Abstract
Genomic research that targets large-scale, prospective birth cohorts constitutes an essential strategy for understanding the influence of genetics and environment on human health1. Nonetheless, such studies remain scarce, particularly in Asia. Here we present the phase I genome study of the Born in Guangzhou Cohort Study2 (BIGCS), which encompasses the sequencing and analysis of 4,053 Chinese individuals, primarily composed of trios or mother–infant duos residing in South China. Our analysis reveals novel genetic variants, a high-quality reference panel, and fine-scale local genetic structure within BIGCS. Notably, we identify previously unreported East Asian-specific genetic associations with maternal total bile acid, gestational weight gain and infant cord blood traits. Additionally, we observe prevalent age-specific genetic effects on lipid levels in mothers and infants. In an exploratory intergenerational Mendelian randomization analysis, we estimate the maternal putatively causal and fetal genetic effects of seven adult phenotypes on seven fetal growth-related measurements. These findings illuminate the genetic links between maternal and early-life traits in an East Asian population and lay the groundwork for future research into the intricate interplay of genetics, intrauterine exposures and early-life experiences in shaping long-term health.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The release of the raw sequencing data by this work is approved by The Ministry of Science and Technology of the People’s Republic of China (permission number 2022BAT2230) at the National Genomics Data Center (https://ngdc.cncb.ac.cn) (accession number HRA002496). Data can be accessed via applications, following the GSA guide (https://ngdc.cncb.ac.cn/gsa-human/document). The access authority can be obtained for academic research use only. Previous published genotype data for ancient individuals were reported by the Reich laboratory in the Allen Ancient DNA Resource (https://reich.hms.harvard.edu/allen-ancient-dna-resource-aadr-downloadable-genotypes-present-day-and-ancient-dna-data, version 54.1). Researchers who are interested in collaborating with the BIGCS group are welcome to contact X.Q. or data.bigcs@bigcs.org.
Code availability
The ilus (code for BIGCS data variant calling) and GDBIGtools are all available in the Github repository using the following links: ilus: https://github.com/ShujiaHuang/ilus. GDBIGtools: https://github.com/BIGCS-Lab/GDBIGtools. Script for distinguishing the parental haplotype alleles from infant genotype and calculation of the genotype/haplotype-based PRS: https://github.com/ShujiaHuang/genotools/blob/master/scripts/mr.py. Script for detecting of age-specific genetic effects on lipid levels among mothers and infants: https://github.com/ShujiaHuang/genotools/blob/master/scripts/twosamplettest.py. Other software and databases used in this study are publicly available, and the URLs are listed below: SOAPnuke (v1.5.6): https://github.com/BGI-flexlab/SOAPnuke. BWA-MEM (v0.7.17): https://github.com/lh3/bwa. verifyBamID2 (v1.0.6): https://github.com/Griffan/VerifyBamID. GATK (v4.1.8.1): https://github.com/broadgsa/gatk/. SAMtools (v1.9): http://samtools.github.io/. BCFtools (v1.9): https://samtools.github.io/bcftools/bcftools.html. bedtools (v2.27.1-65-gc2af1e7-dirty): https://github.com/arq5x/bedtools2/. Variant Effect Predictor (release 95): https://github.com/Ensembl/ensembl-vep. Beagle (v4.0): https://faculty.washington.edu/browning/beagle/beagle.r1399.jar. Minimac3 (v 2.0.1): http://genome.sph.umich.edu/wiki/Minimac3. AdmixTools (v7.0.2): https://github.com/DReichLab/AdmixTools. MSMC2 (v2.1.1): https://github.com/stschiff/msmc2. CrossMap (version 0.2.2): http://crossmap.sourceforge.net/. dbSNP Build 154: http://www.ncbi.nlm.nih.gov/SNP/. GATK bundle (hg38): https://console.cloud.google.com/storage/browser/genomics-public-data/resources/broad/hg38/v0. Human genome reference (GRCh38/hg38):ftp://ftp.ncbi.nlm.nih.gov/genomes/all/GCA/000/001/405/GCA_000001405.15_GRCh38/seqs_for_alignment_pipelines.ucsc_ids/GCA_000001405.15_GRCh38_no_alt_analysis_set.fna.gz. The low-complexity regions of GRCh38: https://github.com/lh3/varcmp/blob/master/scripts/LCR-hs38.bed.gz. The 1000 Genome Project: https://www.internationalgenome.org/. The GWAS Catalogue: https://www.ebi.ac.uk/gwas/. The Human Protein Atlas: https://www.proteinatlas.org/. The public GWAS SNPs used in constructing genotype-based PRS and haplotype-based PRS: https://doi.org/10.1371/journal.pmed.1003305.s003. We used Python (version 3.7.6) and R (version 4.1.1) extensively to analyse data and create plots. The Venn and admixture plots were created by using a Python library: https://github.com/ShujiaHuang/geneview. Supplementary Figs. 17a and 20 were created using: https://gtexportal.org/. Fig. 4d was created using: https://popgen.uchicago.edu/ggv/.
