Abstract
Tumor-specific elucidation of physical and functional oncoprotein interactions could improve tumorigenic mechanism characterization and therapeutic response prediction. Current interaction models and pathways, however, lack context specificity and are not oncoprotein specific. We introduce SigMaps as context-specific networks, comprising modulators, effectors and cognate binding-partners of a specific oncoprotein. SigMaps are reconstructed de novo by integrating diverse evidence sources—including protein structure, gene expression and mutational profiles—via the OncoSig machine learning framework. We first generated a KRAS-specific SigMap for lung adenocarcinoma, which recapitulated published KRAS biology, identified novel synthetic lethal proteins that were experimentally validated in three-dimensional spheroid models and established uncharacterized crosstalk with RAB/RHO. To show that OncoSig is generalizable, we first inferred SigMaps for the ten most mutated human oncoproteins and then for the full repertoire of 715 proteins in the COSMIC Cancer Gene Census. Taken together, these SigMaps show that the cell’s regulatory and signaling architecture is highly tissue specific.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All data are provided in Supplementary Files. Source data are provided with this paper.
Code availability
R scripts and code for Naive Bayes and Random Forest classifiers and input data files to reproduce the results described are freely available at https://github.com/califano-lab/OncoSig.
References
Prahallad, A. et al. Unresponsiveness of colon cancer to BRAF(V600E) inhibition through feedback activation of EGFR. Nature 483, 100–103 (2012).
Bild, A. H. et al. Oncogenic pathway signatures in human cancers as a guide to targeted therapies. Nature 439, 353–357 (2006).
Krogan, N. J., Lippman, S., Agard, D. A., Ashworth, A. & Ideker, T. The Cancer Cell Map Initiative: defining the hallmark networks of cancer. Mol. Cell 58, 690–698 (2015).
Greene, C. S. et al. Understanding multicellular function and disease with human tissue-specific networks. Nat. Genet. 47, 569–576 (2015).
Cancer Genome Atlas Research Networket al. The Cancer Genome Atlas Pan-Cancer analysis project. Nat. Genet. 45, 1113–1120 (2013).
Zhang, Q. C. et al. Structure-based prediction of protein–protein interactions on a genome-wide scale. Nature 490, 556–560 (2012).
Garzon, J. I. et al. A computational interactome and functional annotation for the human proteome. eLife 5, e18715 (2016).
Ashburner, M. et al. Gene Ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat. Genet. 25, 25–29 (2000).
Margolin, A. A. et al. ARACNE: an algorithm for the reconstruction of gene regulatory networks in a mammalian cellular context. BMC Bioinformatics 7 Suppl 1, S7 (2006).
Alvarez, M. J. et al. Functional characterization of somatic mutations in cancer using network-based inference of protein activity. Nat. Genet. 48, 838–847 (2016).
Wang, K. et al. Genome-wide identification of post-translational modulators of transcription factor activity in human B cells. Nat. Biotechnol. 27, 829–839 (2009).
Giorgi, F. M. et al. Inferring protein modulation from gene expression data using conditional mutual information. PLoS ONE 9, e109569 (2014).
Network, C. G. A. R. Comprehensive molecular profiling of lung adenocarcinoma. Nature 511, 543–550 (2014).
Jansen, R. et al. A Bayesian networks approach for predicting protein–protein interactions from genomic data. Science 302, 449–453 (2003).
Breiman, L. Random forests. Machine Learning 45, 5–32 (2001).
Liberzon, A. et al. The molecular signatures database (MSigDB) hallmark gene set collection. Cell Syst. 1, 417–425 (2015).
Kanehisa, M., Sato, Y., Kawashima, M., Furumichi, M. & Tanabe, M. KEGG as a reference resource for gene and protein annotation. Nucleic Acids Res. 44, D457–D462 (2016).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Huttlin, E. L. et al. The BioPlex network: a systematic exploration of the human interactome. Cell 162, 425–440 (2015).
Rolland, T. et al. A proteome-scale map of the human interactome network. Cell 159, 1212–1226 (2014).
Chatr-Aryamontri, A. et al. The BioGRID interaction database: 2017 update. Nucleic Acids Res. 45, D369–D379 (2017).
Corsello, S. M. et al. The Drug Repurposing Hub: a next-generation drug library and information resource. Nat. Med. 23, 405–408 (2017).
