Abstract
Cell types with specialized functions fundamentally regulate animal behaviour, and yet the genetic mechanisms that underlie the emergence of novel cell types and their consequences for behaviour are not well understood1. Here we show that the monogamous oldfield mouse (Peromyscus polionotus) has recently evolved a novel cell type in the adrenal gland that expresses the enzyme AKR1C18, which converts progesterone into 20α-hydroxyprogesterone. We then demonstrate that 20α-hydroxyprogesterone is more abundant in oldfield mice, where it induces monogamous-typical parental behaviours, than in the closely related promiscuous deer mice (Peromyscus maniculatus). Using quantitative trait locus mapping in a cross between these species, we ultimately find interspecific genetic variation that drives expression of the nuclear protein GADD45A and the glycoprotein tenascin N, which contribute to the emergence and function of this cell type in oldfield mice. Our results provide an example by which the recent evolution of a new cell type in a gland outside the brain contributes to the evolution of social behaviour.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Sequence data are available at NCBI Sequence Read Archive under BioProject ID PRJNA1094591. Source data are provided with this paper.
References
Arendt, D. et al. The origin and evolution of cell types. Nat. Rev. Genet. 17, 744–757 (2016).
Tosches, M. A. et al. Evolution of pallium, hippocampus, and cortical cell types revealed by single-cell transcriptomics in reptiles. Science 360, 881–888 (2018).
Bakken, T. E. et al. Single-cell and single-nucleus RNA-seq uncovers shared and distinct axes of variation in dorsal LGN neurons in mice, non-human primates, and humans. eLife 10, e64875 (2021).
Krienen, F. M. et al. Innovations present in the primate interneuron repertoire. Nature 586, 262–269 (2020).
Hain, D. et al. Molecular diversity and evolution of neuron types in the amniote brain. Science 377, eabp8202 (2022).
Woych, J. et al. Cell-type profiling in salamanders identifies innovations in vertebrate forebrain evolution. Science 377, eabp9186 (2022).
Bakken, T. E. et al. Comparative cellular analysis of motor cortex in human, marmoset and mouse. Nature 598, 111–119 (2021).
Callaway, E. M. et al. A multimodal cell census and atlas of the mammalian primary motor cortex. Nature 598, 86–102 (2021).
Knoedler, J. R. et al. A functional cellular framework for sex and estrous cycle-dependent gene expression and behavior. Cell 185, 654–671.e22 (2022).
Kim, D.-W. et al. Multimodal analysis of cell types in a hypothalamic node controlling social behavior. Cell 179, 713–728.e17 (2019).
Moffitt, J. R. et al. Molecular, spatial, and functional single-cell profiling of the hypothalamic preoptic region. Science 362, eaau5324 (2018).
Brückner, A. et al. Evolutionary assembly of cooperating cell types in an animal chemical defense system. Cell 184, 6138–6156.e28 (2021).
Khadraoui, M., Merritt, J. R., Hoekstra, H. E. & Bendesky, A. Post-mating parental behavior trajectories differ across four species of deer mice. PLoS ONE 17, e0276052 (2022).
Melmed, S., Koenig, R., Rosen, C., Auchus, R. & Goldfine, A. Williams Textbook of Endocrinology 14th edn (Elsevier Health Sciences, 2019).
Barresi, M. J. F. & Gilbert, S. F. Developmental Biology 13th edn (Oxford Univ. Press, 2023).
Keeney, D. S., Jenkins, C. M. & Waterman, M. R. Developmentally regulated expression of adrenal 17 α-hydroxylase cytochrome P450 in the mouse embryo. Endocrinology 136, 4872–4879 (1995).
Ogunsua, A. O., de Nicola, A. F., Traikov, H., Birmingham, M. K. & Levine, S. Adrenal steroid biosynthesis by different species of mouselike rodents. Gen. Comp. Endocrinol. 16, 192–199 (1971).
Mao, J., Duan, R. W., Zhong, L., Gibori, G. & Azhar, S. Expression, purification and characterization of the rat luteal 20 α-hydroxysteroid dehydrogenase. Endocrinology 138, 182–190 (1997).