References
Manolio, T. A., Bailey-Wilson, J. E. & Collins, F. S. Genes, environment and the value of prospective cohort studies. Nat. Rev. Genet. 7, 812–820 (2006).
Qiu, X. et al. The Born in Guangzhou Cohort Study (BIGCS). Eur. J. Epidemiol. 32, 337–346 (2017).
Claussnitzer, M. et al. A brief history of human disease genetics. Nature 577, 179–189 (2020).
Auton, A. et al. A global reference for human genetic variation. Nature 526, 68–74 (2015).
Bycroft, C. et al. The UK Biobank resource with deep phenotyping and genomic data. Nature 562, 203–209 (2018).
Denny, J. C. et al. The ‘all of us’ research program. N. Engl. J. Med. 381, 668–676 (2019).
Barker, D. J. P. The fetal and infant origins of adult disease. Br. Med. J. 301, 1111 (1990).
Gaillard, R. & Jaddoe, V. W. V. Maternal cardiovascular disorders before and during pregnancy and offspring cardiovascular risk across the life course. Nat. Rev. Cardiol. 20, 617–630 (2023).
Sirugo, G., Williams, S. M. & Tishkoff, S. A. The missing diversity in human genetic studies. Cell 177, 26–31 (2019).
Fraser, A. et al. Cohort profile: the Avon Longitudinal Study of Parents and Children: ALSPAC mothers cohort. Int. J. Epidemiol. 42, 97–110 (2013).
Magnus, P. et al. Cohort profile update: the Norwegian Mother and Child Cohort Study (MoBa). Int. J. Epidemiol. 45, 382–388 (2016).
Ernst, A. et al. Cohort profile: the puberty cohort in the Danish National Birth Cohort (DNBC). Int. J. Epidemiol. 49, 373–374 (2020).
Kooijman, M. N. et al. The Generation R Study: design and cohort update 2017. Eur. J. Epidemiol. 31, 1243–1264 (2016).
Middeldorp, C. M., Felix, J. F., Mahajan, A. & McCarthy, M. I. The Early Growth Genetics (Egg) and Early Genetics And Lifecourse Epidemiology (eagle) consortia: design, results and future prospects. Eur. J. Epidemiol. 34, 279–300 (2019).
Metzger, B. E. et al. Hyperglycemia and adverse pregnancy outcomes. N. Engl. J. Med. 358, 1991–2002 (2008).
Kishi, R. et al. Birth Cohort Consortium of Asia: current and future perspectives. Epidemiology 28, S19–S34 (2017).
Tao, F. B. et al. Cohort profile: the China–Anhui Birth Cohort Study. Int. J. Epidemiol. 42, 709–721 (2013).
Hu, Z. B. et al. Profile of China National Birth Cohort. Chinese J. Epidemiol. 42, 569–574 (2021).
Yue, W. et al. The China Birth Cohort Study (CBCS). Eur. J. Epidemiol. 37, 295–304 (2022).
Li, Y., Sidore, C., Kang, H. M., Boehnke, M. & Abecasis, G. R. Low-coverage sequencing: Implications for design of complex trait association studies. Genome Res. 21, 940–951 (2011).
DePristo, M. A. et al. A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nat. Genet. 43, 491–498 (2011).
Liu, S. et al. Genomic analyses from non-invasive prenatal testing reveal genetic associations, patterns of viral infections, and Chinese population history. Cell 175, 347–359.e14 (2018).
Cao, Y. et al. The ChinaMAP analytics of deep whole genome sequences in 10,588 individuals. Cell Res. 30, 717–731 (2020).
Karczewski, K. J. et al. The mutational constraint spectrum quantified from variation in 141,456 humans. Nature 581, 434–443 (2020).
