Abstract
Expansions of short tandem repeats are genetic variants that have been implicated in several neuropsychiatric and other disorders, but their assessment remains challenging with current polymerase-based methods1,2,3,4. Here we introduce a CRISPR–Cas-based enrichment strategy for nanopore sequencing combined with an algorithm for raw signal analysis. Our method, termed STRique for short tandem repeat identification, quantification and evaluation, integrates conventional sequence mapping of nanopore reads with raw signal alignment for the localization of repeat boundaries and a hidden Markov model-based repeat counting mechanism. We demonstrate the precise quantification of repeat numbers in conjunction with the determination of CpG methylation states in the repeat expansion and in adjacent regions at the single-molecule level without amplification. Our method enables the study of previously inaccessible genomic regions and their epigenetic marks.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
All sequencing data generated in this study and utilized for the determination of FMR1 CGGn- and C9orf72 (G4C2)n-repeat expansion lengths and methylation status in plasmids, BACs and patient DNA are available in a Figshare repository with identifier 7205666.
Whole-genome sequencing data and associated uncropped Southern blot images, size-marker standard curves and ethidium bromide imaging data from patient-derived cell lines are available from the corresponding author upon reasonable request through a material transfer agreement protecting the participants’ genomic privacy.
Code availability
All custom code developed for this study is under MIT license and is available at https://github.com/giesselmann/STRique.
The RepeatHMM package was forked and modified and is available at https://github.com/giesselmann/RepeatHMM.
References
DeJesus-Hernandez, M. et al. Expanded GGGGCC hexanucleotide repeat in noncoding region of C9ORF72 causes chromosome 9p-linked FTD and ALS. Neuron 72, 245–256 (2011).
Renton, A. E. et al. A hexanucleotide repeat expansion in C9ORF72 is the cause of chromosome 9p21-linked ALS-FTD. Neuron 72, 257–268 (2011).
Crook, A. et al. The C9orf72 hexanucleotide repeat expansion presents a challenge for testing laboratories and genetic counseling. Amyotroph. Lateral Scler. Frontotemporal Degener. 20, 310–316 (2019).
Klepek, H., Goutman, S. A., Quick, A., Kolb, S. J. & Roggenbuck, J. Variable reporting of C9orf72 and a high rate of uncertain results in ALS genetic testing. Neurol. Genet. 5, e301 (2019).
Gatchel, J. R. & Zoghbi, H. Y. Diseases of unstable repeat expansion: mechanisms and common principles. Nat. Rev. Genet. 6, 743–755 (2005).
Paulson, H. Repeat expansion diseases. Handb. Clin. Neurol. 147, 105–123 (2018).
Verkerk, A. J. et al. Identification of a gene (FMR-1) containing a CGG repeat coincident with a breakpoint cluster region exhibiting length variation in fragile X syndrome. Cell 65, 905–914 (1991).
van Blitterswijk, M. et al. Association between repeat sizes and clinical and pathological characteristics in carriers of C9ORF72 repeat expansions (Xpansize-72): a cross-sectional cohort study. Lancet Neurol. 12, 978–988 (2013).
Xi, Z. et al. Hypermethylation of the CpG island near the G4C2 repeat in ALS with a C9orf72 expansion. Am. J. Hum. Genet. 92, 981–989 (2013).
Russ, J. et al. Hypermethylation of repeat expanded C9orf72 is a clinical and molecular disease modifier. Acta Neuropathol. 129, 39–52 (2015).
Deamer, D., Akeson, M. & Branton, D. Three decades of nanopore sequencing. Nat. Biotechnol. 34, 518–524 (2016).
Brown, C. G. & Clarke, J. Nanopore development at Oxford Nanopore. Nat. Biotechnol. 34, 810–811 (2016).
Jain, M. et al. Nanopore sequencing and assembly of a human genome with ultra-long reads. Nat. Biotechnol. 36, 338–345 (2018).
Simpson, J. T. et al. Detecting DNA cytosine methylation using nanopore sequencing. Nat. Methods 14, 407–410 (2017).
Rand, A. C. et al. Mapping DNA methylation with high-throughput nanopore sequencing. Nat. Methods 14, 411–413 (2017).
Mizielinska, S. et al. C9orf72 repeat expansions cause neurodegeneration in Drosophila through arginine-rich proteins. Science 345, 1192–1194 (2014).
