.
Extended Data Fig. 3 Installation of a 38-bp Bxb1 attB site at CCR5 with twinPE.
Spacer pairs targeting the CCR5 locus were designed for twinPE-mediated insertion of the Bxb1 attB attachment site. For each spacer, three pegRNAs were designed having three different PBS lengths and a fixed RT template that encodes the full-length Bxb1 attB sequence (38 bp). Sequences of pegRNAs are listed in Supplementary Table 1. For each spacer pair, a three-by-three matrix of pegRNA combinations was tested by plasmid DNA co-transfection with PE2 in HEK293T cells. Each pegRNA pair is specified below the x-axis. Values reflect single biological replicates.
Extended Data Fig. 4 Installation of a 50-bp Bxb1 attP site at AAVS1 with twinPE.
Spacer pairs targeting the AAVS1 locus were designed for twinPE-mediated insertion of the Bxb1 attP attachment site. For each spacer, three pegRNAs were designed having three different PBS lengths and a fixed RT template that encodes a portion (43-44 bp) of the Bxb1 attP sequence. Sequences of pegRNAs are listed in Supplementary Table 1. For each spacer pair, a three-by-three matrix of pegRNA combinations was tested by plasmid DNA co-transfection with PE2 in HEK293T cells. Each pegRNA pair is specified below the x-axis. Values reflect single biological replicates.
Extended Data Fig. 5 Comparison of twinPE and PE3 for Bxb1 attB insertion at CCR5.
(a) Replacement of endogenous sequence within CCR5 region 1 with the Bxb1 attB site using twinPE or PE3. For PE3 editing systems, pegRNA RT templates were designed to encode the Bxb1 attB sequence and one of three different target-site homology sequence lengths. For PE3 edits, each pegRNA was tested with three nicking sgRNAs. (b) Replacement of endogenous sequence within CCR5 region 2 with the Bxb1 attB sequence using twinPE or PE3. As in (a), PE3 edits were tested with pegRNAs containing RT templates that were designed to encode the Bxb1 attB sequence and one of three different target-site homology sequence lengths and tested with three nicking sgRNAs. Values and error bars in (a) and TwinPE edits, PE3 edits of CCR5_D2_23, CCR5 D2_28 with nicking guide RNA C1 and C1.5 in (b) reflect the mean and s.d. of three independent biological replicates. Values of CCR5 D2_28 with nicking guide RNA C4 and CCR5 D2_34 in (b) reflect the mean of two independent biological replicates.
Extended Data Fig. 6 TwinPE combined with Bxb1 recombinase for targeted knock-in of donor DNA plasmids.
(a) Bxb1-mediated DNA donor knock-in in clonal HEK293T cell lines. Transfection of a HEK293T clonal cell line containing homozygous attB site insertion at CCR5 with varying amounts of Bxb1-expressing plasmid and attP-containing donor DNA plasmid. Knock-in efficiency was quantified by ddPCR. Values and error bars reflect the mean of two independent biological replicates. (b) Assessment of genome-donor junction purity by high-throughput sequencing. Genomic DNA from single-transfection knock-in experiments was amplified with a forward primer that binds the genome and a reverse primer that binds within the donor plasmid (Supplementary Table 2). Values and error bars reflect the mean and s.d. of three independent biological replicates. (c) Assessment of genome-donor junction purity at the other junction by high-throughput sequencing as performed in (b). Values and error bars in 506c + 584b, 509b + 584b, 1077c + 1154c, and 3786c + 3903c reflect the mean and s.d. of three independent biological replicates. Values in 325a + 414b, 513b + 584b, 3786c + 3930c reflect the mean of two independent biological replicates. (d) Multiplexed single-transfection knock-in at AAVS1 and CCR5. HEK293T cells were transfected with plasmids encoding PE2, Bxb1, a pair of pegRNAs for the insertion of attP at AAVS1, an attB-donor, a pegRNA pair for the insertion of one of four attachment sites (attB, attB-GA, attP, or attP-GA) at CCR5, and a corresponding donor. Knock-in was observed at both target loci under all four conditions. Insertion of attP at AAVS1 and attB at CCR5 gave the lowest knock-in efficiencies overall (0.2% at AAVS1, 0.4% at CCR5). Insertion of attP at both sites yielded the highest levels of knock-in at AAVS1 (1.8%) but low levels (0.2%) at CCR5. When an orthogonal edit (attB-GA or attP-GA) was introduced at CCR5, AAVS1 knock-in was 0.7-0.8%. Higher knock-in at CCR5 was observed with attB-GA (1.4%) than with attP-GA (0.4%), consistent with our single locus knock-in results. Values and error bars reflect the mean and s.d. of three independent biological