McCarthy, S. et al. A reference panel of 64,976 haplotypes for genotype imputation. Nat. Genet. 48, 1279–1283 (2016).
Wall, J. D. et al. The GenomeAsia 100 K Project enables genetic discoveries across Asia. Nature 576, 106–111 (2019).
Taliun, D. et al. Sequencing of 53,831 diverse genomes from the NHLBI TOPMed Program. Nature 590, 290–299 (2021).
Zhang, P. et al. NyuWa Genome resource: a deep whole-genome sequencing-based variation profile and reference panel for the Chinese population. Cell Rep. 37, 110017 (2021).
Cong, P. K. et al. Genomic analyses of 10,376 individuals in the Westlake BioBank for Chinese (WBBC) pilot project. Nat. Commun. 13, 2939–15 (2022).
Mallick, S. et al. The Allen Ancient DNA Resource (AADR): A curated compendium of ancient human genomes. Preprint at bioRxiv https://doi.org/10.1101/2023.04.06.535797 (2023).
Mao, X. et al. The deep population hiswwwtory of northern East Asia from the Late Pleistocene to the Holocene. Cell 184, 3256–3266.e13 (2021).
Yang, M. A. et al. Ancient DNA indicates human population shifts and admixture in northern and southern China. Science 369, 282–288 (2020).
Ning, C. et al. Ancient genomes from northern China suggest links between subsistence changes and human migration. Nat. Commun. 11, 2700 (2020).
Wang, T. et al. Human population history at the crossroads of East and Southeast Asia since 11,000 years ago. Cell 184, 3829–3841.e21 (2021).
Buniello, A. et al. The NHGRI-EBI GWAS Catalog of published genome-wide association studies, targeted arrays and summary statistics 2019. Nucleic Acids Res. 47, D1005–D1012 (2019).
Kamat, M. A. et al. PhenoScanner V2: an expanded tool for searching human genotype-phenotype associations. Bioinformatics 35, 4851–4853 (2019).
Hayes, M. G. et al. Identification of HKDC1 and BACE2 as genes influencing glycemic traits during pregnancy through genome-wide association studies. Diabetes 62, 3282–3291 (2013).
Peng, L. et al. The p.Ser267Phe variant in SLC10A1 is associated with resistance to chronic hepatitis B. Hepatology 61, 1251–1260 (2015).
Ovadia, C. et al. Association of adverse perinatal outcomes of intrahepatic cholestasis of pregnancy with biochemical markers: results of aggregate and individual patient data meta-analyses. Lancet 393, 899–909 (2019).
Warrington, N. M. et al. Maternal and fetal genetic contribution to gestational weight gain. Int. J. Obes. 42, 775–784 (2018).
Safran, M. et al. GeneCards version 3: the human gene integrator. Database 2010, baq020 (2010).
Smith, J. R. et al. The Year of the Rat: the Rat Genome Database at 20: a multi-species knowledgebase and analysis platform. Nucleic Acids Res. 48, D731–D742 (2020).
Marissal-Arvy, N. et al. QTLs influencing carbohydrate and fat choice in a LOU/CxFischer 344 F2 rat population. Obesity 22, 565–575 (2014).
Juliusdottir, T. et al. Distinction between the effects of parental and fetal genomes on fetal growth. Nat. Genet. 53, 1135–1142 (2021).
Han, Z., Lutsiv, O., Mulla, S. & McDonald, S. D. Maternal height and the risk of preterm birth and low birth weight: a systematic review and meta-analyses. J. Obstet. Gynaecol. Canada 34, 721–746 (2012).
Voigt, M. et al. Individualized birth length and head circumference percentile charts based on maternal body weight and height. J. Perinat. Med. 48, 656–664 (2020).
Teng, H. et al. Gestational systolic blood pressure trajectories and risk of adverse maternal and perinatal outcomes in Chinese women. BMC Pregnancy Childbirth 21, 155 (2021).
Chen, J. et al. Dissecting maternal and fetal genetic effects underlying the associations between maternal phenotypes, birth outcomes, and adult phenotypes: a Mendelian-randomization and haplotype-based genetic score analysis in 10,734 mother–infant pairs. PLoS Med. 17, e1003305 (2020).
Baker, H. D. R. Language atlas of China. Bull. Sch. Orient. Afr. Stud. 56, 398–399 (1993).