Dashnow, H. et al. STRetch: detecting and discovering pathogenic short tandem repeat expansions. Genome Biol. 19, 121 (2018).
Liu, Q., Zhang, P., Wang, D., Gu, W. & Wang, K. Interrogating the ‘unsequenceable’ genomic trinucleotide repeat disorders by long-read sequencing. Genome Med. 9, 65 (2017).
Wick, R.R., Judd, L.M. & Holt, K.E. Performance of neural network basecalling tools for Oxford Nanopore sequencing. Genome Biol. 20, 129 (2019).
Li, H. Minimap2: pairwise alignment for nucleotide sequences. Bioinformatics 3, 321 (2018).
Schreiber, J. & Karplus, K. Analysis of nanopore data using hidden Markov models. Bioinformatics 31, 1897–1903 (2015).
O’Rourke, J. G. et al. C9orf72 BAC transgenic mice display typical pathologic features of ALS/FTD. Neuron 88, 892–901 (2015).
Zetsche, B. et al. Cpf1 is a single RNA-guided endonuclease of a class 2 CRISPR–Cas system. Cell 163, 759–771 (2015).
Boland, M. J. et al. Molecular analyses of neurogenic defects in a human pluripotent stem cell model of fragile X syndrome. Brain 140, 582–598 (2017).
Pattanayak, V. et al. High-throughput profiling of off-target DNA cleavage reveals RNA-programmed Cas9 nuclease specificity. Nat. Biotechnol. 31, 839–843 (2013).
Hornstra, L. K., Nelson, D. L., Warren, S. T. & Yang, T. P. High resolution methylation analysis of the FMR1 gene trinucleotide repeat region in fragile X syndrome. Hum. Mol. Genet. 2, 1659–1665 (1993).
Xi, Z. et al. The C9orf72 repeat expansion itself is methylated in ALS and FTLD patients. Acta Neuropathol. 129, 715–727 (2015).
Lyons, J. I., Kerr, G. R. & Mueller, P. W. Fragile X syndrome: scientific background and screening technologies. J. Mol. Diagn. 17, 463–471 (2015).
Hansen, R. S., Gartler, S. M., Scott, C. R., Chen, S.-H. & Laird, C. M. Methylation analysis of CGG sites in the CpG island of the human FMR1 gene. Hum. Mol. Genet. 1, 571–578 (1992).
Gabrieli, T. et al. Selective nanopore sequencing of human BRCA1 by Cas9-assisted targeting of chromosome segments (CATCH). Nucleic Acids Res. 46, e87 (2018).
Reinert, K. et al. The SeqAn C++ template library for efficient sequence analysis: a resource for programmers. J. Biotechnol. 261, 157–168 (2017).
Giesselmann, P., Hetzel, S., Müller, F.-J., Meissner, A. & Kretzmer, H. Nanopype: a modular and scalable nanopore data processing pipeline. Bioinformatics 26, 2204 (2019).
Rohrandt, C. et al. Nanopore SimulatION–a raw data simulator for nanopore sequencing. in 2018 IEEE International Conference on Bioinformatics and Biomedicine (BIBM) 1–8 (IEEE, 2019).
R Core Team. R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, 2019).
Gu, Z., Eils, R. & Schlesner, M. Complex heat maps reveal patterns and correlations in multidimensional genomic data. Bioinformatics 32, 2847–2849 (2016).
Gu, Z., Gu, L., Eils, R., Schlesner, M. & Brors, B. Circlize implements and enhances circular visualization in R. Bioinformatics 30, 2811–2812 (2014).
Sambrook, J. & Maniatis, T. Molecular Cloning (Cold Spring Harbor Laboratory Press, 1989).
Labun, K., Montague, T. G., Gagnon, J. A., Thyme, S. B. & Valen, E. CHOPCHOP v2: a web tool for the next generation of CRISPR genome engineering. Nucleic Acids Res. 44, W272–W276 (2016).
Montague, T. G., Cruz, J. M., Gagnon, J. A., Church, G. M. & Valen, E. CHOPCHOP: a CRISPR/Cas9 and TALEN web tool for genome editing. Nucleic Acids Res. 42, W401–W407 (2014).
Chen, J. S. et al. CRISPR–Cas12a target binding unleashes indiscriminate single-stranded DNase activity. Science 360, 436–439 (2018).
Kwok, Y. K. et al. Validation of a robust PCR-based assay for quantifying fragile X CGG repeats. Clin. Chim. Acta 456, 137–143 (2016).