Veliça, P. et al. Lack of functional and expression homology between human and mouse aldo-keto reductase 1C enzymes: implications for modelling human cancers. Mol. Cancer 8, 121 (2009).
Wooldridge, T. B. et al. An enhancer of Agouti contributes to parallel evolution of cryptically colored beach mice. Proc. Natl Acad. Sci. USA 119, e2202862119 (2022).
Kubli-Garfias, C. & Whalen, R. E. Induction of lordosis behavior in female rats by intravenous administration of progestins. Horm. Behav. 9, 380–386 (1977).
Bendesky, A. et al. The genetic basis of parental care evolution in monogamous mice. Nature 544, 434–439 (2017).
Williams, J. R., Catania, K. C. & Carter, C. S. Development of partner preferences in female prairie voles (Microtus ochrogaster): the role of social and sexual experience. Horm. Behav. 26, 339–349 (1992).
Ogle, T. F. & Beyer, B. K. Steroid-binding specificity of the progesterone receptor from rat placenta. J. Steroid Biochem. 16, 147–150 (1982).
Young, P. C. & Cleary, R. E. Characterization and properties of progesterone-binding components in human endometrium. J. Clin. Endocrinol. Metab. 39, 425–439 (1974).
Nowak, F. V., Nuti, K. M. & Karavolas, H. J. Quantitative changes in the metabolism of 20α-hydroxy-4-pregnen-3-one by rat hypothalamus and pituitary during proestrus. Steroids 28, 509–520 (1976).
Nowak, F. V. Distribution and metabolism of 20α-hydroxylated progestins in the female rat. J. Steroid Biochem. Mol. Biol. 80, 469–479 (2002).
Khanna, M., Qin, K.-N. & Cheng, K.-C. Distribution of 3α-hydroxysteroid dehydrogenase in rat brain and molecular cloning of multiple cDNAs encoding structurally related proteins in humans. J. Steroid Biochem. Mol. Biol. 53, 41–46 (1995).
Penning, T. M. et al. Human 3α-hydroxysteroid dehydrogenase isoforms (AKR1C1–AKR1C4) of the aldo-keto reductase superfamily: functional plasticity and tissue distribution reveals roles in the inactivation and formation of male and female sex hormones. Biochem. J 351, 67–77 (2000).
Agís-Balboa, R. C. et al. Characterization of brain neurons that express enzymes mediating neurosteroid biosynthesis. Proc. Natl Acad. Sci. USA 103, 14602–14607 (2006).
Russell, D. W. & Wilson, J. D. Steroid 5 α-reductase: two genes/two enzymes. Annu. Rev. Biochem. 63, 25–61 (1994).
MacKenzie, G. & Maguire, J. Neurosteroids and GABAergic signaling in health and disease. Biomol. Concepts 4, 29–42 (2013).
Rudolph, S. et al. Cerebellum-specific deletion of the GABAA receptor δ subunit leads to sex-specific disruption of behavior. Cell Rep. 33, 108338 (2020).
Kohl, J. & Dulac, C. Neural control of parental behaviors. Curr. Opin. Neurobiol. 49, 116–122 (2018).
Stell, B. M., Brickley, S. G., Tang, C. Y., Farrant, M. & Mody, I. Neuroactive steroids reduce neuronal excitability by selectively enhancing tonic inhibition mediated by δ subunit-containing GABAA receptors. Proc. Natl Acad. Sci. USA 100, 14439–14444 (2003).
Maguire, J. & Mody, I. GABAAR plasticity during pregnancy: relevance to postpartum depression. Neuron 59, 207–213 (2008).
Maguire, J., Ferando, I., Simonsen, C. & Mody, I. Excitability changes related to GABAA receptor plasticity during pregnancy. J. Neurosci. 29, 9592–9601 (2009).
