Abstract
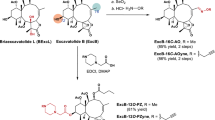
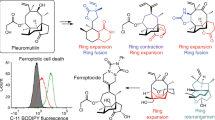
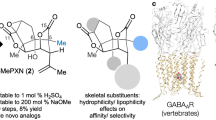
Marine-derived cyclic imine toxins, portimine A and portimine B, have attracted attention because of their chemical structure and notable anti-cancer therapeutic potential1,2,3,4. However, access to large quantities of these toxins is currently not feasible, and the molecular mechanism underlying their potent activity remains unknown until now. To address this, a scalable and concise synthesis of portimines is presented, which benefits from the logic used in the two-phase terpenoid synthesis5,6 along with other tactics such as exploiting ring-chain tautomerization and skeletal reorganization to minimize protecting group chemistry through self-protection. Notably, this total synthesis enabled a structural reassignment of portimine B and an in-depth functional evaluation of portimine A, revealing that it induces apoptosis selectively in human cancer cell lines with high potency and is efficacious in vivo in tumour-clearance models. Finally, practical access to the portimines and their analogues simplified the development of photoaffinity analogues, which were used in chemical proteomic experiments to identify a primary target of portimine A as the 60S ribosomal export protein NMD3.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The data that support the findings in this work are available in the paper and Supplementary Information. Uncropped, full western blot images and gels are provided in Supplementary Figs. 2 and 3. All raw proteomics data files have been deposited to the PRIDE53 repository and are available under the accession PXD041911. Source data are provided with this paper.
Change history
05 October 2023
A Correction to this paper has been published: https://doi.org/10.1038/s41586-023-06699-w
References
Stivala, C. E. et al. Synthesis and biology of cyclic imine toxins, an emerging class of potent, globally distributed marine toxins. Nat. Prod. Rep. 32, 411–435 (2015).
Molgó, J. et al. Cyclic imine toxins from dinoflagellates: a growing family of potent antagonists of the nicotinic acetylcholine receptors. J. Neurochem. 142, 41–51 (2017).
Selwood, A. I. et al. Portimine: a bioactive metabolite from the benthic dinoflagellate Vulcanodinium rugosum. Tetrahedron Lett. 54, 4705–4707 (2013).
Cuddihy, S. L. et al. The marine cytotoxin portimine is a potent and selective inducer of apoptosis. Apoptosis 21, 1447–1452 (2016).
Jørgensen, L. et al. 14-Step synthesis of (+)-ingenol from (+)-3-carene. Science 341, 878–882 (2013).
Kanda, Y. et al. Two-phase synthesis of taxol. J. Am. Chem. Soc. 142, 10526–10533 (2020).
Munday, R., Selwood, A. I. & Rhodes, L. Acute toxicity of pinnatoxins E, F and G to mice. Toxicon 60, 995–999 (2012).
Munday, R. et al. Investigations into the toxicology of spirolides, a group of marine phycotoxins. Toxins 4, 1–14 (2012).
Munday, R. et al. Acute toxicity of gymnodimine to mice. Toxicon 44, 173–178 (2004).
Fribley, A. M. et al. Identification of Portimine B, a new cell permeable spiroimine that induces apoptosis in oral squamous cell carcinoma. ACS Med. Chem. Lett. 10, 175–179 (2019).
Hermawan, I. et al. Kabirimine, a new cyclic imine from an Okinawan dinoflagellate. Mar. Drugs 17, 353 (2019).
MacKinnon, S. L. et al. Biosynthesis of 13-desmethyl spirolide C by the dinoflagellate Alexandrium ostenfeldii. J. Org. Chem. 71, 8724–8731 (2006).
Kellmann, R., Stüken, A., Orr, R. J. S., Svendsen, H. M. & Jakobsen, K. S. Biosynthesis and molecular genetics of polyketides in marine dinoflagellates. Mar. Drugs 8, 1011–1048 (2010).
Van Wagoner, R. M., Satake, M. & Wright, J. L. C. Polyketide biosynthesis in dinoflagellates: what makes it different? Nat. Prod. Rep. 31, 1101–1137 (2014).
McCauley, J. A. et al. Total synthesis of pinnatoxin A. J. Am. Chem. Soc. 120, 7647–7648 (1998).
Stivala, C. E. & Zakarian, A. Total synthesis of (+)-pinnatoxin A. J. Am. Chem. Soc. 130, 3774–3776 (2008).
Nakamura, S., Kikitchi, F. & Hashimoto, S. Total synthesis of pinnatoxin A. Angew. Chem. Int. Ed. 47, 7091–7094 (2008).
