Abstract
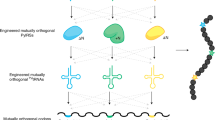
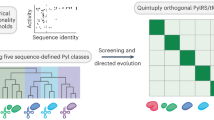
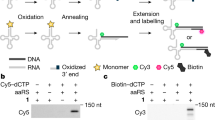
A central challenge in expanding the genetic code of cells to incorporate noncanonical amino acids into proteins is the scalable discovery of aminoacyl-tRNA synthetase (aaRS)–tRNA pairs that are orthogonal in their aminoacylation specificity. Here we computationally identify candidate orthogonal tRNAs from millions of sequences and develop a rapid, scalable approach—named tRNA Extension (tREX)—to determine the in vivo aminoacylation status of tRNAs. Using tREX, we test 243 candidate tRNAs in Escherichia coli and identify 71 orthogonal tRNAs, covering 16 isoacceptor classes, and 23 functional orthogonal tRNA–cognate aaRS pairs. We discover five orthogonal pairs, including three highly active amber suppressors, and evolve new amino acid substrate specificities for two pairs. Finally, we use tREX to characterize a matrix of 64 orthogonal synthetase–orthogonal tRNA specificities. This work expands the number of orthogonal pairs available for genetic code expansion and provides a pipeline for the discovery of additional orthogonal pairs and a foundation for encoding the cellular synthesis of noncanonical biopolymers.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
The tREX screening data used in this study are available in Supplementary Figs. 2–10, with tREX screening data for tRNA orthogonality in E. coli shown in Supplementary Figs. 2–7 and tREX screening data for aaRS activity on cognate tRNAs shown in Supplementary Figs. 8–10. Supplementary Table 4 has a complete list of the tRNAs generated by the filter described. Supplementary Table 5 lists tRNAs that were selected for experimental investigation, including tRNA accession numbers from tRNA-DB-CE and cognate synthetase accession numbers from NCBI Protein together with the sequences of the corresponding tREX probes. Source data for Figs. 3 and 7 are presented with the paper. All other datasets and material generated or analyzed in this study are available from the corresponding author upon reasonable request.
References
Chin, J. W. Expanding and reprogramming the genetic code. Nature 550, 53–60 (2017).
Young, D. D. & Schultz, P. G. Playing with the molecules of life. ACS Chem. Biol. 13, 854–870 (2018).
Liu, C. C. & Schultz, P. G. Adding new chemistries to the genetic code. Annu. Rev. Biochem. 79, 413–444 (2010).
Chin, J. W. Expanding and reprogramming the genetic code of cells and animals. Annu. Rev. Biochem. 83, 379–408 (2014).
Edwards, H. & Schimmel, P. An E. coli aminoacyl-tRNA synthetase can substitute for yeast mitochondrial enzyme function in vivo. Cell 51, 643–649 (1987).
Chin, J. W. et al. An expanded eukaryotic genetic code. Science 301, 964–967 (2003).
Wang, L., Magliery, T. J., Liu, D. R. & Schultz, P. G. A new functional suppressor tRNA/aminoacyl−tRNA synthetase pair for the in vivo incorporation of unnatural amino acids into proteins. J. Am. Chem. Soc. 122, 5010–5011 (2000).
Ambrogelly, A. et al. Pyrrolysine is not hardwired for cotranslational insertion at UAG codons. Proc. Natl Acad. Sci. USA 104, 3141–3146 (2007).
Park, H.-S. et al. Expanding the genetic code of Escherichia coli with phosphoserine. Science 333, 1151–1154 (2011).
Santoro, S. W., Anderson, J. C., Lakshman, V. & Schultz, P. G. An archaebacteria‐derived glutamyl‐tRNA synthetase and tRNA pair for unnatural amino acid mutagenesis of proteins in Escherichia coli. Nucleic Acids Res. 31, 6700–6709 (2003).
Pastrnak, M., Magliery, T. J. & Schultz, P. G. A new orthogonal suppressor tRNA/aminoacyl-tRNA synthetase pair for evolving an organism with an expanded genetic code. Helvetica Chim. Acta 83, 2277–2286 (2000).
Anderson, J. C. & Schultz, P. G. Adaptation of an orthogonal archaeal leucyl-tRNA and synthetase pair for four-base, amber, and opal suppression. Biochemistry 42, 9598–9608 (2003).
Anderson, J. C. et al. An expanded genetic code with a functional quadruplet codon. Proc. Natl Acad. Sci. USA 101, 7566–7571 (2004).
Chatterjee, A., Xiao, H. & Schultz, P. G. Evolution of multiple, mutually orthogonal prolyl-tRNA synthetase/tRNA pairs for unnatural amino acid mutagenesis in Escherichia coli. Proc. Natl Acad. Sci. USA 109, 14841–14846 (2012).
Steer, B. A. & Schimmel, P. Major anticodon-binding region missing from an archaebacterial tRNA synthetase. J. Biol. Chem. 274, 35601–35606 (1999).
