Abstract
To survive bacteriophage (phage) infections, bacteria developed numerous anti-phage defence systems1,2,3,4,5,6,7. Some of them (for example, type III CRISPR–Cas, CBASS, Pycsar and Thoeris) consist of two modules: a sensor responsible for infection recognition and an effector that stops viral replication by destroying key cellular components8,9,10,11,12. In the Thoeris system, a Toll/interleukin-1 receptor (TIR)-domain protein, ThsB, acts as a sensor that synthesizes an isomer of cyclic ADP ribose, 1′′−3′ glycocyclic ADP ribose (gcADPR), which is bound in the Smf/DprA-LOG (SLOG) domain of the ThsA effector and activates the silent information regulator 2 (SIR2)-domain-mediated hydrolysis of a key cell metabolite, NAD+ (refs. 12,13,14). Although the structure of ThsA has been solved15, the ThsA activation mechanism remained incompletely understood. Here we show that 1′′−3′ gcADPR, synthesized in vitro by the dimeric ThsB′ protein, binds to the ThsA SLOG domain, thereby activating ThsA by triggering helical filament assembly of ThsA tetramers. The cryogenic electron microscopy (cryo-EM) structure of activated ThsA revealed that filament assembly stabilizes the active conformation of the ThsA SIR2 domain, enabling rapid NAD+ depletion. Furthermore, we demonstrate that filament formation enables a switch-like response of ThsA to the 1′′−3′ gcADPR signal.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All data are available in the manuscript and the Supplementary Information. The atomic coordinates and structure factors or cryo-EM density maps have been deposited in the PDB and Electron Microscopy Data Bank under accession codes 8BTN (ThsB’ crystal structure); 8BTO and EMD-16233 (ThsA–1′′−3′ gcADPR); and 8BTP and EMD-16234 (ThsA–1′′−3′ gcεADPR). The crystal structures in Fig. 1e (PDB accession code 6LHY), 2g and 3a (6LHX) and Fig. 2h (1ICI) and Extended Data Fig. 2b (7O62, 7UN8) and 6f (4IAO) are publicly available from the PDB. Source data are provided with this paper.
Code availability
No code was used for this study.
References
Doron, S. et al. Systematic discovery of antiphage defense systems in the microbial pangenome. Science 359, eaar4120 (2018).
Gao, L. et al. Diverse enzymatic activities mediate antiviral immunity in prokaryotes. Science 369, 1077–1084 (2020).
Millman, A. et al. An expanded arsenal of immune systems that protect bacteria from phages. Cell Host Microbe 30, 1556–1569 (2022).
Rousset, F. et al. Phages and their satellites encode hotspots of antiviral systems. Cell Host Microbe 30, 740–753 (2022).
Vassallo, C. N., Doering, C. R., Littlehale, M. L., Teodoro, G. I. C. & Laub, M. T. A functional selection reveals previously undetected anti-phage defence systems in the E. coli pangenome. Nat. Microbiol. 7, 1568–1579 (2022).
Pingoud, A., Wilson, G. G. & Wende, W. Type II restriction endonucleases — a historical perspective and more. Nucleic Acids Res. 44, 8011–8011 (2016).
Koonin, E. V., Makarova, K. S. & Zhang, F. Diversity, classification and evolution of CRISPR-Cas systems. Curr. Opin. Microbiol. 37, 67–78 (2017).
Kazlauskiene, M., Kostiuk, G., Venclovas, Č., Tamulaitis, G. & Siksnys, V. A cyclic oligonucleotide signaling pathway in type III CRISPR-Cas systems. Science 357, 605–609 (2017).
Niewoehner, O. et al. Type III CRISPR-Cas systems produce cyclic oligoadenylate second messengers. Nature 548, 543–548 (2017).
Lowey, B. et al. CBASS immunity uses CARF-related effectors to sense 3′–5′- and 2′–5′-linked cyclic oligonucleotide signals and protect bacteria from phage infection. Cell 182, 38–49 (2020).
Tal, N. et al. Cyclic CMP and cyclic UMP mediate bacterial immunity against phages. Cell 184, 5728–5739 (2021).
Ofir, G. et al. Antiviral activity of bacterial TIR domains via immune signalling molecules. Nature 600, 116–120 (2021).
Leavitt, A. et al. Viruses inhibit TIR gcADPR signalling to overcome bacterial defence. Nature 611, 326–331 (2022).
Manik, M. K. et al. Cyclic ADP ribose isomers: production, chemical structures, and immune signaling. Science 377, eadc8969 (2022).