Chen, Y. et al. SOAPnuke: A MapReduce acceleration-supported software for integrated quality control and preprocessing of high-throughput sequencing data. Gigascience 7, gix120 (2018).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Zhang, F. et al. Ancestry-agnostic estimation of DNA sample contamination from sequence reads. Genome Res. 30, 185–194 (2020).
Li, H. A statistical framework for SNP calling, mutation discovery, association mapping and population genetical parameter estimation from sequencing data. Bioinformatics 27, 2987–2993 (2011).
Browning, B. L. & Yu, Z. Simultaneous genotype calling and haplotype phasing improves genotype accuracy and reduces false-positive associations for genome-wide association studies. Am. J. Hum. Genet. 85, 847–861 (2009).
McLaren, W. et al. The Ensembl variant effect predictor. Genome Biol. 17, 122 (2016).
Das, S. et al. Next-generation genotype imputation service and methods. Nat. Genet. 48, 1284–1287 (2016).
Yu, K. et al. Meta-imputation: an efficient method to combine genotype data after imputation with multiple reference panels. Am. J. Hum. Genet. 109, 1007–1015 (2022).
Alexander, D. H., Novembre, J. & Lange, K. Fast model-based estimation of ancestry in unrelated individuals. Genome Res. 19, 1655–1664 (2009).
Chang, C. C. et al. Second-generation PLINK: rising to the challenge of larger and richer datasets. Gigascience 4, https://doi.org/10.1186/s13742-015-0047-8 (2015).
Price, A. L. et al. Principal components analysis corrects for stratification in genome-wide association studies. Nat. Genet. 38, 904–909 (2006).
Patterson, N. et al. Ancient admixture in human history. Genetics 192, 1065–1093 (2012).
Wangkumhang, P., Greenfield, M. & Hellenthal, G. An efficient method to identify, date, and describe admixture events using haplotype information. Genome Res. 32, 1553–1564 (2022).
Hellenthal, G. et al. A genetic atlas of human admixture history. Science 343, 747–751 (2014).
Zhou, W. et al. Scalable generalized linear mixed model for region-based association tests in large biobanks and cohorts. Nat. Genet. 52, 634–639 (2020).
Pruim, R. J. et al. LocusZoom: regional visualization of genome-wide association scan results. Bioinformatics 26, 2336–2337 (2010).
Lonsdale, J. et al. The Genotype–Tissue Expression (GTEx) project. Nat. Genet. 45, 580–585 (2013).
Zhao, H. et al. CrossMap: a versatile tool for coordinate conversion between genome assemblies. Bioinformatics 30, 1006–1007 (2014).
Acknowledgements
This study was supported by the Ministry of Science and Technology of the People’s Republic of China (2022YFC2702903, 2022YFC2704601, 2021ZD0200536), the National Natural Science Foundation of China (81673181, 82173525, 31900487, 82003471, 82273642), the Department of Science and Technology of Guangdong Province (2020B1111170001, 2018B030335001, 2019B030301004, 2022B1212010004), Guangdong Basic and Applied Basic Research Foundation (2022B1515120080, 2020A1515110859), Science and Technology Planning Project of Guangdong Province (2019B020227001, 2019B030316014), the Guangzhou Municipal Science and Technology Bureau (202201020656, 202102010254, 202007030002), the Guangzhou Municipal Health Commission (2023A031001), and Shenzhen Basic Research Foundation (20220818100717002). We are grateful to all the participants in BIGCS project. We thank C. Gao, P. Huang, X. Liu, Y. Hu and all colleagues at GWCMC who have provided invaluable assistance to the BIGCS project; G. Zhang for useful discussions on Mendelian randomization analysis; and the professional technical support service provide by W. Lai, L. Wei and S. Liu for setting up the GDBIG website. We thank the Tianhe-2 Supercomputer Center in Guangzhou for support of computational and storage resources. We would also like to acknowledge the Genotype-Tissue Expression (GTEx) Project, for providing figure data used in Fig. 4b, Supplementary Figs. 17 and 20. The data used for the analyses described in this manuscript were obtained from the GTEx portal on 7 July 2022.
