Abstract
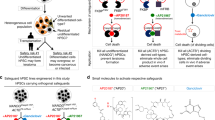
Safeguard mechanisms can ameliorate the potential risks associated with cell therapies but currently rely on the introduction of transgenes. This limits their application owing to immunogenicity or transgene silencing. We aimed to create a control mechanism for human cells that is not mediated by a transgene. Using genome editing methods, we disrupt uridine monophosphate synthetase (UMPS) in the pyrimidine de novo synthesis pathway in cell lines, pluripotent cells and primary human T cells. We show that this makes proliferation dependent on external uridine and enables us to control cell growth by modulating the uridine supply, both in vitro and in vivo after transplantation in xenograft models. Additionally, disrupting this pathway creates resistance to 5-fluoroorotic acid, which enables positive selection of UMPS-knockout cells. We envision that this approach will add an additional level of safety to cell therapies and therefore enable the development of approaches with higher risks, especially those that are intended for limited treatment durations.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All data generated or analyzed during this study are included in this published article and its extended data. Source data are provided with this paper.
Code availability
The custom script used to analyze next-generation sequencing data indelQuantificationFromFastqPaired-1.0.1.pl can be found at https://github.com/piyuranjan/NucleaseIndelActivityScript/blob/master/indelQuantificationFromFastqPaired-1.0.1.pl.
References
Hoggatt, J. Gene therapy for ‘bubble boy’ disease. Cell 166, 263 (2016).
Majzner, R. G., Heitzeneder, S. & Mackall, C. L. Harnessing the immunotherapy revolution for the treatment of childhood cancers. Cancer Cell 31, 476–485 (2017).
Teixeira, A. P. & Fussenegger, M. Engineering mammalian cells for disease diagnosis and treatment. Curr. Opin. Biotechnol. 55, 87–94 (2019).
Chen, Y. Y. & Smolke, C. D. From DNA to targeted therapeutics: bringing synthetic biology to the clinic. Sci. Transl. Med. 3, 106ps42 (2011).
Tang, J., Hubbard-Lucey, V. M., Pearce, L., O’Donnell-Tormey, J. & Shalabi, A. The global landscape of cancer cell therapy. Nat. Rev. Drug Discov. 17, 465–466 (2018).
Bonifant, C. L., Jackson, H. J., Brentjens, R. J. & Curran, K. J. Toxicity and management in CAR T-cell therapy. Mol. Ther. Oncolytics 3, 16011 (2016).
Sadelain, M. Eliminating cells gone astray. N. Engl. J. Med. 365, 1735–1737 (2011).
Porteus, M. Translating the lessons from gene therapy to the development of regenerative medicine. Mol. Ther. 19, 439–441 (2011).
Narayanan, P. et al. A composite MyD88/CD40 switch synergistically activates mouse and human dendritic cells for enhanced antitumor efficacy. J. Clin. Invest. 121, 1524–1534 (2011).
Sockolosky, J. T. et al. Selective targeting of engineered T cells using orthogonal IL-2 cytokine–receptor complexes. Science 359, 1037–1042 (2018).
Tey, S.-K. Adoptive T-cell therapy: adverse events and safety switches. Clin. Transl. Immunol. 3, e17 (2014).
Ciceri, F. et al. Infusion of suicide-gene-engineered donor lymphocytes after family haploidentical haemopoietic stem-cell transplantation for leukaemia (The TK007 Trial): a non-randomised phase I–II study. Lancet Oncol. 10, 489–500 (2009).
Di Stasi, A. et al. Inducible apoptosis as a safety switch for adoptive cell therapy. N. Engl. J. Med. 365, 1673–1683 (2011).
Ben-David, U. & Benvenisty, N. The tumorigenicity of human embryonic and induced pluripotent stem cells. Nat. Rev. Cancer 11, 268–277 (2011).
Lee, A. S., Tang, C., Rao, M. S., Weissman, I. L. & Wu, J. C. Tumorigenicity as a clinical hurdle for pluripotent stem cell therapies. Nat. Med. 19, 998–1004 (2013).
Traversari, C. et al. The potential immunogenicity of the TK suicide gene does not prevent full clinical benefit associated with the use of TK-transduced donor lymphocytes in HSCT for hematologic malignancies. Blood 109, 4708–4715 (2007).
Yagyu, S., Hoyos, V., Del Bufalo, F. & Brenner, M. K. An inducible caspase-9 suicide gene to improve the safety of therapy using human induced pluripotent stem cells. Mol. Ther. 23, 1475–1485 (2015).