Ka, D., Oh, H., Park, E., Kim, J.-H. & Bae, E. Structural and functional evidence of bacterial antiphage protection by Thoeris defense system via NAD+ degradation. Nat. Commun. 11, 2816 (2020).
Athukoralage, J. S. & White, M. F. Cyclic nucleotide signaling in phage defense and counter-defense. Annu. Rev. Virol. 9, 451–468 (2022).
Nimma, S. et al. Structural evolution of TIR-domain signalosomes. Front. Immunol. 12, 784484 (2021).
Holm, L. DALI and the persistence of protein shape. Protein Sci. 29, 128–140 (2020).
Fernández-Lucas, J. et al. Biochemical and structural studies of two tetrameric nucleoside 2′-deoxyribosyltransferases from psychrophilic and mesophilic bacteria: Insights into cold-adaptation. Int. J. Biol. Macromol. 192, 138–150 (2021).
Armstrong, S. R., Cook, W. J., Short, S. A. & Ealick, S. E. Crystal structures of nucleoside 2-deoxyribosyltransferase in native and ligand-bound forms reveal architecture of the active site. Structure 4, 97–107 (1996).
Morehouse, B. R. et al. Cryo-EM structure of an active bacterial TIR–STING filament complex. Nature 608, 803–807 (2022).
Martin, R. et al. Structure of the activated ROQ1 resistosome directly recognizing the pathogen effector XopQ. Science 370, eabd9993 (2020).
Eastman, S. et al. A phytobacterial TIR domain effector manipulates NAD+ to promote virulence. New Phytol. 233, 890–904 (2022).
Kagan, J. C., Magupalli, V. G. & Wu, H. SMOCs: supramolecular organizing centres that control innate immunity. Nat. Rev. Immunol. 14, 821–826 (2014).
Zhang, Q., Bhattacharya, S. & Andersen, M. E. Ultrasensitive response motifs: basic amplifiers in molecular signalling networks. Open Biol. 3, 130031 (2013).
Park, C. K. & Horton, N. C. Structures, functions, and mechanisms of filament forming enzymes: a renaissance of enzyme filamentation. Biophys. Rev. 11, 927–994 (2019).
Hsu, H.-C. et al. Structural basis for allosteric stimulation of Sir2 activity by Sir4 binding. Genes Dev. 27, 64–73 (2013).
Zaremba, M. et al. Short prokaryotic Argonautes provide defence against incoming mobile genetic elements through NAD+ depletion. Nat. Microbiol. 7, 1857–1869 (2022).
Koopal, B. et al. Short prokaryotic Argonaute systems trigger cell death upon detection of invading DNA. Cell 185, 1471–1486 (2022).
Garb, J. et al. Multiple phage resistance systems inhibit infection via SIR2-dependent NAD+ depletion. Nat. Microbiol. 7, 1849–1856 (2022).
Hogrel, G. et al. Cyclic nucleotide-induced helical structure activates a TIR immune effector. Nature 608, 808–812 (2022).
Clabbers, M. T. B. et al. MyD88 TIR domain higher-order assembly interactions revealed by microcrystal electron diffraction and serial femtosecond crystallography. Nat. Commun. 12, 2578 (2021).
Kabsch, W. XDS. Acta Crystallogr. D 66, 125–132 (2010).
Collaborative Computational Project, Number 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D 50, 760–763 (1994).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Mirdita, M. et al. ColabFold: making protein folding accessible to all. Nat. Methods 19, 679–682 (2022).
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004).
Liebschner, D. et al. Macromolecular structure determination using X-rays, neutrons and electrons: recent developments in Phenix. Acta Crystallogr. D 75, 861–877 (2019).
R Core Team. R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, 2020).
Mazzocco, A., Waddell, T. E., Lingohr, E. & Johnson, R. P. in Bacteriophages Vol. 501 (eds Clokie, M. R. J. & Kropinski, A. M.) 81–85 (Humana, 2009).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION−3. eLife 7, e42166 (2018).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Pettersen, E. F. et al. UCSF ChimeraX: structure visualization for researchers, educators, and developers. Protein Sci. 30, 70–82 (2021).
Krissinel, E. Crystal contacts as nature’s docking solutions. J. Comput. Chem. 31, 133–143 (2010).
Min, J., Landry, J., Sternglanz, R. & Xu, R.-M. Crystal structure of a SIR2 homolog–NAD complex. Cell 105, 269–279 (2001).
Morehouse, B. R. et al. STING cyclic dinucleotide sensing originated in bacteria. Nature 586, 429–433 (2020).