Author information
Authors and Affiliations
Consortia
Contributions
Conceptualization: X.Q., S.H. and S. Liu Sample collection and data curation: Y.K., J.L. and X.X. Investigation: S.H., M.H., S. Liu and C.W. Methodology: S.H., S. Liu and Q.F. Formal analysis: S.H., M.H., C.W., S. Liu, T.W., Q.F. and X.F. Visualization: S.H., M.H., C.W. and S. Liu. Software: S.H., M.H. and C.W. Validation: S. Liu, Y. Gu, M.H., S.H., J.L., X.X., Y.K. and J.H. Writing, original draft: S. Liu and S.H. Writing, review and editing: S. Liu, S.H., Q.F., T.W., X.Q., J.H., S. Lin and W.Z. Project administration: X.Q., S.H. and J.H. Supervision: X.Q. and H.X. Resources: X.Q. and H.X.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks Rachel Freathy, Sarah Gagliano Taliun, Chuan-Chao Wang and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer review reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Read alignment, variant calling, filtering, and genotype refinement.
A comprehensive description of the quality control and bioinformatics analysis is available in the Methods and Supplementary Notes.
Extended Data Fig. 2 Assessment of imputation accuracy and coverage of imputed variants compared to true set variants from WGS.
For variants present in the true set but absent from the reference panel, an R2 value of zero was assigned. The X-axis represents allele frequency, estimated based on the BIGCS dataset. In cases where a variant was unavailable in BIGCS, its allele frequency was estimated using data from the 50 WGS samples.
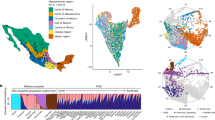
Extended Data Fig. 3 Principal component analysis (PCA) comparing linguistic groups in BIGCS and nine present-day Asian linguistic families in AADR.
(a-b) Geographic distribution of the 4,053 BIGCS participants (a) and all the 836 Chinese samples from the AADR dataset (b). (c-d) PCA was conducted on a merged sample comprising 2,245 present-day unrelated Chinese individuals from the BIGCS dataset, 402 present-day Chinese individuals from the AADR dataset, and 202 present-day Asian groups from the AADR dataset. Each data point on the PCA plot represents one participant, with colors and shapes denoting their linguistic or ethnic groups. In plot (c), nine different shapes were used to represent nine linguistic families. In (d), the shapes remained consistent, with additional colors assigned to each linguistic family to represent linguistic groups. The analysis utilized 258,552 biallelic sites and applied the following pruning and filtration parameters: “--maf 0.01 --geno 0.3 --hwe 1e-6 --vcf-half-call m --indep-pairwise 1000 100 0.9”. The map in panels a and b was sourced from an approved standard map service (http://bzdt.ch.mnr.gov.cn) endorsed by the Ministry of National Resources of the People’s Republic of China (GS YUE(2023)1422).
Extended Data Fig. 4 Genetic structure and admixture of participants in the BIGCS study.
Ancestral components were determined for each participant using ADMIXTURE, considering K values ranging from 2 to 5. This analysis included the five linguistic groups with a sample size greater than 50. The optimal number of ancestral components, determined by the smallest cross-validation error, was found to be K = 3. These three ancestral components were visually represented by the colors green, orange, and blue, which corresponded to ancestral components enriched with Cantonese, Min, and Mandarin respectively. Mandarin speakers were further subdivided into seven groups based on their birthplace: Mandarin_SC (South China), Mandarin_SWC (Southwest China), Mandarin_CC (Central China), Mandarin_EC (East China), Mandarin_NWC (Northwest China), Mandarin_NEC (Northeast China), Mandarin_NC (North China).
Extended Data Fig. 5 LocusZoom plots of the remaining 12 loci reaching study-wide association significance (P < 2.78 × 10−9) besides the SLC10A1 locus.
Detailed information about the lead variants is provided in Table S9. LD r2 calculations were performed using East Asian populations from the 1KGP dataset, except for the TTC28 and SOAT2 loci. As the lead SNPs for these two loci were absent in the 1KGP dataset, LD r2 was computed using the BIGCS reference panel through the pairwise LD research tool available on the BIGCS website (http://gdbig.bigcs.com.cn/ld/cal.html). The LocusZoom plot illustrating the SLC10A1 locus association is presented in Fig. 4a.
Extended Data Fig. 6 Observed phenotypic associations, estimated effects of parentally transmitted alleles, maternal non-transmitted alleles, maternal causal effect, and fetal genetic effect per one-unit change in maternal phenotypes on birth outcomes.
Measurement Units: 1 cm (height), 1 kg/m2 (BMI), 1 mmHg (BP), 1 mmol/L (FPG, TC and TG) and 1 umol/L (TBA).