replicates. (e) and (f) Effects of reducing pegRNA overlap on twinPE efficiency and donor/pegRNA recombination. (e) The editing efficiencies of pairs of pegRNAs for insertion of Bxb1 attB at CCR5 were measured by high-throughput sequencing. The pairs differed in the amount of overlap shared between their flaps, from 38 bp (full-length attB sequence) down to 20 bp. Editing efficiency of the pairs with shorter overlaps was comparable to the pair with full-length overlap. Values and error bars reflect the mean and s.d. of three independent biological replicates. (f) Assessment of recombination between attB-containing pegRNA plasmids and attP-containing donor plasmids. Following transfection of HEK293T cells with the indicated samples, isolated DNA was amplified with a forward primer that binds the pegRNA expression plasmid (TTGAAAAAGTGGCACCGAGT) and a reverse primer that binds the donor plasmid (CTCCCACTCATGATCTA). A positive 256-bp PCR band confirms recombination between the two plasmids. When the pegRNA encodes full-length attB (38-bp) or a truncated version of attB with 30-bp of overlap between flaps, a band is observed; however, recombination is not observed when the pegRNAs encode a truncated attB with only 20-bp of flap overlap. The ‘No PE2’ control uses the 38-bp overlap pegRNA pair. No recombination is observed in the absence of Bxb1 or if the donor and pegRNA plasmids both bear attB (Mismatch, ‘M’). Three independent biological replicates were performed and a representative image from one of the replicates is shown.
Extended Data Fig. 7 Expression of human Factor IX from the ALB promoter following twinPE-recombinase knock-in and characterization of Bxb1 off-target editing.
(a) Huh7 cells were transfected with Bxb1, donor (attP-splice acceptor-cDNA of F9 exons 2-8), PE2, and pegRNAs for installation of attB in the first intron of ALB or at CCR5. Three days post-transfection, cells were split and allowed to grow to confluence. Their media was changed, and they were left to condition the fresh media, with aliquots taken at days 4, 7, and 10. Factor IX was present at detectable levels by ELISA (dashed line represents the lower limit of detection) in two of three samples treated with ALB pegRNAs at Day 4, and in all samples treated with ALB pegRNAs at Day 7 and Day 10. Factor IX was never detected in the conditioned media of any samples treated with CCR5 pegRNAs. Values and error bars reflect the mean and s.d. of two or three independent biological replicates. (b) Targeted amplicon sequencing was performed for each of the five nominated pseudo-sites (OT1-OT5) from seven different samples treated with 5.6-kb donor DNA plasmid, twinPE reagents targeting CCR5 or AAVS1, and Bxb1 recombinase. The indels in all five pseudo-sites are either below the limit of detection (<0.1%) or near-background compared to untreated controls. The integration efficiency at the on-target site was measured by ddPCR as shown in Fig. 3d (c) To capture potential donor plasmid integration events at nominated pseudo-sites, primers were used to amplify predicted integration junctions. The gel depicts PCR reactions performed for each off-target site as indicated in the above legend. Confirmation of on-target donor integration from the samples is shown in the right-most column of the gel. In (b) and (c), two or three independent biological replicates were performed.
Extended Data Fig. 8 TwinPE and Bxb1-mediated inversion in HEK293T GFP reporter cells.
(a) The lentiviral fluorescent reporter construct used to assess inversion efficiency with twinPE and Bxb1 recombinase. The reporter contains an EF1α promoter followed by an inverted H2B-EGFP coding sequence that is flanked by partial AAVS1 DNA sequence, an internal ribosome entry site (IRES), and a puromycin resistance gene. Successful installation of opposite-facing attB (left) and attP (right) sequences at the AAVS1 target sequences and subsequent inversion by Bxb1 corrects the orientation of GFP for functional expression. (b) The fluorescent reporter construct was stably integrated into HEK293T cells via lentiviral transduction and puromycin selection. The polyclonal GFP reporter cell line was then transfected with twinPE plasmid components (PE2 and four pegRNAs) and varying amounts of Bxb1 plasmid for single-transfection inversion. Cells were analyzed by flow cytometry and gated for live single cells. Quantification of GFP positive cells by flow cytometry. Values and error bars reflect the mean of two independent biological replicates.