Acknowledgements
The research has been supported by the Research Council of Lithuania grant S-MIP-21-6 to G.T. The synchrotron X-ray diffraction data were collected at beamline P14 operated by the European Molecular Biology Laboratory, Hamburg at the PETRA III storage ring (Deutsches Elektronen-Synchrotron, Hamburg, Germany). Access to the beamline has been supported by iNEXT-Discovery, project number 871037, funded by the Horizon 2020 programme of the European Commission. We thank I. Drulyte and A. Carabias for discussions on cryo-EM structure determination.
Author information
Authors and Affiliations
Contributions
G.T. and M.Z. designed the research; D.S. and G.O. cloned the proteins and carried out mutagenesis; A.S. carried out the protein purifications; G.S. and G.T. prepared cryo-EM samples, collected cryo-EM data, solved the structures and carried out structure analyses; G.T. and D.S. collected X-ray diffraction data, G.T. solved X-ray structure; D.S. carried out biochemical experiments and crystallized ThsB′; A.R. carried out MS analysis; C.A. carried out phage plaque assays; G.T., D.S., G.S., M.Z., R.S. and V.S. wrote the manuscript with input from all authors. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
R.S. is a scientific cofounder of and adviser for BiomX and Ecophage. All other authors declare no competing interests.
Peer review
Peer review information
Nature thanks Laura Spagnolo and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Characterization of ThasA and ThsB’ proteins.
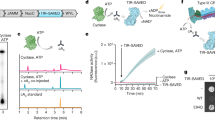
a, SDS-PAGE of ThsA and ThsB’ proteins used in this study. Lane images cropped for clarity. A representative gel of n = 2 independent experiments is shown. For gel source data, see Supplementary Fig. 1. b, Size-exclusion chromatography with multi-angle light scattering analysis tagged and non-tagged ThsB’ indicates that ThsB’ exists as a dimer. Theoretical weights of the dimers are: ThsB’_TwinStrep 52.608 kDa, HisTEV_ThsB’ 49.654 kDa, ThsB’ 45.694 kDa, ThsB_TwinStrep 51.783 kDa. c, ThsB’ uses NAD+ or εNAD+ to produce an isomeric cyclic ADP ribose product. Extracted ion current (EIC) of mass signal corresponding to a cyclic product as well as mass spectra are presented. MS of canonical cADPR is also shown. Shaded peak area represents the analysis time frame of MS spectra shown. ([M-H]− theoretical values: NAD+ 662.1018, cADPR isomer 540.0538, [M-H]+ theoretical values: εNAD+ 688.1164, cεADPR isomer 566.0684). All values are presented in detector counts. Data are representative of n = 3 independent experiments. d, Tagged and non-tagged ThsB’ forms are able to activate ThsA in a εNAD+-based fluorescence assay. Data are presented as mean ± s.d. for n = 3 independent experiments, with individual data points overlaid.
Extended Data Fig. 2 Crystal structure of ThsB’.
a, Topology diagram of ThsB’. Conserved TIR elements are colored light blue, unique ThsB’ structural features are colored light green and lime green. BB-loop is shown in blue. Gray boxes mark the loops that are disordered in both subunits of the asymmetric unit. b, Comparison of the ThsB’ dimer with the dimers of DpNDT (PDB ID 7O6246) and SfSTING (PDB ID 7UN847). Active site residues and complementing residues from the DD loop of the second subunit of DpNDT and SfSTING are shown.
Extended Data Fig. 3 1″−3′ gcADPR binding induced polymerization of ThsA in solution.
Temporal Dynamic Light Scattering (DLS) analysis of the reaction mixtures containing ThsA and ThsB’ started with NAD+. Plotted Z average diameter values qualitatively indicate ThsB’ and NAD+ dependent oligomerization of ThsA into higher order species. Comparison with ThsB’ or NAD+ reaction control indicates that observed oligomerization is by ThsA and not ThsB’ molecules.
Extended Data Fig. 4 The ThsA SLOG domain bound to 1″−3′ gcεADPR.
a, The ThsA SLOG domain dimer, bound to 1″−3′ gcεADPR (pink). 1″−3′ gcADPR (green) is shown for comparison. b, Adenine ring is rotated ~145° in 1″−3′ gcεADPR compared to 1″−3′ gcADPR due to a possible steric clash of two bound 1″−3′ gcεADPR. c, Local refinement unsharpened electrostatic potential map for 1″−3′ gcεADPR and its chemical formula.