Araoz, R. et al. Total synthesis of pinnatoxins A and G and revision of the mode of action of pinnatoxin A. J. Am. Chem. Soc. 133, 10499–10511 (2011).
Kong, K., Moussa, Z., Lee, C. & Romo, D. Total synthesis of the spirocyclic imine marine toxin (−)-gymnodimine and an unnatural C4-epimer. J. Am. Chem. Soc. 133, 19844–19856 (2011).
Saito, T., Fujiwara, K., Kondo, Y., Akiba, U. & Suzuki, T. Synthesis of the cyclohexene segment of portimine. Tetrahedron Lett. 60, 386–389 (2019).
Aitken, H. R. M., Brimble, M. A. & Furkert, D. P. A catalytic asymmetric ene reaction for direct preparation of α-Hydroxy 1,4-diketones as intermediates in natural product synthesis. Synlett 31, 687–690 (2020).
Ding, X.-B., Aitken, H. R. M., Pearl, E. S., Furkert, D. P. & Brimble, M. A. Synthesis of the C4-C16 polyketide fragment of portimines A and B. J. Org. Chem. 86, 12840–12850 (2021).
Li, L., El Khoury, A., Clement, B. O., Wu, C. & Harran, P. G. Asymmetric organocatalysis enables rapid assembly of portimine precursor chains. Org. Lett. 24, 2607–2612 (2022).
Fürstner, A. Alkyne metathesis on the rise. Angew. Chem. Int. Ed. 52, 2794–2819 (2013).
Huang, Y., Iwama, T. & Rawal, V. H. Design and development of highly effective Lewis acid catalysts for enantioselective Diels–Alder reactions. J. Am. Chem. Soc. 124, 5950–5951 (2002).
Corey, E. J. & Beames, D. J. Mixed cuprate reagents of type R1R2CuLi which allow selective group transfer. J. Am. Chem. Soc. 94, 7210–7211 (1972).
Hillenbrand, J. et al. “Canopy catalysts” for alkyne metathesis: molybdenum alkylidyne complexes with a tripodal ligand framework. J. Am. Chem. Soc. 142, 11279–11294 (2020).
Chen, S. et al. Ruthenium-catalyzed oxidation of alkenes at room temperature: a practical and concise approach to α-diketones. Org. Lett. 13, 2274–2277 (2011).
Cummins, C. H. & Coates, R. M. α-Oxygenation of aldehydes and cyclic-ketones by acylation rearrangement of nitrones. J. Org. Chem. 48, 2070–2076 (1983).
Wurdak, H. et al. An RNAi screen identifies TRRAP as a regulator of brain tumor-initiating cell differentiation. Cell Stem Cell 6, 37–47 (2010).
Li, Z. et al. Design and synthesis of minimalist terminal alkyne-containing diazirine photo-crosslinkers and their incorporation into kinase inhibitors for cell- and tissue-based proteome profiling. Angew. Chem. Int. Ed. 52, 8551–8556 (2013).
Parker, C. G. et al. Ligand and target discovery by fragment-based screening in human cells. Cell 168, 527–541 (2017).
Parker, C. G. et al. Chemical proteomics identifies SLC25A20 as a functional target of the ingenol class of actinic keratosis drugs. ACS Cent. Sci. 3, 1276–1285 (2017).
Conway, L. P. et al. Evaluation of fully-functionalized diazirine tags for chemical proteomic applications. Chem. Sci. 12, 7839–7847 (2021).
McAlister, G. C. et al. MultiNotch MS3 enables accurate, sensitive, and multiplexed detection of differential expression across cancer cell line proteomes. Anal. Chem. 86, 7150–7158 (2014).
Martinez Molina, D. et al. Monitoring drug target engagement in cells and tissues using the cellular thermal shift assay. Science 341, 84–87 (2013).
Sengupta, J. et al. Characterization of the nuclear export adaptor protein Nmd3 in association with the 60S ribosomal subunit. J. Cell Biol. 189, 1079–1086 (2010).
Ho, J. H.-N., Kallstrom, G. & Johnson, A. W. Nmd3p is a Crm1p-dependent adapter protein for nuclear export of the large ribosomal subunit. J. Cell Biol. 151, 1057–1066 (2000).
Trotta, C. R., Lund, E., Kahan, L., Johnson, A. W. & Dahlberg, J. E. Coordinated nuclear export of 60S ribosomal subunits and NMD3 in vertebrates. EMBO J. 22, 2841–2851 (2003).
