Abstract
The human reference genome represents only a small number of individuals, which limits its usefulness for genotyping. We present a method named HISAT2 (hierarchical indexing for spliced alignment of transcripts 2) that can align both DNA and RNA sequences using a graph Ferragina Manzini index. We use HISAT2 to represent and search an expanded model of the human reference genome in which over 14.5 million genomic variants in combination with haplotypes are incorporated into the data structure used for searching and alignment. We benchmark HISAT2 using simulated and real datasets to demonstrate that our strategy of representing a population of genomes, together with a fast, memory-efficient search algorithm, provides more detailed and accurate variant analyses than other methods. We apply HISAT2 for HLA typing and DNA fingerprinting; both applications form part of the HISAT-genotype software that enables analysis of haplotype-resolved genes or genomic regions. HISAT-genotype outperforms other computational methods and matches or exceeds the performance of laboratory-based assays.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The CAAPA genome data are available from dbGaP (accession phs001123.v1.p1).
Code availability
HISAT2 and HISAT-genotype are open-source software freely available at https://github.com/DaehwanKimLab/hisat2. The HISAT2 package includes programs and application programming interfaces for C++, Python and JAVA that rapidly retrieve genomic locations from repeat alignments for use in downstream analyses such as variant calling, peak calling and differential gene expression analysis.
References
1000 Genomes Project Consortium A map of human genome variation from population-scale sequencing. Nature 467, 1061–1073 (2010).
1000 Genomes Project Consortium An integrated map of genetic variation from 1,092 human genomes. Nature 491, 56–65 (2012).
GTEx Consortium The Genotype-Tissue Expression (GTEx) project. Nat. Genet. 45, 580–585 (2013).
Lappalainen, T. et al. Transcriptome and genome sequencing uncovers functional variation in humans. Nature 501, 506–511 (2013).
t Hoen, P. A. et al. Reproducibility of high-throughput mRNA and small RNA sequencing across laboratories. Nat. Biotechnol. 31, 1015–1022 (2013).
Sanders, S. J. et al. De novo mutations revealed by whole-exome sequencing are strongly associated with autism. Nature 485, 237–241 (2012).
Krumm, N. et al. Excess of rare, inherited truncating mutations in autism. Nat. Genet. 47, 582–588 (2015).
Leinonen, R., Sugawara, H., Shumway, M. & International Nucleotide Sequence Database Consortium. The sequence read archive. Nucleic Acids Res. 39, D19–D21 (2011).
Lander, E. S. et al. Initial sequencing and analysis of the human genome. Nature 409, 860–921 (2001).
Sherry, S. T. et al. dbSNP: the NCBI database of genetic variation. Nucleic Acids Res. 29, 308–311 (2001).
Lappalainen, I. et al. DbVar and DGVa: public archives for genomic structural variation. Nucleic Acids Res. 41, D936–D941 (2013).
Burrows, M. & Wheeler, D. J. A block sorting lossless data compression algorithm. SRC Research Report 124 (Digital Equipment Corporation, 1994).
Ferragina, P. & Manzini, G. in Proceedings 41st Annual Symposium on Foundations of Computer Science, IEEE Computer Society 390–398 (2000).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Li, R. et al. SOAP2: an improved ultrafast tool for short read alignment. Bioinformatics 25, 1966–1967 (2009).
Kent, W. J. BLAT—the BLAST-like alignment tool. Genome Res. 12, 656–664 (2002).
Li, H., Ruan, J. & Durbin, R. Mapping short DNA sequencing reads and calling variants using mapping quality scores. Genome Res. 18, 1851–1858 (2008).
Garrison, E. et al. Variation graph toolkit improves read mapping by representing genetic variation in the reference. Nat. Biotechnol. 36, 875–879 (2018).
Rakocevic, G. et al. Fast and accurate genomic analyses using genome graphs. Nat. Genet. 51, 354–362 (2019).
Siren, J., Valimaki, N. & Makinen, V. Indexing graphs for path queries with applications in genome research. IEEE-ACM Trans. Comput. Biol. Bioinform. 11, 375–388 (2014).
Kim, D., Langmead, B. & Salzberg, S. L. HISAT: a fast spliced aligner with low memory requirements. Nat. Methods 12, 357–360 (2015).
Robinson, J. et al. The IPD and IMGT/HLA database: allele variant databases. Nucleic Acids Res. 43, D423–D431 (2015).
Hares, D. R. Expanding the CODIS core loci in the United States. Forensic Sci. Int. Genet. 6, e52–e54 (2012).
Luo, R. et al. SOAPdenovo2: an empirically improved memory-efficient short-read de novo assembler. Gigascience 1, 18 (2012).
Compeau, P. E., Pevzner, P. A. & Tesler, G. How to apply de Bruijn graphs to genome assembly. Nat. Biotechnol. 29, 987–991 (2011).
Li, H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. Preprint at https://arxiv.org/abs/1303.3997 (2013).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie2. Nat. Methods 9, 357–359 (2012).
