Abstract
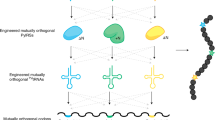
Mutually orthogonal aminoacyl transfer RNA synthetase/transfer RNA pairs provide a foundation for encoding non-canonical amino acids into proteins, and encoded non-canonical polymer and macrocycle synthesis. Here we discover quintuply orthogonal pyrrolysyl-tRNA synthetase (PylRS)/pyrrolysyl-tRNA (tRNAPyl) pairs. We discover empirical sequence identity thresholds for mutual orthogonality and use these for agglomerative clustering of PylRS and tRNAPyl sequences; this defines numerous sequence clusters, spanning five classes of PylRS/tRNAPyl pairs (the existing classes +N, A and B, and newly defined classes C and S). Most of the PylRS clusters belong to classes that were unexplored for orthogonal pair generation. By testing pairs from distinct clusters and classes, and pyrrolysyl-tRNAs with unusual structures, we resolve 80% of the pairwise specificities required to make quintuply orthogonal PylRS/tRNAPyl pairs; we control the remaining specificities by engineering and directed evolution. Overall, we create 924 mutually orthogonal PylRS/tRNAPyl pairs, 1,324 triply orthogonal pairs, 128 quadruply orthogonal pairs and 8 quintuply orthogonal pairs. These advances may provide a key foundation for encoded polymer synthesis.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All materials generated or analysed in this study are available from the corresponding author upon reasonable request. All generated datasets are provided in the Supplementary Information. Protein and nucleotide sequences were obtained from the NCBI Protein and NCBI Nucleotide databases, respectively.
Code availability
The code for PylRS and tRNAPyl clustering and mutually orthogonal PylRS/tRNAPyl pair identification is available at https://github.com/JWChin-Lab/Quint-Pyl.
References
Chin, J. W. Expanding and reprogramming the genetic code. Nature 550, 53–60 (2017).
De La Torre, D. & Chin, J. W. Reprogramming the genetic code. Nat. Rev. Genet. 22, 169–184 (2021).
Robertson, W. E. et al. Sense codon reassignment enables viral resistance and encoded polymer synthesis. Science 372, 1057–1062 (2021).
Spinck, M. et al. Genetically programmed cell-based synthesis of non-natural peptide and depsipeptide macrocycles. Nat. Chem. 15, 61–69 (2023).
Cervettini, D. et al. Rapid discovery and evolution of orthogonal aminoacyl-tRNA synthetase–tRNA pairs.Nat. Biotechnol. 38, 989–999 (2020).
Dunkelmann, D. L., Willis, J. C. W., Beattie, A. T. & Chin, J. W. Engineered triply orthogonal pyrrolysyl–tRNA synthetase/tRNA pairs enable the genetic encoding of three distinct non-canonical amino acids. Nat. Chem. 12, 535–544 (2020).
Willis, J. C. W. & Chin, J. W. Mutually orthogonal pyrrolysyl-tRNA synthetase/tRNA pairs. Nat. Chem. 10, 831–837 (2018).
Srinivasan, G., James, C. M. & Krzycki, J. A. Pyrrolysine encoded by UAG in archaea: charging of a UAG-decoding specialized tRNA. Science 296, 1459–1462 (2002).
Krzycki, J. A. The direct genetic encoding of pyrrolysine. Curr. Opin. Microbiol. 8, 706–712 (2005).
Neumann, H., Peak-Chew, S. Y. & Chin, J. W. Genetically encoding Nε-acetyllysine in recombinant proteins. Nat. Chem. Biol. 4, 232–234 (2008).
Wang, L., Brock, A., Herberich, B. & Schultz, P. G. Expanding the genetic code of Escherichia coli. Science 292, 498–500 (2001).
Borrel, G. et al. Unique characteristics of the pyrrolysine system in the 7th order of methanogens: implications for the evolution of a genetic code expansion cassette. Archaea 2014, 374146 (2014).
Park, H.-S. et al. Expanding the genetic code of Escherichia coli with phosphoserine. Science 333, 1151–1154 (2011).