Chen, G. et al. Chemically defined conditions for human iPSC derivation and culture. Nat. Methods 8, 424–429 (2011).
Mertens, J. et al. APP processing in human pluripotent stem cell-derived neurons is resistant to NSAID-based γ-secretase modulation. Stem Cell Rep. 1, 491–498 (2013).
Zhou, Y., Kumari, D., Sciascia, N. & Usdin, K. CGG-repeat dynamics and FMR1 gene silencing in fragile X syndrome stem cells and stem cell-derived neurons. Mol. Autism 7, 165 (2016).
Acknowledgements
We are deeply thankful for the invaluable support by the patients with c9FTD/ALS and FXS and their families who donated biomaterials for this study. The C9orf72 BAC was generously provided by R. Baloh and S. Bell (Cedars Sinai Medical Center, Los Angeles, CA, USA). We are grateful to J. Loring and A. Zhang (Scripps Research Institute, La Jolla, CA, USA) for providing us with hiPSC lines from a patient with FXS (supported by NIH R33MH087925-03). We thank P. van Damme and W. Robberecht (Laboratory for Neurobiology; VIB-KU Leuven Center for Brain & Disease Research, Belgium) for providing the fibroblasts derived from patients with c9FTD/ALS used for reprogramming. We acknowledge the expert assistance of the technical staff of the Molecular Genetics Laboratory of the Institute of Human Genetics (Ulm, Germany). P.K. and J.L. acknowledge financial support by the Hector Stiftung II gGmbH. F.J.M. and R.T. received funding from the Deutsche Forschungsgemeinschaft (German Research Foundation) under Germany’s Excellence Strategy–EXC 22167-390884018. F.J.M. and B.M.S. were supported by the BMBF (PluriTest2, 13GW0128A). This work was supported by the Max Planck Society. This work was overseen and approved by the Ethics Committee of the Christian-Albrechts-University (Kiel, Germany; reference no. A 145/11). Informed consent was obtained from all donors of cells and tissues used for the generation of hiPSC lines and their subsequent genetic and epigenetic analysis. All materials were donated graciously by our patients.
Author information
Authors and Affiliations
Contributions
P.G., B.M.S. and F.J.M. conceived the project. B.B. and R.T. performed cell culture as well as plasmid and BAC expansion and extraction. P.G. wrote the STRique pipeline. P.G., B.M.S., C.R. and H.K. conducted additional bioinformatic analyses. P.K. and J.L. reprogrammed the c9FTD/ALS hiPSCs from patient fibroblasts used in this study. E.R., R.B., A.H. and J.E.G. developed the Cas12a and Cas9 protocols. B.B. further developed the Cas12a and Cas9 protocols with DNA from patients with c9FTD/ALS and FXS and performed nanopore library preparation and nanopore sequencing for the results presented in this manuscript. R.T. and C.G. worked on the optimization of aspects of the enrichment protocol. G.A. and R.S. conducted diagnostic testing of the repeat expansions by Southern blot and PCR analyses. S.S., R.S., O.A. and G.A. provided clinical and diagnostic advice. P.G., B.B., B.M.S., A.M. and F.J.M. wrote the manuscript. F.J.M. oversaw the study. All authors contributed to the editing and completion of the manuscript.
Corresponding author
Ethics declarations
Competing interests
E.R., R.B., A.H. and J.E.G. are employees of ONT. C.R. was reimbursed for travel costs for an invited talk at the Nanopore Days 2018 conference in Heidelberg (Germany) by ONT. P.G. was reimbursed for travel costs for an invited talk at the London Calling 2019 conference. ONT had no role in the study design, interpretation of results or writing of the manuscript.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Systematic overview over tandem repeats and their nomenclature.
Schematic depiction of terms used in the literature and this manuscript for different classes of tandem repeats. N = nucleotide in tandem repeat motif, lower case number indicates the number of nucleotides in any given tandem repeat motif. Red box indicates the tandem repeat motif classes studied in this work. The referenced studies are in order of the tandem repeat motif length listed in the schematic: [1] Nature Genetics, Ishiura et al., 2018. [2] Molecular Neurodegeneration, Ebbert et al., 2018. [3] bioRxiv preprint, de Roek et al., 2018. [4] Nature Biotechnology, Jain, Olsen et al. 2018.
Supplementary Figure 2 Comparison of timelines between nanopore sequencing and Southern blotting for diagnostic purposes.