Eberle, M. A. et al. A reference data set of 5.4 million phased human variants validated by genetic inheritance from sequencing a three-generation 17-member pedigree. Genome Res. 27, 157–164 (2017).
Erlich, R. L. et al. Next-generation sequencing for HLA typing of class I loci. BMC Genomics 12, 42 (2011).
Lee, H. & Kingsford, C. Kourami: graph-guided assembly for novel human leukocyte antigen allele discovery. Genome Biol. 19, 16 (2018).
Li, H. Minimap and miniasm: fast mapping and de novo assembly for noisy long sequences. Bioinformatics 32, 2103–2110 (2016).
Pachter, L. Models for transcript quantification from RNA-Seq. Preprint at https://arxiv.org/abs/1104.3889 (2011).
Kim, D., Song, L., Breitwieser, F. P. & Salzberg, S. L. Centrifuge: rapid and sensitive classification of metagenomic sequences. Genome Res. 26, 1721–1729 (2016).
Acknowledgements
We would like to express our thanks to K. Barnes and M. Daya for sharing Omixon’s HLA results with us. We would like to thank B. Langmead and J. Pritt for their invaluable contributions to our discussions on HISAT2. We also greatly appreciate the generosity of G. Danuser and D. Reed in providing wet-lab bench space and equipment for us. This work was supported in part by the National Human Genome Research Institute under grants R01-HG006102 and R01-HG006677 to S.L.S. and by the Cancer Prevention Research Institute of Texas under grant RR170068 to D.K. All authors read and approved the final manuscript.
Author information
Authors and Affiliations
Contributions
D.K. and S.L.S. performed the analysis and discussed the results of HISAT2 and HISAT-genotype. D.K. designed and implemented HISAT2 and HISAT-genotype. J.M.P. optimized the index-building algorithm of HISAT2. D.K. and C.P. implemented the repeat-indexing algorithm of HISAT2. D.K., C.P. and C.B. performed the evaluations of the various programs. D.K. performed the wet-lab experiments. D.K., C.B. and S.L.S. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Figure Supplementary 1 Space efficient representation of the table in Figure 1a.
In the two ‘Node rank’ columns on the left, since node ranks are given in consecutive and increasing order, one bit (0 or 1) can be used to represent a node rank instead of 4 bytes (any number between 0 and 4,294,967,295) to manage offsets for a human genome. 1 and 0 are used to indicate a new node rank and to indicate an additional outgoing or incoming edge that a node has, respectively. To retrieve a node rank, simply summing up the 1s gives rise to that node’s rank. Since the labels in the ‘First’ column are already sorted, five numbers are enough to represent the column, two for As, three for Cs, four for Gs, three for Ts, and two for Zs. In the ‘Last’ column, two bits are used to represent each label: 00 for A, 01 for C, 10 for G, and 11 for T. 00 is also used to represent Z. HISAT2 internally resolves whether 00 represents A or Z. The right table is the space efficient representation of the left table after these transformations.
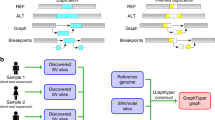
Supplementary Figure 2 Alternative approaches to graph representation.
In a and b, a reference consisting of only alleles of genes of interest can introduce significant mapping bias by mapping reads from regions not included in the restricted reference, as illustrated in more detail in Supplementary Fig. 3. In c, an aligner using the current human reference may not be able to map many reads if they originated from alleles that are substantially different from the human reference allele. In d, a reference consisting of the human reference plus numerous alleles of HLA genes enables mapping of reads from even substantially different alleles. Most HLA-typing methods, such as HLA-VBSeq, HLA*PRG, Kourami, and Graphtyper, are based on c, d, or a combination thereof to initially identify HLA reads, after which HLA-VBSeq uses approach a, and HLA*PRG, Kourami, and Graphtyper use a small-scale graph representation as described in b to perform typing. Kourami assembles only exons of HLA genes, while HISAT-genotype is able to assemble full-length sequences of HLA genes including exons and introns.
Supplementary Figure 3 Incorrect reference leading to mis-alignment and bias.
An illustration of the benefits of using the right reference/index when working with sequencing reads. The figure shows the alignment of reads to the whole genome (upper right) and to one particular genomic region denoted as Region 3 (lower right). When using the whole genome for aligning the six example reads, reads are perfectly aligned to the correct regions (regions 1, 2, 3, 4, 5, and 6). However, if the example reads are aligned using only one particular region (e.g. Region 3), five out of the six reads are incorrectly aligned to that region because alignment programs allow for a few mismatches. For example, in order to identify and extract reads that belong to the HLA-DRB1 gene from whole-genome sequencing reads, one may attempt to align them only to the HLA-DRB1 gene region. In one experiment, we found that this strategy produced 1100 times more reads mapped to HLA-DRB1 than a whole-genome alignment produced, because of the numerous pseudogene copies of DRB1 in the genome.
Supplementary Figure 5 GFM index.