Garin, M. I. et al. Molecular mechanism for ganciclovir resistance in human T lymphocytes transduced with retroviral vectors carrying the herpes simplex virus thymidine kinase gene. Blood 97, 122–129 (2001).
Wu, C. et al. Development of an inducible caspase-9 safety switch for pluripotent stem cell-based therapies. Mol. Ther. Methods Clin. Dev. 1, 14053 (2014).
Sułkowski, M., Konieczny, P., Chlebanowska, P. & Majka, M. Introduction of exogenous HSV-TK suicide gene increases safety of keratinocyte-derived induced pluripotent stem cells by providing genetic ‘emergency exit’ switch. Int. J. Mol. Sci. 19, 197 (2018).
Ando, M. et al. A safeguard system for induced pluripotent stem cell-derived rejuvenated T cell therapy. Stem Cell Rep. 5, 597–608 (2015).
Merkle, F. T. et al. Human pluripotent stem cells recurrently acquire and expand dominant negative p53 mutations. Nature 545, 229–233 (2017).
Frank, O. et al. Tumor cells escape suicide gene therapy by genetic and epigenetic instability. Blood 104, 3543–3549 (2004).
Li, H. & Zhao, Y. Increasing the safety and efficacy of chimeric antigen receptor T cell therapy. Protein Cell 8, 573–589 (2017).
Terazaki, Y. et al. An optimal therapeutic expression level is crucial for suicide gene therapy for hepatic metastatic cancer in mice. Hepatology 37, 155–163 (2003).
van Galen, P. et al. The unfolded protein response governs integrity of the haematopoietic stem-cell pool during stress. Nature 510, 268–272 (2014).
Murray, P. J. Amino acid auxotrophy as a system of immunological control nodes. Nat. Immunol. 17, 132–139 (2016).
Grohmann, U. et al. Amino-acid sensing and degrading pathways in immune regulation. Cytokine Growth Factor Rev. 35, 37–45 (2017).
Fung, M. K. L. & Chan, G. C.-F. Drug-induced amino acid deprivation as strategy for cancer therapy. J. Hematol. Oncol. 10, 144 (2017).
Hill, J. M. et al. l-Asparaginase therapy for leukemia and other malignant neoplasms. JAMA 202, 882–888 (1967).
Kato, Y. An engineered bacterium auxotrophic for an unnatural amino acid: a novel biological containment system. PeerJ 3, e1247 (2015).
Steidler, L. et al. Biological containment of genetically modified Lactococcus lactis for intestinal delivery of human interleukin 10. Nat. Biotechnol. 21, 785–789 (2003).
Hendel, A. et al. Chemically modified guide RNAs enhance CRISPR–Cas genome editing in human primary cells. Nat. Biotechnol. 33, 985–989 (2015).
Porteus, M. H. & Baltimore, D. Chimeric nucleases stimulate gene targeting in human cells. Science 300, 763 (2003).
Bak, R. O., Dever, D. P. & Porteus, M. H. CRISPR/Cas9 genome editing in human hematopoietic stem cells. Nat. Protoc. 13, 358–376 (2018).
Rose, M. & Winston, F. Identification of a Ty insertion within the coding sequence of the S. cerevisiae URA3 gene. Mol. Gen. Genet. 193, 557–560 (1984).
Fallon, H. J., Smith, L. H., Graham, J. B. & Burnett, C. H. A genetic study of hereditary orotic aciduria. N. Engl. J. Med. 270, 878–881 (1964).
Okesli, A., Khosla, C. & Bassik, M. C. Human pyrimidine nucleotide biosynthesis as a target for antiviral chemotherapy. Curr. Opin. Biotechnol. 48, 127–134 (2017).
Cradick, T. J., Qiu, P., Lee, C. M., Fine, E. J. & Bao, G. COSMID: a web-based tool for identifying and validating CRISPR/Cas off-target sites. Mol. Ther. Nucleic Acids 3, e214 (2014).
Vakulskas, C. A. et al. A high-fidelity Cas9 mutant delivered as a ribonucleoprotein complex enables efficient gene editing in human hematopoietic stem and progenitor cells. Nat. Med. 24, 1216–1224 (2018).
Brinkman, E. K., Chen, T., Amendola, M. & van Steensel, B. Easy quantitative assessment of genome editing by sequence trace decomposition. Nucleic Acids Res. 42, e168 (2014).