Extended Data Fig. 9 TwinPE and Bxb1 recombinase-mediated inversion between IDS and IDS2.
(a) Assessment of the inverted IDS junction purity by high-throughput sequencing in HEK293T cells. Frequency of expected junction sequences containing attR and attL recombination products after twinPE and BxB1-mediated single-step inversion. The product purities range from 81-89%. Values and error bars reflect the mean and s.d. of three independent biological replicates. (b) Schematic diagram of the designed PCR strategies for quantifying IDS inversion efficiency. Primer pair 1 (green forward and blue reverse primer) can amplify the unedited alleles (403 bp), twinPE-edited alleles (337 bp), and the inverted alleles (326 bp) at junction 1 in a single PCR reaction. Due to the size difference, a UMI protocol was applied to eliminate PCR bias during quantification of inversion efficiency. Similarly, using primer pair 2 (red forward and blue reverse primer), the unedited alleles (346 bp), twinPE-edited alleles (326 bp), and inverted alleles (320 bp) at junction 2 can be amplified in a single PCR reaction. Amplicons can then be sequenced by standard high-throughput sequencing protocols for amplicon sequencing. (c) Screening of pegRNA pairs for the insertion of Bxb1 attB and attP sequences at IDS and IDS2. TwinPE editing was tested with standard pegRNAs and epegRNAs containing a 3’ evoPreQ1 motif. Values and error bars reflect the mean and s.d. of three independent biological replicates.
Extended Data Fig. 10 Twin prime editing mediated insertion in CCR5 region 2 in HEK293T cells, twin prime editing in multiple human cell lines, and editing activity of Cas9 nickase and PE2-dead RT variants.
(a) TwinPE-mediated endogenous sequence replacement with Bxb1 attB attachment site in CCR5 region 2 in HEK293T cells. (b) TwinPE-mediated endogenous sequence replacement with attP, attB, or 22-nt DNA sequences in multiple human cell lines. Six different pegRNA pairs targeting five loci were tested in HEK293T, HeLa, U2OS and K562 cells. HEK293T and HeLa cell were transfected with PE2 and pegRNA plasmids via Lipofectamine 2000 (Thermo Fisher) and TransIT-HeLaMonster (Mirus), respectively. U2OS and K562 cells were nucleofected using Lonza 4D-Nucleofector and SE kit. DNA loci and the specified insertion edits are shown in the x-axis. (c) HEK293T cells were transfected with twinPE pegRNA pairs and either Cas9–H840A nickase (nCas9), PE2-dRT (a PE2 variant that contains K103L and R110S inactivating mutations to the RT domain), or PE2. Treatment with either nCas9 or PE2-dRT did not result in desired edits, while PE2 installed the specified edits as indicated. Values and error bars in (a) and (c) reflect the mean and s.d. of three independent biological replicates. Values and error bars in (b) reflect the mean and s.d. of at least two independent biological replicates except editing in IDS2 in HeLa cells, editing in U2OS cells, and editing in MYC in K562 cells, which represent two independent biological replicates.
Supplementary information
Supplementary Information
Supplementary Tables 1–5, Notes 1–4 and Sequences 1–3.
Source data
Source Data Extended Data Fig. 6
Unprocessed agarose gel.
Source Data Extended Data Fig. 7
Unprocessed agarose gel.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Anzalone, A.V., Gao, X.D., Podracky, C.J. et al. Programmable deletion, replacement, integration and inversion of large DNA sequences with twin prime editing. Nat Biotechnol 40, 731–740 (2022). https://doi.org/10.1038/s41587-021-01133-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41587-021-01133-w
This article is cited by
-
Gene editing technology to improve antitumor T-cell functions in adoptive immunotherapy
Inflammation and Regeneration (2024)
-
Improving CRISPR–Cas9 directed faithful transgene integration outcomes by reducing unwanted random DNA integration
Journal of Biomedical Science (2024)
-
Precise integration of large DNA sequences in plant genomes using PrimeRoot editors
Nature Biotechnology (2024)
-
CRISPR technologies for genome, epigenome and transcriptome editing
Nature Reviews Molecular Cell Biology (2024)
-
Exonuclease-enhanced prime editors
Nature Methods (2024)