Rogerson, D. T. et al. Efficient genetic encoding of phosphoserine and its nonhydrolyzable analog. Nat. Chem. Biol. 11, 496–503 (2015).
Hughes, R. A. & Ellington, A. D. Rational design of an orthogonal tryptophanyl nonsense suppressor tRNA. Nucleic Acids Res. 38, 6813–6830 (2010).
Chatterjee, A., Sun, S. B., Furman, J. L., Xiao, H. & Schultz, P. G. A versatile platform for single- and multiple-unnatural amino acid mutagenesis in Escherichia coli. Biochemistry 52, 1828–1837 (2013).
Italia, J. S. et al. Mutually orthogonal nonsense-suppression systems and conjugation chemistries for precise protein labeling at up to three distinct sites. J. Am. Chem. Soc. 141, 6204–6212 (2019).
Neumann, H., Wang, K., Davis, L., Garcia-Alai, M. & Chin, J. W. Encoding multiple unnatural amino acids via evolution of a quadruplet-decoding ribosome. Nature 464, 441–444 (2010).
Wang, K. et al. Optimized orthogonal translation of unnatural amino acids enables spontaneous protein double-labelling and FRET. Nat. Chem. 6, 393–403 (2014).
Anderson, J. C. et al. An expanded genetic code with a functional quadruplet codon. Proc. Natl Acad. Sci. USA 101, 7566–7571 (2004).
Dunkelmann, D. L., Oehm, S. B., Beattie, A. T. & Chin, J. W. A 68-codon genetic code to incorporate four distinct non-canonical amino acids enabled by automated orthogonal mRNA design. Nat. Chem. 13, 1110–1117 (2021).
Malyshev, D. A. et al. A semi-synthetic organism with an expanded genetic alphabet. Nature 509, 385–388 (2014).
Fischer, E. C. et al. New codons for efficient production of unnatural proteins in a semisynthetic organism. Nat. Chem. Biol. 16, 570–576 (2020).
Zhang, Y. et al. A semi-synthetic organism that stores and retrieves increased genetic information. Nature 551, 644–647 (2017).
Fredens, J. et al. Total synthesis of Escherichia coli with a recoded genome. Nature 569, 514–518 (2019).
Wang, K. et al. Defining synonymous codon compression schemes by genome recoding. Nature 539, 59–64 (2016).
Neumann, H., Slusarczyk, A. L. & Chin, J. W. De novo generation of mutually orthogonal aminoacyl-tRNA synthetase/tRNA pairs. J. Am. Chem. Soc. 132, 2142–2144 (2010).
Beranek, V., Willis, J. C. W. & Chin, J. W. An evolved Methanomethylophilus alvus pyrrolysyl-tRNA synthetase/tRNA pair is highly active and orthogonal in mammalian cells. Biochemistry 58, 387–390 (2019).
Chatterjee, A., Xiao, H. & Schultz, P. G. Evolution of multiple, mutually orthogonal prolyl-tRNA synthetase/tRNA pairs for unnatural amino acid mutagenesis in Escherichia coli. Proc. Natl Acad. Sci. USA 109, 14841–14846 (2012).
Italia, J. S. et al. An orthogonalized platform for genetic code expansion in both bacteria and eukaryotes. Nat. Chem. Biol. 13, 446–450 (2017).
Chin, J. W. Expanding and reprogramming the genetic code of cells and animals. Annu. Rev. Biochem. 83, 379–408 (2014).
Ambrogelly, A. et al. Pyrrolysine is not hardwired for cotranslational insertion at UAG codons. Proc. Natl Acad. Sci. USA 104, 3141–3146 (2007).
Elliott, T. S. et al. Proteome labeling and protein identification in specific tissues and at specific developmental stages in an animal. Nat. Biotechnol. 32, 465–472 (2014).
Suzuki, T. et al. Crystal structures reveal an elusive functional domain of pyrrolysyl-tRNA synthetase. Nat. Chem. Biol. 13, 1261–1266 (2017).