Bak, R. O. et al. Multiplexed genetic engineering of human hematopoietic stem and progenitor cells using CRISPR/Cas9 and AAV6. eLife 6, e27873 (2017).
Wang, X. et al. A transgene-encoded cell surface polypeptide for selection, in vivo tracking, and ablation of engineered cells. Blood 118, 1255–1263 (2011).
Wiebking, V. et al. Genome editing of donor-derived T-cells to generate allogenic chimeric antigen receptor-modified T cells: optimizing αβ T cell-depleted haploidentical hematopoietic stem cell transplantation. Haematologica https://doi.org/10.3324/haematol.2019.233882 (2020).
van Groeningen, C. J., Peters, G. J. & Pinedo, H. M. Reversal of 5-fluorouracil-induced toxicity by oral administration of uridine. Ann. Oncol. 4, 317–320 (1993).
Becroft, D. M., Phillips, L. I. & Simmonds, A. Hereditary orotic aciduria: long-term therapy with uridine and a trial of uracil. J. Pediatr. 75, 885–891 (1969).
Gasser, T., Moyer, J. D. & Handschumacher, R. E. Novel single-pass exchange of circulating uridine in rat liver. Science 213, 777–778 (1981).
Weinberg, M. E. et al. Enhanced uridine bioavailability following administration of a triacetyluridine-rich nutritional supplement. PLoS ONE 6, e14709 (2011).
Ison, G. et al. FDA approval: uridine triacetate for the treatment of patients following fluorouracil or capecitabine overdose or exhibiting early-onset severe toxicities following administration of these drugs. Clin. Cancer Res. 22, 4545–4549 (2016).
Garcia, R. A. G. et al. Severe cytochrome c oxidase inhibition in vivo does not induce a pyrimidine deficiency; neuroprotective action of oral uridine prodrug PN401 requires supraphysiological levels of uridine. Brain Res. 1066, 164–171 (2005).
Karle, J. M., Anderson, L. W., Dietrick, D. D. & Cysyk, R. L. Determination of serum and plasma uridine levels in mice, rats, and humans by high-pressure liquid chromatography. Anal. Biochem. 109, 41–46 (1980).
Ivics, Z. Self-destruct genetic switch to safeguard iPS cells. Mol. Ther. 23, 1417–1420 (2015).
D’Antonio, M. et al. Insights into the mutational burden of human induced pluripotent stem cells from an integrative multi-omics approach. Cell Rep. 24, 883–894 (2018).
Gornalusse, G. G. et al. HLA-E-expressing pluripotent stem cells escape allogeneic responses and lysis by NK cells. Nat. Biotechnol. 35, 765–772 (2017).
Deuse, T. et al. Hypoimmunogenic derivatives of induced pluripotent stem cells evade immune rejection in fully immunocompetent allogeneic recipients. Nat. Biotechnol. 37, 252–258 (2019).
Watanabe, K., Kuramitsu, S., Posey, A. D. & June, C. H. Expanding the therapeutic window for CAR T cell therapy in solid tumors: the knowns and unknowns of CAR T cell biology. Front. Immunol. 9, 2486 (2018).
Lee, J. W., Chan, C. T. Y., Slomovic, S. & Collins, J. J. Next-generation biocontainment systems for engineered organisms. Nat. Chem. Biol. 14, 530–537 (2018).
Fischer, A., Hacein-Bey Abina, S., Touzot, F. & Cavazzana, M. Gene therapy for primary immunodeficiencies. Clin. Genet. 88, 507–515 (2015).
Gaj, T., Gersbach, C. A. & Barbas, C. F. ZFN, TALEN, and CRISPR/Cas-based methods for genome engineering. Trends Biotechnol. 31, 397–405 (2013).
Duong, M. T. et al. Two-dimensional regulation of CAR-T cell therapy with orthogonal switches. Mol. Ther. Oncolytics 12, 124–137 (2019).
Huang, M. & Graves, L. M. De novo synthesis of pyrimidine nucleotides; emerging interfaces with signal transduction pathways. Cell. Mol. Life Sci. 60, 321–336 (2003).
Haeussler, M. et al. Evaluation of off-target and on-target scoring algorithms and integration into the guide RNA selection tool CRISPOR. Genome Biol. 17, 148 (2016).
Dever, D. P. et al. CRISPR/Cas9 β-globin gene targeting in human haematopoietic stem cells. Nature 539, 384–389 (2016).