Franceschini, A. et al. STRING v9.1: protein–protein interaction networks, with increased coverage and integration. Nucleic Acids Res. 41, D808–D815 (2013).
Lee, I., Blom, U. M., Wang, P. I., Shim, J. E. & Marcotte, E. M. Prioritizing candidate disease genes by network-based boosting of genome-wide association data. Genome Res. 21, 1109–1121 (2011).
Barbie, D. A. et al. Systematic RNA interference reveals that oncogenic KRAS-driven cancers require TBK1. Nature 462, 108–112 (2009).
Kim, J. et al. XPO1-dependent nuclear export is a druggable vulnerability in KRAS-mutant lung cancer. Nature 538, 114–117 (2016).
Corcoran, R. B. et al. Synthetic lethal interaction of combined BCL-XL and MEK inhibition promotes tumor regressions in KRAS mutant cancer models. Cancer Cell 23, 121–128 (2013).
Hayes, T. K. et al. Long-term ERK inhibition in KRAS-mutant pancreatic cancer is associated with MYC degradation and senescence-like growth suppression. Cancer Cell 29, 75–89 (2016).
Shaw, A. T. et al. Selective killing of K-ras mutant cancer cells by small molecule inducers of oxidative stress. Proc. Natl Acad. Sci. USA 108, 8773–8778 (2011).
Liu, Z., Xiao, T., Peng, X., Li, G. & Hu, F. APPLs: more than just adiponectin receptor binding proteins. Cell. Signal. 32, 76–84 (2017).
Tzeng, H. T. & Wang, Y. C. Rab-mediated vesicle trafficking in cancer. J. Biomed. Sci. 23, 70 (2016).
Thomas, J. D. et al. Rab1A is an mTORC1 activator and a colorectal oncogene. Cancer Cell 26, 754–769 (2014).
Gabernet-Castello, C., O’Reilly, A. J., Dacks, J. B. & Field, M. C. Evolution of Tre-2/Bub2/Cdc16 (TBC) Rab GTPase-activating proteins. Mol. Biol. Cell 24, 1574–1583 (2013).
Lu, W. et al. Downregulation of ARHGDIA contributes to human glioma progression through activation of Rho GTPase signaling pathway. Tumour Biol. 37, 15783–15793 (2016).
Hornstein, I., Alcover, A. & Katzav, S. Vav proteins, masters of the world of cytoskeleton organization. Cell. Signal. 16, 1–11 (2004).
Oliver, A. W. et al. The HPV16 E6 binding protein Tip-1 interacts with ARHGEF16, which activates Cdc42. Br. J. Cancer 104, 324–331 (2011).
Boulter, E., Estrach, S., Garcia-Mata, R. & Feral, C. C. Off the beaten paths: alternative and crosstalk regulation of Rho GTPases. FASEB J. 26, 469–479 (2012).
Cox, A. D. & Der, C. J. Ras history: the saga continues. Small GTPases 1, 2–27 (2010).
Prior, I. A. & Hancock, J. F. Ras trafficking, localization and compartmentalized signalling. Semin. Cell Dev. Biol. 23, 145–153 (2012).
Bhuin, T. & Roy, J. K. Rab proteins: the key regulators of intracellular vesicle transport. Exp. Cell Res. 328, 1–19 (2014).
Fukuda, M. TBC proteins: GAPs for mammalian small GTPase Rab? Biosci. Rep. 31, 159–168 (2011).
Hwang, J. & Pallas, D. C. STRIPAK complexes: structure, biological function, and involvement in human diseases. Int. J. Biochem. Cell Biol. 47, 118–148 (2014).
Skrzypski, M. et al. Three-gene expression signature predicts survival in early-stage squamous cell carcinoma of the lung. Clin. Cancer Res. 14, 4794–4799 (2008).
Li, N. & Li, S. RASAL2 promotes lung cancer metastasis through epithelial–mesenchymal transition. Biochem. Biophys. Res. Commun. 455, 358–362 (2014).
Yu, F. et al. IFITM1 promotes the metastasis of human colorectal cancer via CAV-1. Cancer Lett. 368, 135–143 (2015).
Weinberg, F. D. & Ramnath, N. Targeting IL22: a potential therapeutic approach for Kras mutant lung cancer? Transl. Lung Cancer Res. 7, S243–S247 (2018).