Kobayashi, T., Yanagisawa, T., Sakamoto, K. & Yokoyama, S. Recognition of non-α-amino aubstrates by pyrrolysyl-tRNA synthetase. J. Mol. Biol. 385, 1352–1360 (2009).
Polycarpo, C. R. et al. Pyrrolysine analogues as substrates for pyrrolysyl-tRNA synthetase. FEBS Lett. 580, 6695–6700 (2006).
Bindman, N. A., Bobeica, S. C., Liu, W. R. & Van Der Donk, W. A. Facile removal of leader peptides from lanthipeptides by incorporation of a hydroxy acid. J. Am. Chem. Soc. 137, 6975–6978 (2015).
Li, Y.-M. et al. Ligation of expressed protein α-hydrazides via genetic incorporation of an α-hydroxy acid. ACS Chem. Biol. 7, 1015–1022 (2012).
Ohtake, K. et al. Engineering an automaturing transglutaminase with enhanced thermostability by genetic code expansion with two codon reassignments. ACS Synth. Biol. 7, 2170–2176 (2018).
Polycarpo, C. et al. An aminoacyl-tRNA synthetase that specifically activates pyrrolysine. Proc. Natl Acad. Sci. USA 101, 12450–12454 (2004).
Nozawa, K. et al. Pyrrolysyl-tRNA synthetase–tRNAPyl structure reveals the molecular basis of orthogonality. Nature 457, 1163–1167 (2008).
Herring, S. et al. The amino-terminal domain of pyrrolysyl-tRNA synthetase is dispensable in vitro but required for in vivo activity. FEBS Lett. 581, 3197–3203 (2007).
Jiang, R. & Krzycki, J. A. PylSn and the homologous N-terminal domain of pyrrolysyl-tRNA synthetase bind the tRNA that is essential for the genetic encoding of pyrrolysine. J. Biol. Chem. 287, 32738–32746 (2012).
Meineke, B., Heimgärtner, J., Eirich, J., Landreh, M. & Elsässer, S. J. Site-specific incorporation of two ncAAs for two-color bioorthogonal labeling and crosslinking of proteins on live mammalian cells. Cell Rep. 31, 107811 (2020).
Meineke, B., Heimgärtner, J., Lafranchi, L. & Elsässer, S. J. Methanomethylophilus alvus Mx1201 provides basis for mutual orthogonal pyrrolysyl tRNA/aminoacyl-tRNA synthetase pairs in mammalian cells. ACS Chem. Biol. 13, 3087–3096 (2018).
Zhang, H. et al. The tRNA discriminator base defines the mutual orthogonality of two distinct pyrrolysyl-tRNA synthetase/tRNAPyl pairs in the same organism. Nucleic Acids Res. 50, 4601–4615 (2022).
Fischer, J. T., Söll, D. & Tharp, J. M. Directed evolution of Methanomethylophilus alvus pyrrolysyl-tRNA synthetase generates a hyperactive and highly selective variant. Front. Mol. Biosci. 9, 850613 (2022).
Tharp, J. M., Vargas-Rodriguez, O., Schepartz, A. & Söll, D. Genetic encoding of three distinct noncanonical amino acids using reprogrammed initiator and nonsense codons. ACS Chem. Biol. 16, 766–774 (2021).
Laslett, D. ARAGORN, a program to detect tRNA genes and tmRNA genes in nucleotide sequences. Nucleic Acids Res. 32, 11–16 (2004).
Katayama, H., Nozawa, K., Nureki, O., Nakahara, Y. & Hojo, H. Pyrrolysine analogs as substrates for bacterial pyrrolysyl-tRNA synthetase in vitro and in vivo. Biosci. Biotechnol. Biochem. 76, 205–208 (2012).
Varani, G. & McClain, W. H. The G·U wobble base pair. EMBO Rep. 1, 18–23 (2000).
Sievers, F. et al. Fast, scalable generation of high‐quality protein multiple sequence alignments using Clustal Omega. Mol. Syst. Biol. 7, 539 (2011).