Priestley, C. M., Williamson, E. M., Wafford, K. A. & Sattelle, D. B. Thymol, a constituent of thyme essential oil, is a positive allosteric modulator of human GABAA receptors and a homo-oligomeric GABA receptor from Drosophila melanogaster. Br. J. Pharmacol. 140, 1363–1372 (2003).
Belelli, D. & Gee, K. W. 5α-pregnan-3α,20α-diol behaves like a partial agonist in the modulation of GABA-stimulated chlride ion uptake by synaptoneurosomes. Eur. J. Pharmacol. 167, 173–176 (1989).
Farrant, M. & Nusser, Z. Variations on an inhibitory theme: phasic and tonic activation of GABAA receptors. Nat. Rev. Neurosci. 6, 215–229 (2005).
Bixo, M. et al. Treatment of premenstrual dysphoric disorder with the GABAA receptor modulating steroid antagonist Sepranolone (UC1010)-A randomized controlled trial. Psychoneuroendocrinology 80, 46–55 (2017).
Legesse, D. H. et al. Structural insights into opposing actions of neurosteroids on GABAA receptors. Nat. Commun. 14, 5091 (2023).
Dolfi, B. et al. Unravelling the sex-specific diversity and functions of adrenal gland macrophages. Cell Rep. 39, 110949 (2022).
Hanemaaijer, E. S. et al. Single-cell atlas of developing murine adrenal gland reveals relation of Schwann cell precursor signature to neuroblastoma phenotype. Proc. Natl Acad. Sci. USA 118, e2022350118 (2021).
Huang, L. et al. Single-cell transcriptomes reveal characteristic features of cell types within the human adrenal microenvironment. J. Cell. Physiol. 236, 7308–7321 (2021).
Lai, S. et al. Mapping a mammalian adult adrenal gland hierarchy across species by microwell-seq. Cell Regen. 9, 11 (2020).
Mitani, F. Functional zonation of the rat adrenal cortex: the development and maintenance. Proc. Jpn. Acad. Ser. B 90, 163–183 (2014).
Barreto, G. et al. Gadd45a promotes epigenetic gene activation by repair-mediated DNA demethylation. Nature 445, 671–675 (2007).
Wingert, S. et al. DNA-damage response gene GADD45A induces differentiation in hematopoietic stem cells without inhibiting cell cycle or survival. Stem Cells 34, 699–710 (2016).
Zhang, R., Shao, J. & Xiang, L. GADD45A protein plays an essential role in active DNA demethylation during terminal osteogenic differentiation of adipose-derived mesenchymal stem cells. J. Biol. Chem. 286, 41083–41094 (2011).
Tucker, R. P. & Degen, M. The expression and possible functions of tenascin-W during development and disease. Front. Cell Dev. Biol. 7, 1–10 (2019).
Merritt, J. R. et al. A supergene-linked estrogen receptor drives alternative phenotypes in a polymorphic songbird. Proc. Natl Acad. Sci. USA 117, 21673–21680 (2020).
Florensa, E., Harrison, R., Johnson, M. & Youssefnejadian, E. Plasma 20α-dihydroprogesterone, progesterone and 17-hydroxyprogesterone in normal human pregnancy. Acta Endocrinol. 86, 634–640 (1977).
Abdel-Khalik, J., Björklund, E. & Hansen, M. Simultaneous determination of endogenous steroid hormones in human and animal plasma and serum by liquid or gas chromatography coupled to tandem mass spectrometry. J. Chromatogr. B 928, 58–77 (2013).
Jensen, C. C. Quantitative determination of urinary pregnanediol and allopregnanediol for clinical use. Eur. J. Endocrinol. 18, 281–287 (1955).
Patterson, R., Balan, I., Morrow, A. L. & Meltzer-Brody, S. Novel neurosteroid therapeutics for post-partum depression: perspectives on clinical trials, program development, active research, and future directions. Neuropsychopharmacology 49, 67–72 (2023).
Hobert, O. & Kratsios, P. Neuronal identity control by terminal selectors in worms, flies, and chordates. Curr. Opin. Neurobiol. 56, 97–105 (2019).