Guillon, A. et al. Interleukin-22 receptor is overexpressed in nonsmall cell lung cancer and portends a poor prognosis. Eur. Respir. J. 47, 1277–1280 (2016).
Janne, P. A. et al. Selumetinib plus docetaxel for KRAS-mutant advanced non-small-cell lung cancer: a randomised, multicentre, placebo-controlled, phase 2 study. Lancet Oncol. 14, 38–47 (2013).
Migliardi, G. et al. Inhibition of MEK and PI3K/mTOR suppresses tumor growth but does not cause tumor regression in patient-derived xenografts of RAS-mutant colorectal carcinomas. Clin. Cancer Res. 18, 2515–2525 (2012).
Adjei, A. A. et al. Phase I pharmacokinetic and pharmacodynamic study of the oral, small-molecule mitogen-activated protein kinase kinase 1/2 inhibitor AZD6244 (ARRY-142886) in patients with advanced cancers. J. Clin. Oncol. 26, 2139–2146 (2008).
Sustic, T., Bosdriesz, E., van Wageningen, S., Wessels, L. F. A. & Bernards, R. RUNX2/CBFB modulates the response to MEK inhibitors through activation of receptor tyrosine kinases in KRAS-mutant colorectal cancer. Transl. Oncol. 13, 201–211 (2019).
Astsaturov, I. et al. Synthetic lethal screen of an EGFR-centered network to improve targeted therapies. Sci. Signal. 3, ra67 (2010).
Cancer Genome Atlas Research Network. Comprehensive molecular profiling of lung adenocarcinoma. Nature 511, 543–550 (2014).
Tate, J. G. et al. COSMIC: the catalogue of somatic mutations in cancer. Nucleic Acids Res. 47, D941–D947 (2019).
Narlikar, G. J., Sundaramoorthy, R. & Owen-Hughes, T. Mechanisms and functions of ATP-dependent chromatin-remodeling enzymes. Cell 154, 490–503 (2013).
Morozumi, Y. et al. Atad2 is a generalist facilitator of chromatin dynamics in embryonic stem cells. J. Mol. Cell Biol. 8, 349–362 (2016).
Sharma, V. M., Li, B. & Reese, J. C. SWI/SNF-dependent chromatin remodeling of RNR3 requires TAF(II)s and the general transcription machinery. Genes Dev. 17, 502–515 (2003).
Mi, H., Muruganujan, A., Ebert, D., Huang, X. & Thomas, P. D. PANTHER version 14: more genomes, a new PANTHER GO-slim and improvements in enrichment analysis tools. Nucleic Acids Res. 47, D419–D426 (2019).
Gong, F. & Miller, K. M. Double duty: ZMYND8 in the DNA damage response and cancer. Cell Cycle 17, 414–420 (2018).
Sridhara, S. C. et al. Transcription dynamics prevent RNA-mediated genomic instability through SRPK2-dependent DDX23 phosphorylation. Cell Rep. 18, 334–343 (2017).
Allemand, E. et al. A broad set of chromatin factors influences splicing. PLoS Genet. 12, e1006318 (2016).
Blume-Jensen, P. & Hunter, T. Oncogenic kinase signalling. Nature 411, 355–365 (2001).
Lemmon, M. A. & Schlessinger, J. Cell signaling by receptor tyrosine kinases. Cell 141, 1117–1134 (2010).
Organ, S. L. & Tsao, M. S. An overview of the c-MET signaling pathway. Ther. Adv. Med. Oncol. 3, S7–S19 (2011).
Meissl, K., Macho-Maschler, S., Muller, M. & Strobl, B. The good and the bad faces of STAT1 in solid tumours. Cytokine 89, 12–20 (2017).
Zhang, Y. & Liu, Z. STAT1 in cancer: friend or foe? Discov. Med. 24, 19–29 (2017).
Balbin, O. A. et al. Reconstructing targetable pathways in lung cancer by integrating diverse omics data. Nat. Commun. 4, 2617 (2013).
Downward, J. RAS synthetic lethal screens revisited: still seeking the elusive prize? Clin. Cancer Res. 21, 1802–1809 (2015).
Luo, J. et al. A genome-wide RNAi screen identifies multiple synthetic lethal interactions with the Ras oncogene. Cell 137, 835–848 (2009).