Aurnhammer, C. et al. Universal real-time PCR for the detection and quantification of adeno-associated virus serotype 2-derived inverted terminal repeat sequences. Hum. Gene Ther. Methods 23, 18–28 (2012).
Martin, R. M. et al. Highly efficient and marker-free genome editing of human pluripotent stem cells by CRISPR–Cas9 RNP and AAV6 donor-mediated homologous recombination. Cell Stem Cell 24, 821–828 (2019).
Lee, C. M., Cradick, T. J. & Bao, G. The Neisseria meningitidis CRISPR–Cas9 system enables specific genome editing in mammalian cells. Mol. Ther. 24, 645–654 (2016).
Lin, Y. et al. CRISPR/Cas9 systems have off-target activity with insertions or deletions between target DNA and guide RNA sequences. Nucleic Acids Res. 42, 7473–7485 (2014).
Acknowledgements
We thank the Stanford Small Animal Imaging Facility and the FACS Core of the Institute for Stem Cell Biology and Regenerative Medicine at Stanford University for providing access to equipment and training. We thank the Stanford Medicine Veterinary Service Center for excellent animal welfare and husbandry. We thank L. Nguyen (Stanford University) for excellent laboratory management and administration. We thank Synthego for providing modified sgRNAs and IDT for providing early access to high-fidelity Cas9 protein. V.W. gratefully acknowledges receiving research fellowships from the Deutsche Forschungsgemeinschaft (DFG) and the Care-For-Rare Foundation, Germany. We thank the Amon G. Carter Foundation and the Laurie Kraus Lacob Faculty Scholar Award in Pediatric Translational Research for support of this work.
Author information
Authors and Affiliations
Contributions
Conceptualization: V.W., J.O.P. and M.H.P.; methodology: V.W., C.M.L. and M.H.P.; validation: V.W. and C.M.L.; formal analysis: V.W.; investigation: V.W., R.M., M.K.C. and C.M.L.; resources: V.W., W.S., M.K.C., C.M.L., G.B. and M.H.P.; writing–original draft: V.W.; writing–review and editing: V.W. and M.H.P.; visualization: V.W.; supervision: G.B. and M.H.P.; project administration: V.W. and M.H.P.; funding acquisition: V.W. and M.H.P.
Corresponding author
Ethics declarations
Competing interests
The authors declare the following competing interests: V.W., J.O.P. and M.H.P. are inventors on intellectual property related to this work. V.W., M.H.P. and J.O.P. own shares of and J.O.P. is the director of Auxolytic Ltd, a company that owns intellectual property related to this work. J.O.P. declares that he is bound by confidentiality agreements that prevent him from disclosing additional competing interests in this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Predicted specificity for sgRNAs in UMPS exon 1.
a, sgRNAs in the target region of the UMPS gene are shown with their attributed symbol, the genomic target sequence and the specificity score. sgRNAs are ranked by their predicted specificity. b, Numbers of predicted off-target sites as evaluated by COSMID are shown in a stacked bar graph. Grey shades encode the COSMID score as annotated in the legend. Sites with lower scores are predicted to be more relevant (higher chance of endonuclease activity). Y-axis is broken to better illustrate results for sgRNAs with <10 predicted off-target sites.
Extended Data Fig. 2 Proliferation of bulk edited T cell populations and the InDel spectrum after gene editing.
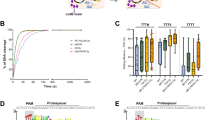
a, Proliferation of bulk T cell populations either mock treated or edited at the CCR5 or UMPS loci and cultured without supplementation of UMP or uridine. Graphs show viable cell counts as measured by trypan blue exclusion. b, Proliferation of a bulk T cell population after electroporation of an RNP targeting UMPS cultured either with high concentrations of uridine or UMP, or without nutrient supplementation. Data points in a, b represent means of 3 biological replicates. Data is from the same experiment shown in Fig. 1f. For statistical analysis on day 5 see Fig. 1f. c, InDel spectrum on day 5 after electroporation of T cells with RNP targeting UMPS, for samples cultured either with or without nutrients as indicated.
Extended Data Fig. 3 Gene targeting approaches to create UMPSKO/KO cells and to knock-in Luciferase-GFP into a safe-harbor locus.