Lynch, V. J. et al. Adaptive changes in the transcription factor HoxA-11 are essential for the evolution of pregnancy in mammals. Proc. Natl Acad. Sci. USA 105, 14928–14933 (2008).
Lynch, V. J., May, G. & Wagner, G. P. Regulatory evolution through divergence of a phosphoswitch in the transcription factor CEBPB. Nature 480, 383–386 (2011).
Chiquet-Ehrismann, R., Orend, G., Chiquet, M., Tucker, R. P. & Midwood, K. S. Tenascins in stem cell niches. Matrix Biol. 37, 112–123 (2014).
Pesheva, P., Gloor, S., Schachner, M. & Probstmeier, R. Tenascin-R is an intrinsic autocrine factor for oligodendrocyte differentiation and promotes cell adhesion by a sulfatide-mediated mechanism. J. Neurosci. 17, 4642–4651 (1997).
Kimura, H., Akiyama, H., Nakamura, T. & Crombrugghe, B. Tenascin-W inhibits proliferation and differentiation of preosteoblasts during endochondral bone formation. Biochem. Biophys. Res. Commun. 356, 935–941 (2007).
Czopka, T., Von Holst, A., Schmidt, G., Ffrench-Constant, C. & Faissner, A. Tenascin C and tenascin R similarly prevent the formation of myelin membranes in a RhoA-dependent manner, but antagonistically regulate the expression of myelin basic protein via a separate pathway. Glia 57, 1790–1801 (2009).
Uhlén, M. et al. Tissue-based map of the human proteome. Science 347, 1260419 (2015).
Choi, H. M. T. et al. Third-generation in situ hybridization chain reaction: multiplexed, quantitative, sensitive, versatile, robust. Development 145, dev165753 (2018).
Kuehn, E. et al. Segment number threshold determines juvenile onset of germline cluster expansion in Platynereis dumerilii. J. Exp. Zoolog. B 338, 225–240 (2022).
Renier, N. et al. iDISCO: A simple, rapid method to immunolabel large tissue samples for volume imaging. Cell 159, 896–910 (2014).
Bedford, N. L. et al. Automated tracking reveals the social network of beach mice and their burrows. Preprint at bioRxiv https://doi.org/10.1101/2021.08.07.455531 (2021).
Kingsley, E. P., Kozak, K. M., Pfeifer, S. P., Yang, D.-S. & Hoekstra, H. E. The ultimate and proximate mechanisms driving the evolution of long tails in forest deer mice. Evolution 71, 261–273 (2017).
Pallares, L. F., Picard, S. & Ayroles, J. F. TM3′seq: a tagmentation-mediated 3′ sequencing approach for improving scalability of RNA-seq experiments. G3 10, 143–150 (2020).
Picelli, S. et al. Tn5 transposase and tagmentation procedures for massively scaled sequencing projects. Genome Res. 24, 2033–2040 (2014).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Li, B. & Dewey, C. N. RSEM: accurate transcript quantification from RNA-seq data with or without a reference genome. BMC Bioinformatics 12, 323 (2011).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Bray, N. L., Pimentel, H., Melsted, P. & Pachter, L. Near-optimal probabilistic RNA-seq quantification. Nat. Biotechnol. 34, 525–527 (2016).
Turro, E. et al. Haplotype and isoform specific expression estimation using multi-mapping RNA-seq reads. Genome Biol. 12, R13 (2011).
Detlefsen, A. J., Wangtrakuldee, P. & Penning, T. M. Characterization of the major single nucleotide polymorphic variants of aldo-keto reductase 1C3 (type 5 17β-hydroxysteroid dehydrogenase). J. Steroid Biochem. Mol. Biol. 221, 106121 (2022).
Friard, O. & Gamba, M. BORIS: a free, versatile open-source event-logging software for video/audio coding and live observations. Methods Ecol. Evol. 7, 1325–1330 (2016).
Bright, D. P. & Smart, T. G. Methods for recording and measuring tonic GABAA receptor-mediated inhibition. Front. Neural Circuits 7, 193 (2013).
