Abstract
Heterosis, or hybrid vigor, is exploited by breeders to produce elite high-yielding crop lines, but beneficial phenotypes are lost in subsequent generations owing to genetic segregation. Clonal propagation through seeds would enable self-propagation of F1 hybrids. Here we report a strategy to enable clonal reproduction of F1 rice hybrids through seeds. We fixed the heterozygosity of F1 hybrid rice by multiplex CRISPR–Cas9 genome editing of the REC8, PAIR1 and OSD1 meiotic genes to produce clonal diploid gametes and tetraploid seeds. Next, we demonstrated that editing the MATRILINEAL (MTL) gene (involved in fertilization) could induce formation of haploid seeds in hybrid rice. Finally, we combined fixation of heterozygosity and haploid induction by simultaneous editing of all four genes (REC8, PAIR1, OSD1 and MTL) in hybrid rice and obtained plants that could propagate clonally through seeds. Application of our method may enable self-propagation of a broad range of elite F1 hybrid crops.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
All data supporting the findings of this study are available in the article and its supplementary figures and tables or are available from the corresponding author on request. For sequence data, rice LOC_Os identifiers (http://rice.plantbiology.msu.edu/): LOC_Os02g37850 (OSD1), LOC_Os03g01590 (PAIR1), LOC_Os05g50410 (REC8) and LOC_Os03g27610 (MTL). Whole-genome sequencing data are deposited in the NCBI Sequence Read Archive with accession codes SRP149641 and SRP149677.
References
Schnable, P. S. & Springer, N. M. Progress toward understanding heterosis in crop plants. Annu. Rev. Plant Biol. 64, 71–88 (2013).
Birchler, J. A., Auger, D. L. & Riddle, N. C. In search of the molecular basis of heterosis. Plant Cell 15, 2236–2239 (2003).
Spillane, C., Curtis, M. D. & Grossniklaus, U. Apomixis technology development-virgin births in farmers’ fields? Nat. Biotechnol. 22, 687–691 (2004).
Sailer, C., Schmid, B. & Grossniklaus, U. Apomixis allows the transgenerational fixation of phenotypes in hybrid plants. Curr. Biol. 26, 331–337 (2016).
d’Erfurth, I. et al. Turning meiosis into mitosis. PLoS Biol. 7, e1000124 (2009).
Mieulet, D. et al. Turning rice meiosis into mitosis. Cell Res. 26, 1242–1254 (2016).
Marimuthu, M. P. A. et al. Synthetic clonal reproduction through seeds. Science 331, 876 (2011).
Karimi-Ashtiyani, R. et al. Point mutation impairs centromeric CENH3 loading and induces haploid plants. Proc. Natl Acad. Sci. USA 112, 11211–11216 (2015).
Wang, C., Shen, L., Fu, Y., Yan, C. & Wang, K. A simple CRISPR/Cas9 system for multiplex genome editing in rice. J. Genet. Genomics 42, 703–706 (2015).
Kelliher, T. et al. Matrilineal, a sperm-specific phospholipase, triggers maize haploid induction. Nature 542, 105–109 (2017).
Li, X. et al. Single nucleus sequencing reveals spermatid chromosome fragmentation as a possible cause of maize haploid induction. Nat. Commun. 8, 991 (2017).
Gilles, L. M. et al. Loss of pollen-specific phospholipase NOT LIKE DAD triggers gynogenesis in maize. EMBO J. 36, 707–717 (2017).
Liu, C. et al. A 4-bp insertion at ZmPLA1 encoding a putative phospholipase a generates haploid induction in maize. Mol. Plant 10, 520–522 (2017).
Yao, L. et al. OsMATL mutation induces haploid seed formation in indica rice. Nat. Plants 4, 530–533 (2018).
Hu, X., Meng, X., Liu, Q., Li, J. & Wang, K. Increasing the efficiency of CRISPR-Cas9-VQR precise genome editing in rice. Plant. Biotechnol. J. 16, 292–297 (2018).
Ma, X., Chen, L., Zhu, Q., Chen, Y. & Liu, Y. G. Rapid decoding of sequence-specific nuclease-induced heterozygous and biallelic mutations by direct sequencing of PCR products. Mol. Plant 8, 1285–1287 (2015).
Zhang, W. et al. The transcribed 165-bp CentO satellite is the major functional centromeric element in the wild rice species Oryza punctata. Plant Physiol. 139, 306–315 (2005).
Patel, R. K. & Jain, M. NGS QC Toolkit: a toolkit for quality control of next generation sequencing data. PLoS ONE 7, e30619 (2012).
Li, R. et al. SOAP2: an improved ultrafast tool for short read alignment. Bioinformatics 25, 1966–1967 (2009).
Li, R. et al. SNP detection for massively parallel whole-genome resequencing. Genome Res. 19, 1124–1132 (2009).
Acknowledgements
This research was supported by the Agricultural Science and Technology Innovation Program of the Chinese Academy of Agricultural Sciences (to K.W.), and the National Natural Science Foundation of China (No. 31871703 to C.W.).