Wang, T. et al. Gene essentiality profiling reveals gene networks and synthetic lethal interactions with oncogenic Ras. Cell 168, 890–903 (2017).
McDonald, E. R. III et al. Project DRIVE: a compendium of cancer dependencies and synthetic lethal relationships uncovered by large-scale, deep RNAi screening. Cell 170, 577–592 (2017).
Aguirre, A. J. & Hahn, W. C. Synthetic lethal vulnerabilities in KRAS-mutant cancers. Cold Spring Harb. Perspect. Med. 8, a031518 (2018).
Woo, J. H. et al. Elucidating compound mechanism of action by network perturbation analysis. Cell 162, 441–451 (2015).
Duan, Q. et al. LINCS Canvas Browser: interactive web app to query, browse and interrogate LINCS L1000 gene expression signatures. Nucleic Acids Res. 42, W449–W460 (2014).
Kramer, A., Green, J., Pollard, J. Jr. & Tugendreich, S. Causal analysis approaches in Ingenuity Pathway Analysis. Bioinformatics 30, 523–530 (2014).
Li, W. & Godzik, A. Cd-hit: a fast program for clustering and comparing large sets of protein or nucleotide sequences. Bioinformatics 22, 1658–1659 (2006).
Arlot, S. & Celisse, A. A survey of cross-validation procedures for model selection. Statist. Surv. 4, 40–79 (2010).
Torres, J. Z., Miller, J. J. & Jackson, P. K. High-throughput generation of tagged stable cell lines for proteomic analysis. Proteomics 9, 2888–2891 (2009).
Shevchenko, A., Tomas, H., Havlis, J., Olsen, J. V. & Mann, M. In-gel digestion for mass spectrometric characterization of proteins and proteomes. Nat. Protoc. 1, 2856–2860 (2006).
Zheng, Y. et al. A rare population of CD24(+)ITGB4(+)Notch(hi) cells drives tumor propagation in NSCLC and requires Notch3 for self-renewal. Cancer Cell 24, 59–74 (2013).
Dai, Z. et al. edgeR: a versatile tool for the analysis of shRNA-seq and CRISPR–Cas9 genetic screens. F1000Res 3, 95 (2014).
Acknowledgements
This work was supported by the National Cancer Institute (NCI) Outstanding Investigator Award R35CA197745 to A.C.; the NCI Cancer Target Discovery and Development Program U01CA168426 to A.C.; the NCI Research Centers for Cancer Systems Biology Consortium U54CA209997 to A.C. and B.H.; National Institute of General Medical Sciences grant R01GM30518 to B.H.; NCI grant R01CA129562 to E.A.S.C.; Innovative Research Grant from Stand Up to Cancer to E.A.S.C.; National Institutes of Health High-End Instrumentation Program grant S10OD012351 to A.C.; and NIH Shared Instrumentation Program grant S10OD021764 to A.C. J.B. was supported, in part, by the Ruth L. Kirschstein National Research Service Award Institutional Research Training Grant T32GM082797. D.R.S. was supported by the Ruth L. Kirschstein National Research Service Award Institutional Research Training Grant T32CA09302. Relevant ethical compliance was ensured by the Institutional Review Board of the Human Research Protection Program at the University of California, San Francisco.
Author information
Authors and Affiliations
Contributions
J.B., D.M., A.T.G., A.L. and F.M.G. performed computational analysis. J.B. performed machine learning. D.S. and P.K.J. performed, respectively, the knockdown and AP–MS experiments. E.O.P, B.W.C. and S.T. created oncosig.org. J.B., D.M. and S.J.J. compiled the codebook. J.B., D.S., E.A.S.C., A.C. and B.H. analyzed knockdown experiments. S.D.M., B.H., A.C. and E.A.S.C. designed research. D.M., B.H. and A.C. designed computational and experimental work and analyzed data. D.M., D.S., J.B., A.C. and B.H. assembled the data and wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
A.C. is the founder and an equity holder in DarwinHealth, a company that has licensed some of the algorithms used in this study from Columbia University. Columbia University is also an equity holder in DarwinHealth.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 The top 40 predictions from the OncoSigNB algorithm for the KRAS LUAD SigMap were chosen for validation.