Fleming, S. J. et al. Unsupervised removal of systematic background noise from droplet-based single-cell experiments using CellBender. Nat. Methods 20, 1323–1335 (2023).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587.e29 (2021).
Huang, Y., McCarthy, D. J. & Stegle, O. Vireo: Bayesian demultiplexing of pooled single-cell RNA-seq data without genotype reference. Genome Biol. 20, 273 (2019).
Hao, Y. et al. Dictionary learning for integrative, multimodal and scalable single-cell analysis. Nat. Biotechnol. 42, 293–304 (2023).
Mi, H. et al. Protocol update for large-scale genome and gene function analysis with the PANTHER classification system (v.14.0). Nat. Protoc. 14, 703–721 (2019).
Kirilenko, B. M. et al. Integrating gene annotation with orthology inference at scale. Science 380, eabn3107 (2023).
Corbett-Detig, R. & Nielsen, R. A hidden Markov model approach for simultaneously estimating local ancestry and admixture time using next generation sequence data in samples of arbitrary ploidy. PLoS Genet. 13, e1006529 (2017).
Broman, K. W., Wu, H., Sen, Ś. & Churchill, G. A. R/qtl: QTL mapping in experimental crosses. Bioinformatics 19, 889–890 (2003).
Wickham, H. Ggplot2: Elegant Graphics for Data Analysis (Springer-Verlag, 2016).
Acknowledgements
This work was supported by the following grants: Searle Scholarship, Klingenstein-Simons Fellowship in Neuroscience, Sloan Foundation Fellowship, and National Institutes of Health (NIH) R00HD084732, R21HD106241 and R35GM143051 to A.B.; the Simons Society of Fellows Junior Fellowship 855220 to J.R.M.; Canadian Institutes of Health Research Project Grant 426405 to K.K.S.; Tishman Fellowship to S.L.; and SFARI Bridge to Independence award and Rose F. Kennedy Intellectual and Developmental Disabilities Research Center pilot grant to S.R. Imaging was performed with support from the Zuckerman Institute’s Cellular Imaging platform, which received grant S10OD023587-01 from the NIH. Columbia University’s Shared Research Computing Facility, where computations were performed, was supported by NIH grant G20RR030893-01 and NYSTAR contract C090171. L. Hammond and H. Ibarra provided training and advice on microscope imaging, C. Zhang shared reagents, L. Remedio shared reagents and advice on tissue clearing, H. Pan and M. Hiller helped run TOGA to find orthologues, and H. Hoesktra provided beach mice and cloudland mice adrenals. M. Tosches, R. Axel, S. Siegelbaum and D. Bambah-Mukku and his students provided comments on the manuscript. C. Everett created the diagram in Fig. 4f. Schematics in Fig. 2a,i and Extended Data Fig. 3 were created using BioRender.com.
Author information
Authors and Affiliations
Contributions
N.N. and A.B. conceived and designed the study. N.N. conducted all transcriptomic and genetic experiments and analyses, with contribution from M.U. and J.R.M. J.R.M. performed behaviour assays with contributions from V.S.E., I.B.B., K.H., M.U. and N.N. N.N. and M.U. performed adrenal histology. N.N. and E.L. characterized adrenal mass and volume. K.H. developed the pair-bonding assay. C.G. and J.R.M. built behaviour testing apparatus. S.A.W. performed AKR1C18 biochemistry experiments. K.K.S. and A.P. developed LC–MS/MS steroid quantification methods and quantified steroid levels. S.R. and S.L. conducted electrophysiology experiments and analysis. N.N., J.R.M. and A.B. wrote the paper with input from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks Tali Kimchi and Jessica Tollkuhn for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Anatomical, molecular, and biochemical characterization of deer- and oldfield adrenal glands.