Upper timeline depicts steps involved in nanopore sequencing with Cas-based enrichment with a total duration of approximately 16 hours. Lower timeline depicts steps involved in Southern blot analysis of the same repeat expansion with a total duration of about three days. For the detailed protocols see Methods.
Supplementary Figure 3 Nanopore signal processing with STRique.
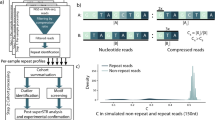
a) Signal alignment detecting prefix and suffix flanking the repeat using a capped distance score function and a semi-global alignment. A compound profile HMM of prefix, a single repeat and the suffix sequence assigns either prefix, repeat or suffix label to each signal value. Repeat counts are obtained through dummy states between repeat and suffix. b) Nanopore signal profile HMM with normal distributed match state and uniform distributed insertion state emission probabilities.
Supplementary Figure 4 Evaluation of repeat quantification approaches.
a) Correlation of manual counted repeat lengths with sequence base methods. Decoy alignment against reference with 3-100 repeats with Albacore (window 10k and 16k), Guppy (fast and hac mode, 1k and 16k window size) and Flappie base-calling (n=204). b) Correlation of manual count with RepeatHMM and STRique results (n=204). c) Manual counted set of plasmid reads on y-axis correlating with guppy base-calling and decoy alignment approach, RepeatHMM and STRique raw signal pipeline on x-axis. Only data points shown which could be evaluated with all four methods (n=15, 49, 45, 48, 47; Pearson correlation).
Supplementary Figure 5 Strand bias in sequence based repeat counts.
Comparison of repeat counts from STRique, decoy alignment based on guppy (high accuracy model, 16k window size) and repeatHMM based on guppy (high accuracy model, 16k window size) for BAC data. One dot (n=5004) per read passing all three approaches and colored by strand.
Supplementary Figure 6 Nanopore raw signal of the C9orf72 STR in NA12878 cells.
Compound multi signal HMM alignment of publicly available raw traces from two template and eight complement reads from the NA12878 cell line shows matching signal pattern in all reads [Nature Biotechnology, Jain, Koren et al., 2018]. Displayed are the current measurements as dots and the model signal as black line. Blue dots indicate current measurements identified as prefix or suffix (see Supplementary Fig. 1). Red dots indicate raw current measurements identified by STRique as belonging to the C9orf72-(G4C2)n-STR. STRique detects in this case five (G4C2)-repeats.
Supplementary Figure 7 Repeat count cluster stability over experiments.
a) C9orf72 target enrichment flow cells for patient 24/5#2 b) FMR1 enrichment flow cells of SC105iPS6/iPS7. (FA*: MinION, PAD*: PromethION, number of reads per boxplot in (a-b) are in Supplementary Tables 5-6 column wt and exp) c) Mean coverage on target per flow cell (dots) compared to genome wide mean coverage of 100k tiles for WGS, Cas12 and Cas9 enrichment on MinION and PromethION (boxplots, n=30971 tiles). Data in (a-c) presented as boxplots (centerline, median; box limits, first and third quartiles; whiskers, 1.5× interquartile range; outliers not shown).
Supplementary Figure 8 Southern blot analysis of c9orf72 and FMR1 repeat expansion.
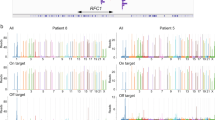
a) Autoradiographic Southern blot from controls and C9orf72 expanded allele carriers. Unmodified scan of a Southern blot analysis of the cell line used for this study. b) Autoradiographic Southern blot from controls and C9orf72 expanded allele carriers with labels. Same scan of a Southern blot analysis of the cell line used for this study as in Supplementary Fig. 9a with labels indicating the sample names and function (e.g. negative control, mutation carrier). Legend: hiPSC = human induced pluripotent stem cell; HMW-DNA = high molecular weight DNA; -m = methylated allele; wt = wildtype; yellow text/lines = hiPSC lines used in this study c) Autoradiographic Southern blot from controls and FMR1 expanded allele carriers. Unmodified scan of a Southern blot analysis of the cell line used for this study. d) Autoradiographic Southern blot from controls and FMR1 expanded allele carriers with labels. Same scan of a Southern blot analysis of the cell line used for this study as in Supplementary Fig. 6e with labels indicating the sample names and function (e.g. negative control, mutation carrier). Fragments covering the FMR1 CGG-STR migrate in this assay depending on their methylation status: methylated fragments as the NruI restriction enzyme does not cut if its 5′-TCGCGA-3′ restriction site is methylated at the CpG site. As a result, the restriction fragment is longer and spans from the two adjacent HindIII restriction sites instead from the HindIII –NruI sites. Legend: hiPSC = human induced pluripotent stem cell; HMW-DNA = high molecular weight DNA; -m = methylated allele; wt = wildtype; -u = unmethylated allele; yellow text/lines = hiPSC lines used in this study.