The size of each block is 128 bytes, consisting of 32 4-byte cells. Each block stores: (1) four 4-byte numbers for the accumulated numbers of occurrences of A, C, G, and T up to that block, (2) one 4-byte number for the accumulated number of 1s up to the block in the Node rank (Outgoing edges) of the right table in Fig. 1a, (3) one 4-byte number for the row number of the Node rank (incoming edge) corresponding to the accumulated number of 1s indicated in (2), (4) 208 labels (or nucleotides) corresponding to the Last column of the right table in Fig. 1a, and (5) 208 ‘OUT’ bits and 208 ‘IN’ bits corresponding to the Node rank columns of the table.
Supplementary Figure 6 Alignment of a small 3-bp query using a GFM index.
How to align a 3-bp query, TAG, whose TG corresponds to the last two nucleotides of the original reference sequence, GAGCTG, and where A is a 1-bp insertion in the query. Searching from the right end of the query to the left, the nodes labeled ‘G’ are first selected (node IDs ‘4’, ‘5’, ‘6’, and ‘7’). Then the incoming edges of those nodes are examined to identify which has a preceding base ‘A’. Nodes ‘5’ and ‘7’ qualify, with preceding nodes ‘1’ and ‘2’. These in turn are examined to determine which of these nodes is preceded by a base ‘T’. Only one of the two nodes, node ‘2’, has a preceding node, ‘8’, whose label corresponds to ‘T’. Node ‘8’ is chosen as a mapped location for the query. This is the final alignment of the query shown in the prefix-sorted graph, and additional algorithms convert it to the corresponding alignment in the original graph.
Supplementary Figure 7 Construction of repeat sequences specifically for alignment purposes.
Given a de Bruijn graph in which k = 3 and each k-mer is present at least C times (e.g. five times) in the genome, a k-mer is chosen (e.g. the most frequently occurring k-mer) and extended in the left and right directions. Note that a de Bruijn graph can be easily constructed from a k-mer table. The extended sequence consisting of k-mers is called a repeat sequence. If there is a branch during extension, one of the k-mers is chosen (e.g. the most frequently occurring k-mer at the branch). In a, for example, a k-mer, TTT, is chosen and shown in yellow, and then extended in both directions until there is no extension possible, resulting in a repeat sequence, CCGTTTAC. In order to find the next repeat sequence, the k-mers belonging to the previously identified repeat sequence are removed as shown in b. A k-mer, CTT, is chosen and not extended in b as it has no k-mers to extend. In c, TAT is initially chosen and extended into TATTGT in orange. Finally, in d, TGC is chosen and not extended. Each repeat sequence consists of sub-sequences that exist in the reference genome, and a sub-sequence consists of one or more consecutive k-mers. We only store one location per sub-sequence, instead of per k-mer.
Supplementary information
Supplementary Information
Supplementary Figs. 1–7, Supplementary Tables 1–9 and Supplementary Note 1
Supplementary Dataset 1
HLA-A gene assembly of PG genome NA12878
Supplementary Dataset 2
HISAT-genotype’s HLA typing results for 17 PG genomes on HLA-A, HLA-B, HLA-C, HLA-DQA1, HLA-DQB1 and HLA-DRB1
Supplementary Dataset 3
Description: HLA-A gene assembly of PG genome NA12892
Supplementary Dataset 4
HISAT-genotype’s HLA typing results for 917 CAAPA genomes on HLA-A, HLA-B, HLA-C, HLA-DQA1, HLA-DQB1 and HLA-DRB1
Supplementary Dataset 5
Comparisons of HISAT-genotype and Kourami for HLA typing using simulated reads (see Supplementary Note 1 for description)
Supplementary Dataset 6
HISAT-genotype initial DNA fingerprinting results for 17 PG genomes
Supplementary Dataset 7
PowerPlex Fusion results for 17 PG genomes (raw signal image data)
Supplementary Dataset 8
List of alleles for 13 DNA fingerprinting loci and the amelogenin locus from the NIST short tandem repeat database
Supplementary Dataset 9
List of eight additionally incorporated alleles for four DNA fingerprinting loci D8S1179, D13S317, VWA and D21S11
Supplementary Dataset 10
HLA-A gene assembly of CAAPA genome LP6005093-DNA_E03
Rights and permissions
About this article
Cite this article
Kim, D., Paggi, J.M., Park, C. et al. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat Biotechnol 37, 907–915 (2019). https://doi.org/10.1038/s41587-019-0201-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41587-019-0201-4
This article is cited by
-
Asymmetric and parallel subgenome selection co-shape common carp domestication
BMC Biology (2024)
-
Exploration and bioinformatic prediction for profile of mRNA bound to circular RNA BTBD7_hsa_circ_0000563 in coronary artery disease
BMC Cardiovascular Disorders (2024)
-
Measuring, visualizing, and diagnosing reference bias with biastools
Genome Biology (2024)
-
Karyotype and LTR-RTs analysis provide insights into oak genomic evolution
BMC Genomics (2024)
-
Pangenome characterization and analysis of the NAC gene family reveals genes for Sclerotinia sclerotiorum resistance in sunflower (Helianthus annuus)
BMC Genomic Data (2024)