a, Performance of OncoSigNB at recovering the Ingenuity-derived positive gold standard set (PGSS) as a function of LRPost. At an LRPost = 53 (probability = 0.50) (vertical gray line), the OncoSigNB LUAD-specific KRAS SigMap contains 10% of the PGSS (horizontal gray line). The vertical red line corresponds to LRPost = 240, the cutoff used to obtain candidates for experimental validation. b, ROC curve analysis, evaluated as the recovery of the Ingenuity-derived PGSS (FPR ≤ 0.05), for 1) OncoSigNB (green curve, N = 1028)), 2) Pearson’s correlation between mRNA expression of KRAS and mRNA expression of other proteins in LUAD (blue curve), and 3) random performance (black curve). Recovery using 2-fold cross-validation (green) is essentially indistinguishable from recovery using 100-fold Monte-Carlo Cross-validation (not shown). 393 OncoSigNB LUAD-specific KRAS SigMap predictions are made for LRPost ≥ 53, which corresponds to probability ≥ 0.50 and FPR ≤ 0.019 (purple dot). 40 OncoSigNB LUAD-specific KRAS SigMap predictions are made for LRPost ≥ 240, which corresponds to probability ≥ 0.82 and FPR ≤ 0.0018 (yellow dot). The top 40 predictions are listed by gene name in (c). c, Orange and blue boxes contain, respectively, known upstream regulators and downstream effectors that are successfully recovered by OncoSigNB. Italicized text indicates proteins known to interact with KRAS via a physical protein-protein interaction. The box titled “validated predictions” shows the novel OncoSigNB predictions tested with the RNAi negative screen; those that were experimentally found to affect cell growth in a KRAS-dependent context are highlighted in bold text.
Extended Data Fig. 2 The OncoSigRF and OncoSigNB algorithms produce highly similar KRAS LUAD SigMaps.
a, Comparison of ROC curves (FPR ≤ 0.05) for LUAD-specific KRAS SigMaps predicted by OncoSigNB (green and blue curves) and OncoSigRF (orange and red curve) trained on the Ingenuity PGSS and the MSigDB PGSS, respectively. b, Gene Set Enrichment Analysis (GSEA) of the top 100 OncoSigNB LUAD-specific KRAS SigMap predictions at the top of the OncoSigRF LUAD-specific KRAS SigMap predictions. Ranking is based on OncoSigRF score (SRF). Both the OncoSigNB predictions tested in the knockdown experiments (red lines) and the remaining top 100 OncoSigNB predictions (blue lines) are highly enriched at the top of the OncoSigRF predictions (p = 5.6 ×10−8 and p = 1.7 ×10-19, respectively).
Extended Data Fig. 3 The log2FC of shRNA abundance is plotted against the novel proteins tested in the KRAS negative selection screen.
The 3–5 points plotted for a given protein are shRNAs that target the mRNA for that protein (N = 100). The X-axis is sorted by mean log2FC for all shRNAs targeting each gene. Colors change from red to green with mean log2FC.
Extended Data Fig. 4 OncoSigRF predictions are highly enriched in oncogenic KRASMut dependencies.
a, GSEA of KRASMut synthetic lethal partners25 (blue lines, N = 216) and the top 500 OncoSigRF LUAD-specific KRAS SigMap predictions obtained by training on a modified PGSS for which the intersection with the synthetic lethal set was removed. Inset is the GSEA using all OncoSigRF predictions obtained in this way, where the ranking is OncoSigRF score. Enrichment analysis was performed with the aREA (analytic Rank-based Enrichment Analysis) algorithm10. b, Enrichment of the protein resistance-signature to ERK inhibitor SCH77298428 (blue lines, N = 24) within OncoSigRF LUAD-specific KRAS SigMap predictions10. c, Enrichment of proteins involved in response to Reactive Oxygen Species (GO:0000302)8,29 (blue lines, N = 276) within OncoSigRF LUAD-specific KRAS SigMap predictions10.