Supplementary Figure 9 Nanopore single read methylation in BAC data.
a) Methylation status of c9orf72 region in BAC data for repeats < 200 (WT), 200-750 (Cluster1,orange) and > 750 (Cluster2,red) and control (Hues64, WGBS) b) Single read methylation on a sample of 500 BAC minus strand reads sorted by repeat count (row split 200 and 750 repeats, n=423,63,14). c) Difference in mean CGI methylation of intron and promoter per read on minus strand. Reads binned by detected repeat length for BAC (n=2066 WT; 315 Cluster1; 72 Cluster2) and patient 24/5#2 (n=925 WT; 362 Cluster1; 153 Cluster2). Two sided Wilcoxon rank sum test, corrected for multiple testing (Holm), q-vals: * 0.05 - 0.01; ** 0.01 - 0.001; *** < 0.001. Median methylation differences between promoter CGI [95%CI] for WT -2.3e-5 [CI: -5.6e-6:-1.5e-5, q=7.4e-3] and Cluster1 -0.01 [CI: -7.1e-5:-3.4e-2, q=1.4e-17] and Cluster2 -0.46 [CI: -0.58:-0.37, q=1.0e-26].
Supplementary Figure 10 Region and repeat methylation detection.
a) FMR1 region methylation in SC105iPS6/iPS7 compared to Hues64 WGBS and patient sample 24/5#2. b) CGG mean repeat methylation status detected by STRique for SC105 (n=197) and synthetic plasmid control with 99 repeats treated with M.SssI+/− (5mC level on minus strand, n=1232 M.SssI+; n=11991 M.SssI-). c) GGGGCC repeat methylation status for plasmid control with 76 repeats treated with M.SssI+/− (n=2939 M.SssI+; n=31280 M.SssI-) and patient sample 24/5#2 treated with M.SssI+ (5mC level on minus strand, n=52 WT and n=6 Cluster1). Data in (b-c) presented as violin plots with overlayed boxplots (centerline, median; box limits, first and third quartiles; whiskers, 1.5× interquartile range; outliers not shown).
Supplementary Figure 11 Characterization of patient-derived hiPSC line 24/5#2.
a) IPS cell colonies from cell line 24/5#2 show the typical morphology of human pluripotent stem cells, b) stain positive for alkaline phosphatase and express the pluripotency-associated surface proteins c) Tra1-60 and d) Tra1-81. e) high-resolution SNP-karyotyping was performed to exclude major karyotypic abnormalities induced by the reprogramming process. Depicted are B-allele frequency and Log R ratio for each chromosome.
Supplementary Figure 12 Sequencing throughput per enrichment protocol and flow cell.
Number of reads on target evaluated by the STRique pipeline for all flow cells used in this study (FA*: MinION flow cell, PAD*: PromethION flow cell). a) Overhang protocol. b) Cas12 enrichment c) Cas9 enrichment.
Supplementary information
Supplementary Information
Supplementary Figs. 1–12, Supplementary Tables 1–8 and Supplementary Note 1.
Rights and permissions
About this article
Cite this article
Giesselmann, P., Brändl, B., Raimondeau, E. et al. Analysis of short tandem repeat expansions and their methylation state with nanopore sequencing. Nat Biotechnol 37, 1478–1481 (2019). https://doi.org/10.1038/s41587-019-0293-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41587-019-0293-x
This article is cited by
-
Sequencing and characterizing short tandem repeats in the human genome
Nature Reviews Genetics (2024)
-
Validated assays for the quantification of C9orf72 human pathology
Scientific Reports (2024)
-
Characterization and visualization of tandem repeats at genome scale
Nature Biotechnology (2024)
-
HycDemux: a hybrid unsupervised approach for accurate barcoded sample demultiplexing in nanopore sequencing
Genome Biology (2023)
-
A survey of algorithms for the detection of genomic structural variants from long-read sequencing data
Nature Methods (2023)