Guan, Y., Haroon, M. F., Alam, I., Ferry, J. G. & Stingl, U. Single-cell genomics reveals pyrrolysine-encoding potential in members of uncultivated archaeal candidate division MSBL1. Environ. Microbiol. Rep. 9, 404–410 (2017).
Cock, P. J. A. et al. Biopython: freely available Python tools for computational molecular biology and bioinformatics. Bioinformatics 25, 1422–1423 (2009).
Gruber, A. R., Lorenz, R., Bernhart, S. H., Neubock, R. & Hofacker, I. L. The Vienna RNA websuite. Nucleic Acids Res. 36, W70–W74 (2008).
Lorenz, R. et al. ViennaRNA package 2.0. Algorithms Mol. Biol. 6, 26 (2011).
Pedregosa, F. et al. Scikit-learn: machine learning in Python. J.Mach. Learn. Res. 12, 2825–2830 (2011).
Reis, A. C. & Salis, H. M. An automated model test system for systematic development and improvement of gene expression models. ACS Synth. Biol. 9, 3145–3156 (2020).
Cetnar, D. P. & Salis, H. M. Systematic quantification of sequence and structural determinants controlling mRNA stability in bacterial operons. ACS Synth. Biol. 10, 318–332 (2021).
Espah Borujeni, A. et al. Precise quantification of translation inhibition by mRNA structures that overlap with the ribosomal footprint in N-terminal coding sequences. Nucleic Acids Res. 45, 5437–5448 (2017).
Espah Borujeni, A. & Salis, H. M. Translation initiation is controlled by RNA folding kinetics via a ribosome drafting mechanism. J. Am. Chem. Soc. 138, 7016–7023 (2016).
Espah Borujeni, A., Channarasappa, A. S. & Salis, H. M. Translation rate is controlled by coupled trade-offs between site accessibility, selective RNA unfolding and sliding at upstream standby sites. Nucleic Acids Res. 42, 2646–2659 (2013).
Salis, H. M., Mirsky, E. A. & Voigt, C. A. Automated design of synthetic ribosome binding sites to control protein expression. Nat. Biotechnol. 27, 946–950 (2009).
Acknowledgements
This work was supported by the Medical Research Council (MRC), UK (MC_U105181009 and MC_UP_A024_1008) and an ERC Advanced Grant SGCR, all to J.W.C. For the purpose of Open Access, the MRC Laboratory of Molecular Biology has applied a CC BY public copyright licence to any Author Accepted Manuscript (AAM) version arising from this submission. D.L.D. was supported by the Boehringer Ingelheim Fonds and Magdalene College, Cambridge.
Author information
Authors and Affiliations
Contributions
D.L.D., A.T.B. and J.W.C. designed the project. A.T.B. and D.L.D. performed the experiments. A.T.B. generated the computational discovery pipeline with inputs from D.L.D. A.T.B., D.L.D and J.W.C. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare the following competing financial interest: J.W.C. is a founder of the company Constructive Bio. The Medical Research Council have filed a patent application on the basis of this work.
Peer review
Peer review information
Nature Chemistry thanks Heinz Neumann, Jiangyun Wang and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Structures of non-canonical amino acids used in this study.
a. Chemical structure of Nε-tert-butyloxycarbonyl-l-lysine BocK 1. b. Chemical structure of N6-((allyloxy)carbonyl)-l-lysine (AllocK) 2.
Extended Data Fig. 2 Screening class N, A and B Pyl tRNAs for orthogonality to synthetases from other classes.
a. Screening of previously reported class N Pyl tRNAs against key active PylRS enzymes from each class (and S∆ PylRS variants). Most class N Pyl tRNAs tested are highly specific to N+-MmPylRS. Each heatmap value represents the average of three biological replicates. All numerical values and bar charts including error bars showing s.d. are provided (Supplementary Table 2). b. Screening of previously reported A-AlvtRNAPyl mutants against key active PylRS enzymes from each class (and S∆ PylRS variants). Multiple mutants are highly specific to A∆-1R26PylRS. Each heatmap value represents the average of three biological replicates. All numerical values and bar charts including error bars showing s.d. are provided (Supplementary Table 2). c. Screening of previously reported B-InttRNAPyl mutants against key active PylRS enzymes from each class (and S∆ PylRS variants). The activity of B∆-Lum1PylRS with class B Pyl tRNAs is closely matched by S∆-ClosPylRS, S∆-I2PylRS, and – most problematically – C∆-NitraPylRS. Each heatmap value represents the average of three biological replicates. All numerical values and bar charts including error bars showing s.d. are provided (Supplementary Table 2).