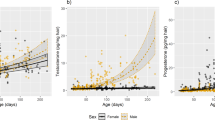
a, Adrenal weight at postnatal day 0.5. Lines at median. b, Adult adrenal weight by age. P-values by generalized linear model. Band is 95% confidence interval. c, Adrenal medulla volume. Lines at median. P-values by generalized linear model. d, Representative adrenal section from the zona fasciculata (zF) of the deer- and oldfield mouse adrenal cortex. Cyp11b1 (green) labeled by in situ hybridization and counterstained with DAPI (red). Five independent biological replicates per species yielded similar results. Scale bar, 20 µm. e, Scatterplot of gene expression by species and sex. f, Boxplots of the expression of steroidogenic enzymes in the corticosterone- and 20α-OHP synthesis pathways in the adrenal. Boxplot hinges are 25th and 75th quartiles, whiskers are 1.5× interquartile range, line at median. g, Total steroid levels in the adrenal gland of deer and oldfield mice. Lines at median. P-values by generalized linear model. DOC, deoxycorticosterone. h, Expression of Akr1c18 in the ovaries and testes of oldfield and deer mice. Lines at median. P-values by two-sided t-test. i, Akr1c18 (red) labeled by in situ hybridization in the ovary of a deer mouse and counterstained with DAPI (blue). Three independent biological replicates per species yielded similar results. Scale bar, 0.5 mm. j, Circulating 20α-OHP levels in the blood of oldfield mice after adrenalectomy. Lines at median. P-values by generalized linear model.
Extended Data Fig. 2 Effects of 20α-OHP on alloparental and parental care.
Male and female care for pups as measured by proportion of time spent huddling pups, proportion of time spent grooming pups, fraction of pups retrieved to the nest, and nest quality score (from 0 to 4) in a, unmated deer mice, b, unmated oldfield mice, and c, deer mouse fathers. Boxplot hinges are 25th and 75th quartiles, whiskers are 1.5× interquartile range, line at median. P-values by generalized linear model for proportion of time huddling, proportion of time grooming the pup, proportion of pups retrieved, and nest quality. P-values for proportion of animals that retrieved at least one pup by Fisher’s exact test. d, Proportion of pup attacks by species, treatment, and reproductive experience.
Extended Data Fig. 3 Partner preference is not affected by 20α-OHP.
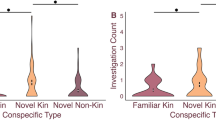
Top: Schematic of the experimental design of the partner preference test. Bottom: Observations from the same breeding pairs are connected by lines (10 min partner preference test). P-values by generalized linear model (effect of trial type).
Extended Data Fig. 4 20α-OHP is converted to allo-diol in the brain of deer- and oldfield mice.
a, Concentration of allo-diol, allopregnanolone and progesterone in deer- and oldfield mouse cerebellum and hypothalamus after incubation with 20α-OHP, as measured by LC-MS/MS. P-values by two-sided t-test.
Extended Data Fig. 5 Effects of allo-diol on alloparental and parental care, and on δGABAAR.
Male and female care for pups as measured by proportion of time spent huddling pups, proportion of time spent grooming pups, fraction of pups retrieved to the nest, and nest quality score in a, unmated deer mice, b, unmated oldfield mice, and c, oldfield parents. Boxplot hinges are 25th and 75th quartiles, whiskers are 1.5× interquartile range, line at median. P-values by generalized linear model for proportion of time huddling, proportion of time grooming the pup, proportion of pups retrieved, and nest quality. P-values for proportion of animals that retrieved at least one pup by Fisher’s exact test. d, Proportion of pup attacks by species, treatment, and reproductive experience. e, Baseline tonic GABA receptor currents with leak current subtracted, after gabazine application. f, Input resistance (Ri) under different pharmacological conditions. No change in Ri in the presence of allo-diol and 20α-OHP suggests no effect on GABA receptor currents. A decrease in Ri in the presence of THIP is consistent with an increase in GABA receptor currents. This effect is diminished when THIP is co-applied with allo-diol but not with 20α-OHP. Ri increases in gabazine when all GABA receptors are blocked. P-values by two-sided one-sample t-test (μ = 0) and two-sample two-sided t-tests. Bars denote the mean ± SEM.