Ceci, M. et al. Release of eIF6 (p27BBP) from the 60S subunit allows 80S ribosome assembly. Nature 426, 579–584 (2003).
Malyutin, A. G., Musalgaonkar, S., Patchett, S., Frank, J. & Johnson, A. W. Nmd3 is a structural mimic of eIF5A, and activates the cpGTPase Lsg1 during 60S ribosome biogenesis. EMBO J. 36, 854–868 (2017).
Koga, Y. et al. Discovery of C13-aminobenzoyl cycloheximide derivatives that potently inhibit translation elongation. J. Am. Chem. Soc. 143, 13473–13477 (2021).
Tang, R. et al. Semisynthetic homoharringtonine induces apoptosis via inhibition of protein synthesis and triggers rapid myeloid cell leukemia-1 down-regulation in myeloid leukemia cells. Mol. Cancer Ther. 5, 723–731 (2006).
Chen, R. et al. Homoharringtonine reduced Mcl-1 expression and induced apoptosis in chronic lymphocytic leukemia. Blood 117, 156–164 (2011).
Zhang, X. et al. Targeting translation initiation by synthetic rocaglates for treating MYC-driven lymphomas. Leukemia 34, 138–150 (2020).
Schneider-Poetsch, T. et al. Inhibition of eukaryotic translation elongation by cycloheximide and lactimidomycin. Nat. Chem. Biol. 6, 209–217 (2010).
Manier, S. et al. Inhibiting the oncogenic translation program is an effective therapeutic strategy in multiple myeloma. Sci. Transl. Med. 9, eaal2668 (2017).
Alinari, L. et al. Dual targeting of the cyclin/Rb/E2F and mitochondrial pathways in mantle cell lymphoma with the translation inhibitor silvestrol. Clin. Cancer Res. 18, 4600–4611 (2012).
Lindqvist, L. M. et al. Translation inhibitors induce cell death by multiple mechanisms and Mcl-1 reduction is only a minor contributor. Cell Death Dis. 3, e409(2012).
Chen, R., Gandhi, V. & Plunkett, W. A sequential blockade strategy for the design of combination therapies to overcome oncogene addiction in chronic myelogenous leukemia. Cancer Res. 66, 10959–10966 (2006).
Peters, D. S. et al. Ideality in context: motivations for total synthesis. Acc. Chem. Res. 54, 605–617 (2021).
Wang, M. Y. et al. Mammalian target of rapamycin inhibition promotes response to epidermal growth factor receptor kinase inhibitors in PTEN-deficient and PTEN-intact glioblastoma cells. Cancer Res. 66, 7864–7869 (2006).
Perez-Riverol, Y. et al. The PRIDE database resources in 2022: A hub for mass spectrometry-based proteomics evidences. Nucleic Acids Res. 50, 543–552 (2022).
Acknowledgements
We thank J. Williamson and A. Popova (Scripps Research) for their discussions and technical assistance with polysome profiling experiments. We thank J. Teijaro and D. Lazar (Scripps Research) for their assistance with PBMC isolation. We are grateful to D.-H. Huang and L. Pasternack (Scripps Research) for NMR spectroscopic assistance and B. Sanchez, Q. N. Wong and CoreService Team (Scripps Research) for analytical support. We thank E. Esquenazi, T. Schwent and P. Stout (Sirenas Marine Discovery) for first alerting us to the structure and bioactivity of PA, as well as providing NMR spectra of the authentic sample. We thank A. Fürstner and J. Hillenbrand (Max-Planck-Institut für Kohlenforschung) for providing RCAM catalysts. Financial support for this work was provided by NIGMS (GM118176).
Author information
Authors and Affiliations
Contributions
The conceptualization was done by J.T., W.L., L.L.L., C.G.P. and P.S.B. The experimental investigation was carried out by J.T., W.L., T.-Y.C., F.M.-P., Z.L., C.T.C., Q.W., N.G., T.J.W. and Y. Y. S. The data analysis was done by J.T., W.L., T.-Y.C., F.M.-P., Q.W., N.G., L.L.L., C.G.P. and P.S.B. The manuscript was written by J.T., W.L., L.L.L., C.G.P. and P.S.B. The funding was acquired by P.S.B. The project administration was done by L.L.L., C.G.P. and P.S.B. The supervision was carried out by L.L.L., C.G.P. and P.S.B.
Corresponding authors
Ethics declarations
Competing interests
Scripps Research filed a US patent application on 17 January 2023 covering the chemical structures described in this Article and their use. P.S.B., C.G.P., L.L.L., J.T., W.L., F.M.-P. and T.-Y.C. are listed as inventors on this patent. The other authors declare no other competing interests.