Extended Data Fig. 3 Families of quadruply orthogonal pairs containing both S∆B and S∆C PylRSs.
a. Activity heatmap of the two (overlapping) quadruplet families with both class B and class C PylRS enzymes substituted by S∆ PylRS variants (labelled in two shades of blue), shown along with the most orthogonal fifth pair from the final class (class N or class S). The tRNAPyl that requires engineering or replacement to abolish unwanted cross-reactions is labelled in red, while the Pyl tRNAs that already satisfy all necessary orthogonality requirements are labelled in green. Each heatmap value represents the average of three biological replicates. All numerical values and bar charts including error bars showing s.d. are provided (Supplementary Table 2).b. Schematic of the interactions within the quadruplets from a (both o.c. 2.9). For both, cross-reactivity (red arrow) between class N and class S must be eliminated to yield quintuply orthogonal pairs. In addition, although the problematic cross-reaction between class C PylRS and class B tRNAPyl has been diminished by substitution of both class B and C pairs by pairs formed with a S∆ PylRS variant, the remaining cross-reactivity (yellow arrow) limits the o.c. of these sets. Each heatmap value represents the average of three biological replicates. All numerical values and bar charts including error bars showing s.d. are provided (Supplementary Table 2).
Extended Data Fig. 4 Directed evolution of Pyl tRNAs and resulting quadruply orthogonal pairs.
a. Heatmap showing the activity of hits obtained from positive selection of the library with S∆B-ClosPylRS followed by successive negative screening with the PylRS enzymes from classes N, A, C and S. Each heatmap value represents the average of three biological replicates. All numerical values and bar charts including error bars showing s.d. are provided (Supplementary Table 2). b. Heatmap showing the activity of the tRNAPyl hit S-I2tRNAPyl-S52 obtained from positive selection of the library with S+-DebPylRS followed by successive negative screening with the PylRS enzymes from the other classes. This tRNAPyl exhibits selective activity with S+-DebPylRS. Each heatmap value represents the average of three biological replicates. All numerical values and bar charts including error bars showing s.d. are provided (Supplementary Table 2). c. Activity heatmaps from each family of quadruply orthogonal PylRS/tRNAPyl pairs obtained following the tRNAPyl directed evolution strategy; the quadruplets with the highest o.c. are shown. Dark grey box: quadruply orthogonal families discovered in Fig. 5. Light grey box: quadruply orthogonal family discovered in Fig. 5, but which has a higher o.c. when incorporating a newly evolved tRNAPyl. Orthogonality coefficient, o.c., is shown in grey. Each heatmap value represents the average of three biological replicates. All numerical values and bar charts including error bars showing s.d. are provided (Supplementary Table 2).
Supplementary information
Supplementary Information
Supplementary Notes 1–3, Figs. 1–10, and Tables 4 and 5.
Supplementary Tables
Supplementary Tables 1 (sheets 1–4), 2 (sheets 5–11), 3 (sheets 12–14), 6 (sheets 15 and 16) and 7 (sheets 17 and 18).
Supplementary Code 1
Custom scripts used for aaRS and tRNA clustering, and identification of sets of mutually orthogonal aaRS/tRNA pairs.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Beattie, A.T., Dunkelmann, D.L. & Chin, J.W. Quintuply orthogonal pyrrolysyl-tRNA synthetase/tRNAPyl pairs. Nat. Chem. 15, 948–959 (2023). https://doi.org/10.1038/s41557-023-01232-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41557-023-01232-y