Supplementary information
Supplementary Information
This document offers comprehensive details on variant calling, population genetic analysis, the assessment of age-specific genetic effects, and intergenerational Mendelian randomization conducted on the BIGCS dataset. It includes Supplementary Notes, Supplementary Figs. 1–21, and a reference guide for Supplementary Tables 1–20.
Supplementary Table 1
Geographic distribution of BIGCS cohort samples investigated in this study. Related to Fig. 1.
Supplementary Table 2
Ethnicity distribution of the BIGCS cohort samples investigated in this study. Related to Fig. 1.
Supplementary Table 3
Summary statistics of sequencing data and variant detected in each BIGCS individual.
Supplementary Table 4
Number of variants and average imputation accuracy for a range of Minimac3 estimated R-squares thresholds and reference panels.
Supplementary Table 5
Evaluation of mean imputation accuracy of the BIGCS reference panel imputation, in comparison with four commonly used reference panels.
Supplementary Table 6
Supplementary Table 6a: Geographic distribution of Chinese sample form AADR samples investigated in this study. Related to Extended Data Fig. 3b. Supplementary Table 6b: Information on 56 present-day and ancient Asian groups from AADR used in admixture analysis.
Supplementary Table 7
Supplementary Table 7a: Symmetric f4 test comparing each BIGCS linguistic groups with Neolithic and Pre-neolithic northern and southern Asian groups. Supplementary Table 7b: Symmetric f4 test comparing each BIGCS linguistic groups with six deep Asian lineages and two present-day Chinese groups. Supplementary Table 7c. Successful qpAdm results for the BIGCS linguistic groups assuming one, two and three sources.
Supplementary Table 8
Summary of Chromopainter and Globetrotter analysis for the five major linguistic groups in BIGCS.
Supplementary Table 9
Supplementary Table 9a: Genome-wide associations signals reaching study-wide significance threshold (P < 2.78e-9). Related to Figs. 3–5. Supplementary Table 9b: Genome-wide association signals reaching genome-wide significance threshold (P < 5 × 10−8). Related to Figs. 3–5.
Supplementary Table 10
Nearby variants with linkage disequilibrium r2 > 0.2 surrounding the 4-bp deletion (rs3840091) associated with GWG and comparison with a GWG GWAS in the European population. Related to Fig. 3.
Supplementary Table 11
Genome-wide association analysis of the four lipid traits using adult and infant samples jointly with SAIGE.
Supplementary Table 12
Supplementary Table 12a: Ratio estimates for intergenerational Mendelian randomization analysis without restriction of instrumental variables (primary analysis). Supplementary Table 12b: Ratio estimates for intergenerational Mendelian randomization analysis excluding specific instrumental variables (secondary analysis).
Supplementary Table 13
Supplementary Table 13a: Intergenerational Mendelian randomization analysis for normalized maternal trait measurements and normalized fetal growth measurements without restriction of instrumental variables. Supplementary Table 13b:Intergenerational Mendelian randomization analysis for normalized maternal trait measurements and normalized fetal growth measurements excluding specific instrumental variables.
Supplementary Table 14
GWAS SNPs used to calculate the genetic scores for maternal height (POS: GRCh38 coordinate; A: effect allele).
Supplementary Table 15
GWAS SNPs used to calculate the genetic scores for maternal pre-pregnancy BMI (POS: GRCh38 coordinate; A: effect allele).
Supplementary Table 16
GWAS SNPs used to calculate the genetic scores for fasting plasma glucose (FPG) (POS: GRCh38 coordinate; A: effect allele).
Supplementary Table 17
GWAS SNPs used to calculate the genetic scores for blood pressure (POS: GRCh38 coordinate; A: effect allele).
Supplementary Table 18
GWAS SNPs used to calculate the genetic scores for TBA (POS: GRCh38 coordinate; A: effect allele).
Supplementary Table 19
GWAS SNPs used to calculate the genetic scores for triglyceride (POS: GRCh38 coordinate; A: effect allele).
Supplementary Table 20
GWAS SNPs used to calculate the genetic scores for total cholesterol (POS: GRCh38 coordinate; A: effect allele).
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Huang, S., Liu, S., Huang, M. et al. The Born in Guangzhou Cohort Study enables generational genetic discoveries. Nature 626, 565–573 (2024). https://doi.org/10.1038/s41586-023-06988-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-023-06988-4
This article is cited by
-
An early look at birth cohort genetics in China
Nature (2024)
-
New genetic variants found in large Chinese mother–baby study
Nature (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.