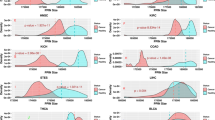
Extended Data Fig. 5 OncoSigRF KRAS SigMaps exhibit tissue context specificity.
a, ROC curves for the OncoSigRF KRAS SigMaps in LUAD (red), LUSC (gray) COAD (brown), and PAAD (orange) for FPR ≤ 0.05. Performance is evaluated as the recovery of established KRAS pathway proteins. b, Gene set enrichment analysis (GSEA) of KRASMut synthetic lethal partners, as determined by Corcoran et al27 (N = 48, blue lines). To avoid training and testing on the same proteins, OncoSigRF predictions for COAD-specific KRAS SigMap proteins were obtained by training on a modified PGSS from which any established KRASMut synthetic lethal protein had been previously removed. Enrichment analysis was performed with the aREA (analytic Rank-based Enrichment Analysis) algorithm10. c, Scatterplot of OncoSigRF scores for KRAS SigMap proteins in PAAD-vs-LUAD (N = 19,789). Each dot represents the scores for one protein. Darker colored points have high scores (SRF ≥ 0.5) in at least one context, and lighter colored points score poorly in both contexts (SRF ≤ 0.5). R2PAAD/LUAD = 0.037. d, OncoSigRF COAD-specific KRAS SigMap in the form depicted conceptually in Fig. 1a. To prevent visual cluttering, only the top 33 OncoSigRF predictions (FPR ≤ 0.01) that are also VIPER-inferred KRAS interactors (p ≤ 0.01), PrePPI-predicted KRAS physical interactors, or both, are depicted. Bold and regular text node labels represent established and novel predictions, respectively; orange and blue node colors represent upstream regulators and downstream effectors, respectively; red, blue, and black node borders represent predictions that are druggable (Drug Repurposing Hub22), KRASMut synthetic lethal from the literature and validated here (see text), and both, respectively; orange and blue solid lines and gray nodes represent PrePPI-predicted physical interactors of KRAS.
Extended Data Fig. 6 OncoSigRF SigMaps for hypermutated oncoproteins are retrospectively validated.
a, Pairwise overlap of established pathway proteins (left) and the OncoSigRF LUAD-specific SigMaps (FPR ≤ 0.01, right) for the ten hyper-mutated oncoproteins (names of columns and rows). Percent overlap is color-coded according to the scale at top. b, SigMap predictions are highly enriched in 600 EGFR-centric network proteins52 (p = 2.3 ×10−43). Enrichment analysis was performed with the aREA (analytic Rank-based Enrichment Analysis) algorithm10. c, Box plots of the OncoSigRF LUAD-specific EGFR SigMap scores for two subsets of the curated EGFR pathway proteins from Astsaturov et al:52 those identified as EGFR synthetic lethal partners (red, N = 58) and those not identified as synthetic lethal (grey, N = 542). The p-value (2 ×10−4) was calculated using Welch’s two sample t-test.
Source data
Source Data Fig. 1c
Statistical Source Data
Source Data Fig. 2a
Statistical Source Data
Source Data Fig. 2b
Statistical Source Data
Source Data Fig. 3d
Statistical Source Data
Source Data Fig. 4a
Statistical Source Data
Source Data Fig. 4b
Statistical Source Data
Source Data Fig. 5a
Statistical Source Data
Source Data Fig. 5b
Statistical Source Data
Source Data Fig. 6a
Statistical Source Data
Source Data Fig. 3d
Layered vector image Powerpoint
Source Data Fig. 6bcd
Layered vector image Powerpoint
Source Data Extended Data Fig. 1a
Statistical Source Data
Source Data Extended Data Fig. 1b
Statistical Source Data
Source Data Extended Data Fig. 2a
Statistical Source Data
Source Data Extended Data Fig. 5a
Statistical Source Data
Source Data Extended Data Fig. 5c
Statistical Source Data
Source Data Extended Data Fig. 6a
Statistical Source Data
Rights and permissions
About this article
Cite this article
Broyde, J., Simpson, D.R., Murray, D. et al. Oncoprotein-specific molecular interaction maps (SigMaps) for cancer network analyses. Nat Biotechnol 39, 215–224 (2021). https://doi.org/10.1038/s41587-020-0652-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41587-020-0652-7
This article is cited by
-
Pancancer network analysis reveals key master regulators for cancer invasiveness
Journal of Translational Medicine (2023)
-
UHRF1 is a mediator of KRAS driven oncogenesis in lung adenocarcinoma
Nature Communications (2023)
-
The application of patient-derived organoid in the research of lung cancer
Cellular Oncology (2023)
-
Multiplexed screens identify RAS paralogues HRAS and NRAS as suppressors of KRAS-driven lung cancer growth
Nature Cell Biology (2023)