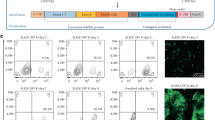
a, Gene targeting approach used to create UMPS-/- Nalm6 cells and UMPS-/- T cells. Dual-allelic gene targeting using constructs carrying tNGFR and tEGFR allow for the purification of the dual-positive population by either fluorescence-activated or magnetic beads-activated cell sorting (FACS or MACS). b, Representative FACS plots of Nalm6 cells that have undergone dual-allelic gene targeting at the UMPS locus and after sorting of the NGFR+/EGFR+ population. Controls are shown that were only mock electroporated or AAV transduced without RNP electroporation. Data from a single experiment. c, Approach used to create cells expressing Luciferase and GFP from a safe harbor. The depicted donor construct was used to target the HBB gene locus considered a safe harbor in non-erythropoietic cells. d, FACS analysis of K562 cells targeted with the approach in c and the relevant controls, before and after sorting of GFP+ cells. The procedure was performed once.
Extended Data Fig. 4 Enrichment of UMPS knockout T cells through 5-FOA.
a, Outline of the experiment used to simulate a mixed population of T cells with intact UMPS or bi-allelic UMPS knockout. Pure cell populations with the respective genotypes were stained with different tracking dyes (eFluor670 or CFSE), mixed and cultured in the presence or absence of 5-FOA during continuous T cell stimulation. b, Representative FACS plots (out of the 3 experiments) showing the mixed T cell populations before and 3 days after culture with 5-FOA. The relative percentages of the populations and the genotype of the cell populations identified by the dyes are annotated. c, Representative plots used for gating of viable T cells (plots 1 and 2) upstream of identification using the tracking dyes (plot 3) are shown. PI, propidium iodide. KO, knockout. WT, wild-type.
Extended Data Fig. 5 Dual-sgRNA editing of UMPS to increase full gene knockout in T cells and analysis strategy for in vivo samples.
a, Illustration of the genomic region with UMPS exon 1. The sgRNA binding sites in exon 1 are indicated, with dashed lines illustrating the cleavage sites. The cut sites of the two sgRNAs UMPS-1 and UMPS-7 are 89 bp apart in order to create a frameshift deletion, as UMPS is an enzyme consisting of 2 separate enzymatic functions and InDels in exon 1 that keep the open reading frame intact will not disrupt the function of ODC, which is encoded by the downstream part of the gene. b, InDel frequencies in human T cells electroporated with RNPs using sgRNA UMPS-7 (blue) or both UMPS-7 and UMPS-1 (red). The graph and error bars represent means and standard deviations. Statistical significance by comparison using two-sided t tests is annotated. Data from n = 4 (UMPS-7) and n = 3 (UMPS-7 + UMPS-1) biological replicates. c, InDel spectrum of T cells edited with one or two RNPs. The relatively low frequency of frameshift-mutations using only 1 sgRNA is due to the high frequency of 6 bp deletions. The increase in frameshift InDels is explained primarily by the high frequency of deletions of the 89-bp fragment between the two cut sites. Shown is 1 representative sample per condition. d, Gating strategy used to detect and quantify human T cells in mouse peripheral blood. Plot 1 shows gating of beads to be used as counting reference. Plot 2 shows all events excluding beads and gates on cells without debris. Plot 3 excludes dead cells. Plot 4 excludes events not expressing CD45. Plots 5 and 6 show the same population (CD45+ viable cells) with different axes combinations. huCD45, human CD45. mCD45, murine CD45. huTCR, human T cell receptor.
Supplementary information
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Rights and permissions
About this article
Cite this article
Wiebking, V., Patterson, J.O., Martin, R. et al. Metabolic engineering generates a transgene-free safety switch for cell therapy. Nat Biotechnol 38, 1441–1450 (2020). https://doi.org/10.1038/s41587-020-0580-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41587-020-0580-6
This article is cited by
-
Transient inhibition of 53BP1 increases the frequency of targeted integration in human hematopoietic stem and progenitor cells
Nature Communications (2024)
-
Tuning CARs: recent advances in modulating chimeric antigen receptor (CAR) T cell activity for improved safety, efficacy, and flexibility
Journal of Translational Medicine (2023)
-
Homology-independent targeted insertion (HITI) enables guided CAR knock-in and efficient clinical scale CAR-T cell manufacturing
Molecular Cancer (2023)
-
Stable expression of large transgenes via the knock-in of an integrase-deficient lentivirus
Nature Biomedical Engineering (2023)
-
CAR immune cells: design principles, resistance and the next generation
Nature (2023)