Author information
Authors and Affiliations
Contributions
C.W. and K.W. conceived and designed the study. C.W., Y.S., Y.H. and Z.C. performed the lab experiments. Q.L. and T.S. conducted the computational analyses. Y.H. and J.W. carried out the field experiments. J.L. and M.W. provided the rice varieties and helped with the field management. C.W., R.M. and K.W. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors have submitted a patent application (application no. 201810325528.4 and 201811205889.1) based on the results reported in this paper.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 The morphology of inter-subspecific hybrid rice variety ‘Chunyou84’ (CY84) and its parents.
The female parent is ‘Chunjiang 16A’ (abbreviated to 16A), a late japonica male sterile line, and the male parent is ‘C84’, an indica-japonica intermediate type restorer line with wide compatibility. The maintainer line ‘Chunjiang 16B’ (16B) is used to cross with the near-isogenic 16A to produce male sterile 16A seeds. Scale bars, 5 cm.
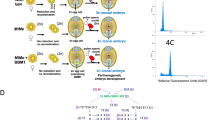
Supplementary Figure 2 Genome editing results of MiMe.
(a) Genotypes of the T0 transgenic plants of MiMe. The letters A–C represent genes OSD1, PAIR1 and REC8, respectively. Upper case letters indicate no mutation, while lower case letters indicate frameshift mutation; lower case* indicates in-frame mutation. (b) Sequencing assay of mutations around the targets of OSD1, PAIR1 and REC8 of MiMe-#7, MiMe-#8 and MiMe-#21. The protospacer adjacent motif and target sequence of wild type (WT) are denoted in red and blue, respectively. The symbol - indicates deleted nucleotides.
Supplementary Figure 3 The morphology of wild type CY84, MiMe and mtl.
The MiMe and mtl mutants showed normal vegetative growth. Scale bars, 5 cm.
Supplementary Figure 4 Chromosome spreads of male meiosis in pollen mother cells of wild type CY84, MiMe and Fix.
(a–f) CY84 (n = 32). (a) Metaphase I with 12 aligned bivalents. (b) Anaphase I. (c) Telophase I. (d) Metaphase II. (e) Anaphase II. (f) Telophase II. (g–i) MiMe (n = 45). (g) Metaphase I with 24 aligned univalents. (h) Anaphase I with segregation of 24 pairs of chromatids. (i) Telophase I. (j–l) Fix (n = 52). (j) Metaphase I with 24 aligned univalents. (k) Anaphase I with segregation of 24 pairs of chromatids. (l) Telophase I. Scale bars, 5 μm.
Supplementary Figure 5 Genome editing results of mtl.
(a) Genotypes of the T0 transgenic plant of mtl. D indicates no mutation, d indicates frameshift mutations, and d* indicates in-frame mutations in MTL. (b) Sequencing assay of mutations around the targets of MTL of mtl-#1, mtl-#2 and mtl-#3. The protospacer adjacent motif and target sequence of the wild type (WT) are denoted in red and blue, respectively. The symbol - indicates deleted nucleotides.
Supplementary Figure 6 Genome editing results of Fix.
(a) Genotypes of the T0 transgenic plants of Fix. The letters A–D represent genes OSD1, PAIR1, REC8 and MTL, respectively. Upper case letters indicate no mutation, while lower case letters indicate frameshift mutations; lower case* indicates in-frame mutations. (b) Sequencing assay of mutations around the targets of OSD1, PAIR1, REC8 and MTL of Fix-#6, Fix-#10 and Fix-#22. The protospacer adjacent motif and target sequence of wild type (WT) are denoted in red and blue, respectively. The symbol - indicates deleted nucleotides.
Supplementary information
Supplementary Information
Supplementary Figures 1–6 and Supplementary Tables 1 and 2.
Supplementary Data 1
Flow cytometry determination of ploidy in leaf cell nuclei
Source Data
Source Data Figure 2
Display of full-length gel images of cropped gel images which appear in main text
Rights and permissions
About this article
Cite this article
Wang, C., Liu, Q., Shen, Y. et al. Clonal seeds from hybrid rice by simultaneous genome engineering of meiosis and fertilization genes. Nat Biotechnol 37, 283–286 (2019). https://doi.org/10.1038/s41587-018-0003-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41587-018-0003-0
This article is cited by
-
CRISPR-Cas9 based molecular breeding in crop plants: a review
Molecular Biology Reports (2024)
-
One-line hybrid rice with high-efficiency synthetic apomixis and near-normal fertility
Plant Cell Reports (2024)
-
Potential gene editing targets for developing haploid inducer stocks in rice and wheat with high haploid induction frequency
3 Biotech (2024)
-
Prediction of protein–protein interactions between anti-CRISPR and CRISPR-Cas using machine learning technique
Journal of Plant Biochemistry and Biotechnology (2023)
-
The Female Gametophyte Characteristics and Gene Expression Analysis Involved in Apomixis of Wild Germplasm Materials of Kentucky Bluegrass in Gansu Province of China
Journal of Plant Growth Regulation (2023)