Neumann, H., Peak-Chew, S. Y. & Chin, J. W. Genetically encoding N ε-acetyllysine in recombinant proteins. Nat. Chem. Biol. 4, 232–234 (2008).
Rogerson, D. T. et al. Efficient genetic encoding of phosphoserine and its nonhydrolyzable analog. Nat. Chem. Biol. 11, 496–503 (2015).
Hughes, R. A. & Ellington, A. D. Rational design of an orthogonal tryptophanyl nonsense suppressor tRNA. Nucleic Acids Res. 38, 6813–6830 (2010).
Chatterjee, A., Xiao, H., Yang, P. Y., Soundararajan, G. & Schultz, P. G. A tryptophanyl-tRNA synthetase/tRNA pair for unnatural amino acid mutagenesis in E. coli. Angew. Chem. Int. Ed. Engl. 52, 5106–5109 (2013).
Willis, J. C. W. & Chin, J. W. Mutually orthogonal pyrrolysyl-tRNA synthetase/tRNA pairs. Nat. Chem. 10, 831–837 (2018).
Wu, N., Deiters, A., Cropp, T. A., King, D. & Schultz, P. G. A genetically encoded photocaged amino acid. J. Am. Chem. Soc. 126, 14306–14307 (2004).
Abe, T. et al. tRNADB-CE: tRNA gene database well-timed in the era of big sequence data. Front. Genet. 5, 114 (2014).
Giegé, R., Sissler, M. & Florentz, C. Universal rules and idiosyncratic features in tRNA identity. Nucleic Acids Res. 26, 5017–5035 (1998).
Aldinger, C. A., Leisinger, A. K. & Igloi, G. L. The influence of identity elements on the aminoacylation of tRNAArg by plant and Escherichia coli arginyl-tRNA synthetases. FEBS J. 279, 3622–3638 (2012).
Larkin, D. C., Williams, A. M., Martinis, S. A. & Fox, G. E. Identification of essential domains for Escherichia coli tRNALeu aminoacylation and amino acid editing using minimalist RNA molecules. Nucleic Acids Res. 30, 2103–2113 (2002).
Quinn, C. L., Tao, N. & Schimmel, P. Species-specific microhelix aminoacylation by a eukaryotic pathogen tRNA synthetase dependent on a single base pair. Biochemistry 34, 12489–12495 (1995).
Nguyen, D. P., Garcia Alai, M. M., Kapadnis, P. B., Neumann, H. & Chin, J. W. Genetically encoding N ε-methyl-l-lysine in recombinant histones. J. Am. Chem. Soc. 131, 14194–14195 (2009).
Zamecnik, P. C., Stephenson, M. L. & Scott, J. F. Partial purification of soluble RNA. Proc. Natl Acad. Sci. USA 46, 811–822 (1960).
Rizzino, A. A. & Freundlich, M. Estimation of in vivo aminoacylation by periodate oxidation: tRNA alterations and iodate inhibition. Anal. Biochem. 66, 446–449 (1975).
Lodemann, E., Niedenthal, I. & Wacker, A. Influence of pH on the stability of some aminoacyl transfer ribonucleic acids and their elution pattern in chromatography on columns of methylated albumin adsorbed on kieselguhr. Z. Naturforsch. B 25, 845–848 (1970).
Hentzen, D., Mandel, P. & Garel, J.-P. Relation between aminoacyl-tRNA stability and the fixed amino acid. Biochim. Biophys. 281, 228–232 (1972).
Eiler, S., Dock-Bregeon, A., Moulinier, L., Thierry, J. C. & Moras, D. Synthesis of aspartyl-tRNAAsp in Escherichia coli—a snapshot of the second step. EMBO J. 18, 6532–6541 (1999).
Pedelacq, J. D., Cabantous, S., Tran, T., Terwilliger, T. C. & Waldo, G. S. Engineering and characterization of a superfolder green fluorescent protein. Nat. Biotechnol. 24, 79–88 (2006).
Kobayashi, T. et al. Structural basis for orthogonal tRNA specificities of tyrosyl-tRNA synthetases for genetic code expansion. Nat. Struct. Mol. Biol. 10, 425–432 (2003).
Wang, L., Xie, J., Deniz, A. A. & Schultz, P. G. Unnatural amino acid mutagenesis of green fluorescent protein. J. Org. Chem. 68, 174–176 (2003).
Kuratani, M. et al. Crystal structures of tyrosyl-tRNA synthetases from Archaea. J. Mol. Biol. 355, 395–408 (2006).
Strack, R. L. et al. A rapidly maturing far-red derivative of DsRed-Express2 for whole-cell labeling. Biochemistry 48, 8279–8281 (2009).
Brustad, E. et al. A genetically encoded boronate-containing amino acid. Angew. Chem. Int. Ed. Engl. 47, 8220–8223 (2008).