Extended Data Fig. 5 Characterization of the ThsA mutants.
a, CD spectroscopy results of WT and mutant ThsA proteins. Mutant proteins and WT exhibited CD spectra indicative of properly folded proteins. b, SEC-MALS analysis of WT and mutant ThsA proteins. Calculated molecular masses of mutant and WT proteins eluting at a single peak indicate tetrameric oligomeric state (ThsA WT monomer theoretical Mw 55.39 kDa). ThsA R371A exhibits polydispersity with multiple elution peaks. c, Temporal dynamic light scattering analysis of mutant ThsA proteins activated using ThsB’ and NAD+.
Extended Data Fig. 6 Structural and biochemical features of the helical filament.
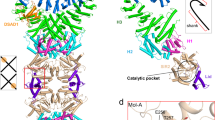
a, Cryo-EM map showing 12 subunits in different colors. Each subunit makes contacts with 5 other subunits. Subunit A makes dimerization contacts with subunit B involving both SLOG and SIR2 domains, also makes tetramerization contacts with C subunit and stabilizing helical interactions with E, I and J subunits. Interactions A-E and I-A are symmetric. b, List of the residues making hydrogen bonds in the tetramerization and helical interfaces (calculated by PDBePISA45). Residues from the elements undergoing conformational changes in SIR2 domain (shown in Fig. 2g) are shown on a light orange background. Helical interface residues NQN are colored magenta, RDR residues are colored blue. c, MS of purified 1″−3′ gcADPR used for further experiments. d, DLS analysis of ThsA−1″−3′ gcADPR complex. Data are represented as boxplots (n = 30 independent measurements) where the middle line is the median, the lower and upper hinges correspond to the first and third quartiles, the upper whisker extends from the hinge to the largest value no further than 1.5 × IQR from the hinge (where IQR is the inter-quartile range) and the lower whisker extends from the hinge to the smallest value at most 1.5 × IQR of the hinge, while data beyond the end of the whiskers are outlying points that are plotted individually as solid dots. Individual data points are overlaid as open dots. An inlet shows a representative micrograph (n = 600) of ThsA−1″−3′ gcADPR 1:1 complex. e, HPCL analysis of 1″−3′ gcADPR after incubation with ThsA. f, Structural superposition of yeast Sir2-Sir4 complex (PDB ID 4IAO) with the ThsA A subunit SIR2 domain. Sir2 is omitted for clarity, Sir4 is colored magenta and shown is in sphere representation. ThsA subunits are colored as in (a), the surface of the ThsA A subunit (violet) is shown.
Extended Data Fig. 7 Cryo-EM helical reconstruction workflow of ThsA-1″−3′ gcADPR complex.
The final electron density maps showing local resolution, masks from the local refinement jobs, directional distribution plots and FSC (Fourier shell correlation) plots of helical (left) and local refinement (A subunit, right) are shown in black rectangles.
Extended Data Fig. 8 Cryo-EM helical reconstruction workflow of ThsA-1″−3′ gcεADPR complex.
The final electron density maps showing local resolution, masks from the local refinement jobs, directional distribution plots and FSC (Fourier shell correlation) plots of helical (left) and local refinement (A subunit, right) are shown in black rectangles.
Supplementary information
Supplementary Information
Supplementary Fig. 1 (uncropped images for Extended Data Fig. 1a) and Tables 1 and 2 (lists of oligonucleotides, strains and plasmids used in the work).
Supplementary Video 1
Helical filament cryo-EM map. Helical refinement cryo-EM map of the activated ThsA helical filament (video associated with Fig. 1).
Supplementary Video 2
SLOG domain structure. The ThsA SLOG domain bound to 1′′–3′ gcADPR and its comparison with the apo-ThsA SLOG domain. The local refinement cryo-EM map is shown (video associated with Fig. 2).
Supplementary Video 3
SIR2 domain structure. The ThsA SIR2 domain bound to NAD+ fragment and its structure comparison with the apo-ThsA SIR2 domain. Loop 34–39, which blocks NAD+ binding in the apo structure, is coloured red. The local refinement cryo-EM map is shown (video associated with Fig. 2).
Supplementary Video 4
ThsA subunit structure. Conformational differences between the apo and 1′′–3′ gcADPR-bound ThsA subunit from the helical filament (video associated with Fig. 3).
Supplementary Video 5
ThsA tetramer structure. Conformational differences between the apo and 1′′–3′ gcADPR-bound ThsA tetramer from the helical filament (video associated with Fig. 3).
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Tamulaitiene, G., Sabonis, D., Sasnauskas, G. et al. Activation of Thoeris antiviral system via SIR2 effector filament assembly. Nature 627, 431–436 (2024). https://doi.org/10.1038/s41586-024-07092-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-024-07092-x
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.