Extended Data Fig. 6 UMAP visualization of the house-, deer-, and oldfield mouse adrenal.
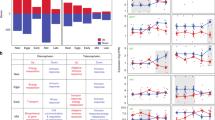
a, UMAP of integrated analysis of house mouse, deer mouse, and oldfield mouse adrenal nuclei. b, UMAP showing Akr1c18 expression in adrenal cells, marking the X zone in house mice and the zona inaudita in oldfield mice. c, UMAP of integrated analysis of deer mouse and oldfield mouse nuclei, then split by sex. d, Volcano plot of differential adrenal gene expression between deer- and oldfield mouse (purple: higher in deer mouse, green: higher in oldfield mouse, grey: n.s.). Akr1c18 and extracellular matrix gene markers of the zona inaudita (from Fig. 3g) are highlighted. False Discovery Rate=0.05. A medulla, adrenergic medulla; NA medulla, noradrenergic medulla.
Extended Data Fig. 7 Expression of top marker genes of the adrenal zones of deer mice and oldfield mice.
Violin plots denoting the top two markers of each adrenal cell type.
Extended Data Fig. 8 Dot plots of marker genes of the zona inaudita of oldfield mice and the X zone of house mice.
Expression of transcription factors (TFs), extracellular matrix (ECM) genes, and other genes upregulated in the zona inaudita or X zone. Note that in adults, the X zone is only present in unmated females.
Extended Data Fig. 9 Cis-regulation of Gadd45a contributes to transcription factor module expression.
a, Expression of Akr1c18 in the adrenal of female and male deer × oldfield F2 hybrids. b, Dot plot of Gadd45a, Tnn, and Akr1c18 expression in adrenal cortex cell types. c, Gadd45a expression by genotype at the Gadd45a locus in F2-hybrid males. Boxplot hinges are 25th and 75th quartiles, whiskers are 1.5× interquartile range, line at median. P-value by ANOVA. Logarithm of the odds (LOD) across the genome of d, Hif1a and e, Runx2 expression. f, LOD across the genome for the TF module and the TF module excluding Gadd45a. g, LOD of the TF module and the TF module controlling for Gadd45a expression. Dashed lines denote genome-wide threshold of significance (α = 0.05).
Extended Data Fig. 10 Cis-regulation of Tnn contributes to extracellular matrix module expression.
a, Expression of Tnn in deer and oldfield mice. P-value by two-sided t-test. b, Allele-specific expression of Tnn in deer × oldfield F1 hybrids. P-value by paired two-sided t-test. c, Correlation between Akr1c18 expression and Tnn expression across development of oldfield mice. P-value by bivariate correlation, R denotes Pearson’s correlation coefficient. d, Logarithm of the odds (LOD) across the genome of Cdkn2a, Podnl1, Serpine1, and Timp1 expression. e, LOD of the ECM module and the ECM module without Tnn. f, LOD of the ECM module and the ECM module controlling for Tnn expression. g, LOD of Akr1c18 expression and Akr1c18 expression controlling for Tnn expression. Dashed lines denote genome-wide threshold of significance (α = 0.05). h, Tnn expression as a function of genotype at the Tnn locus in F2-hybrid males. Akr1c18 expression as a function of genotype at the Tnn locus (i) or the Akr1c18 locus (j) in F2-hybrid males. h,i,j Boxplot hinges are 25th and 75th quartiles, whiskers are 1.5× interquartile range, line at median. P-values by ANOVA.
Supplementary information
Supplementary Information
Supplementary Methods, Supplementary Table 3, Supplementary Notes 1 and 2, Supplementary References and Supplementary Figs. 1–3.
Supplementary Table 1
Adrenal cell cluster markers
Supplementary Table 2
One-to-one orthologues
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Niepoth, N., Merritt, J.R., Uminski, M. et al. Evolution of a novel adrenal cell type that promotes parental care. Nature (2024). https://doi.org/10.1038/s41586-024-07423-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41586-024-07423-y
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.