Wang, L., Brock, A., Herberich, B. & Schultz, P. G. Expanding the genetic code of Escherichia coli. Science 292, 498–500 (2001).
Xie, J. et al. The site-specific incorporation of p-iodo-l-phenylalanine into proteins for structure determination. Nat. Biotechnol. 22, 1297–1301 (2004).
Chin, J. W. et al. Addition of p-azido-l-phenylalanine to the genetic code of Escherichia coli. J. Am. Chem. Soc. 124, 9026–9027 (2002).
Zhang, M. S. et al. Biosynthesis and genetic encoding of phosphothreonine through parallel selection and deep sequencing. Nat. Methods 14, 729–736 (2017).
Varshney, U., Lee, C. P. & RajBhandary, U. L. Direct analysis of aminoacylation levels of tRNAs in vivo. Application to studying recognition of Escherichia coli initiator tRNA mutants by glutaminyl-tRNA synthetase. J. Biol. Chem. 266, 24712–24718 (1991).
Stenum, T. S., Sorensen, M. A. & Svenningsen, S. L. Quantification of the abundance and charging levels of transfer RNAs in Escherichia coli. J. Vis. Exp. https://doi.org/10.3791/56212 (2017).
Walker, S. E. & Fredrick, K. Preparation and evaluation of acylated tRNAs. Methods 44, 81–86 (2008).
Neumann, H., Wang, K., Davis, L., Garcia-Alai, M. & Chin, J. W. Encoding multiple unnatural amino acids via evolution of a quadruplet-decoding ribosome. Nature 464, 441–444 (2010).
Zhang, Y. et al. A semi-synthetic organism that stores and retrieves increased genetic information. Nature 551, 644–647 (2017).
Fredens, J. et al. Total synthesis of Escherichia coli with a recoded genome. Nature 569, 514–518 (2019).
Schmied, W. H. et al. Controlling orthogonal ribosome subunit interactions enables evolution of new function. Nature 564, 444–448 (2018).
Brubaker, L. H. & McCorquodale, D. J. The preparation of amino acid-transfer ribonucleic acid from Escherichia coli by direct phenol extraction of intact cells. Biochim. Biophys. Acta 76, 48–53 (1963).
Stemmer, W. P. & Morris, S. K. Enzymatic inverse PCR: a restriction site independent, single-fragment method for high-efficiency, site-directed mutagenesis. Biotechniques 13, 214–220 (1992).
Acknowledgements
This work was supported by the UK Medical Research Council (MRC; MC_U105181009 and MC_UP_A024_1008) and ERC-Advanced Grant SGCR, all to J.W.C. We thank W. Schmied, W. Robertson and R. Hegde for helpful discussions.
Author information
Authors and Affiliations
Contributions
D.C. designed and implemented tREX. D.C., S.T., J.C.W.W. and L.F.H.F. performed the tREX screening. D.C. and S.T. performed tRNA and synthetase characterization and engineering. S.D.F. and D.C. performed tRNA sequence analysis, with initial input from L.J.C. J.W.C. set the direction of research. J.W.C. and D.C. wrote the manuscript with input from the other authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–19, Table 1–7 legends and References
Supplementary Table 1
Complete list of identity elements used in this study, as described in the literature, thought to be recognized by the E. coli aaRSs.
Supplementary Table 3
Alignment of the D loop of each possible sequence in the tRNA-DB-CE database.
Supplementary Table 4
Complete list of tRNAs passing our filtering scheme, sorted by isoacceptor class.
Supplementary Table 5
tRNA scoring, tRNA detection, tRNA orthogonality, and synthetase and tREX probe sequences for experimentally tested tRNAs.
Supplementary Table 6
Raw data for quantification of sfGFP150TAG expression resulting from amber suppression by a given tRNA and aaRS.
Supplementary Table 7
Theoretical mass of the sfGFP variant used in this study after maturation of the chromophore.
Source data
Source Data Fig. 3
Full gel for the gel shown in Fig. 3c.
Source Data Fig. 7
Full gels for the gels shown in Fig. 7.
Rights and permissions
About this article
Cite this article
Cervettini, D., Tang, S., Fried, S.D. et al. Rapid discovery and evolution of orthogonal aminoacyl-tRNA synthetase–tRNA pairs. Nat Biotechnol 38, 989–999 (2020). https://doi.org/10.1038/s41587-020-0479-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41587-020-0479-2
This article is cited by
-
Adding α,α-disubstituted and β-linked monomers to the genetic code of an organism
Nature (2024)
-
tRNA therapeutics for genetic diseases
Nature Reviews Drug Discovery (2024)
-
Bioorthogonal Reactions in Bioimaging
Topics in Current Chemistry (2024)
-
Quintuply orthogonal pyrrolysyl-tRNA synthetase/tRNAPyl pairs
Nature Chemistry (2023)
-
Genetically programmed cell-based synthesis of non-natural peptide and depsipeptide macrocycles
Nature Chemistry (2023)