Peer review
Peer review information
Nature thanks Raphaël Rodriquez, Nikolai Slavov and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Portimine A (PA) inhibits the growth of human and mouse cell lines in the low nanomolar range.
Concentration−response curves for PA and IC50 (95% CI) values in human (a-q) and mouse (r-t) cell lines: a) Jurkat (leukemia). b) RD (rhabdomyosarcoma). c) HT-1080 (fibrosarcoma). d) A673 (Ewings sarcoma). e) HCC1806 (acantholytic squamous cell carcinoma). f) HeLa (cervical adenocarcinoma). g) MCF-7 (breast adenocarcinoma). h) MDA-MB-231 (breast adenocarcinoma). i) SUM159 (breast adenocarcinoma). j) HepG2 (hepatocellular carcinoma). k) LNCaP (prostate carcinoma). l) 786-O (renal adenocarcinoma). m) GBM-A (glioblastoma). n) GBM-F (glioblastoma). o) LN229 (glioblastoma). p) U87EGFRvIII (glioblastoma). q) U118 (glioblastoma). r) MC38 (colon carcinoma). s) B16-F10 (melanoma). t) 4T1 (mammary carcinoma). Cells were treated with PA at different concentrations for 72 h, followed by the CellTiter-Glo proliferation assay. Data represents mean ± s.d. of three biologically replicated experiments.
Extended Data Fig. 2 Portimine A triggers apoptosis and cell cycle arrest in Jurkat cells.
(a) PA induced-toxicity (24 h) can be rescued by caspase pan-inhibitor Z-VAD-FMK (50 μM) in Jurkat cells. Data are mean ± s.d. (n = 6 biologically independent samples). Statistical analysis performed using multiple unpaired two-tailed Student t-test; P-values are shown. (b-e) Cell cycle analysis in Jurkat cells. Jurkat cells were treated with 1 nM of indicated compounds for 12 h and propodium iodide used to identify different stages of cells by flow cytometry. Shown are G1 (b), S (c), G2 (d) and SubG1 (e) phase distributions. Data are mean ± s.d of three independent biologically replicated experiments. Statistical analysis was performed using one-way ANOVA analysis with multiple comparisons. P-values are shown.
Extended Data Fig. 3 Portimine A and analogs do not affect cell viability or induce apoptosis in freshly isolated human PBMCs.
(a) PA displays minimal viability effects on human PBMCs. ePA displays no toxicity to both Jurkat and PBMCs at the concentration range displayed. All presented data as the mean ± s.d. of biological replicated experiments (n = 3). (b) FACS analysis of apoptosis using annexin V/ eFluor 780 viability dye after PA and analogs (12 h) treatment in PBMCs. All data presented as mean ± s.d. of biological replicated experiments (n = 3). (c) Immunoblot of caspase-3 and PARP1 in PBMCs with indicated conditions (n = 3 biologically independent samples). For uncropped immunoblot images, see Supplementary Fig. 3.
Extended Data Fig. 4 Portimine A mouse pharmacokinetic properties and fast acting in vitro cell-based target engagement properties based on compound wash-out.
(a) Pharmacokinetic studies of mouse intraperitoneal (i.p.) and oral (p.o.) administration for portimine. Data presented as mean ± s.d. (n = 3 biologically independent samples). (b-d) Washout experiment performed in b) Jurkat c) MC38 and d) HT-1080 cells showed exposure-dependent decrease in IC50 and revealed PA has a fast-acting cytotoxicity mechanism. Data presented as mean ± s.d. (n = 3 biologically independent samples). (e) FACS analysis of apoptosis using annexin V/ eFluor 780 viability dye after PA and analogs (24 h) treatment in MC38 cell line. Data presented as mean ± s.d. of biological replicated experiments (n = 3 biologically independent samples). Statistical analysis was performed using one-way ANOVA analysis with multiple comparisons ; P-values are shown. (f) Kaplan-Myer survival curve of WT C57BL/6 MC38 tumour-bearing mice (n = 6 mice per group) after treatment with PA 0.3 or 1 mg kg−1 intraperitoneally.
Extended Data Fig. 5 Chemical proteomic analysis reveals NMD3 is the target of portimine A.
(a) Chemical proteomic profiling with PA-DA in HCC1806. X-axis corresponds to PA-DA (500 nM) enriched proteins competed by PA (PA, 8×); y-axis corresponds to proteins enriched by PA-DA over ePA-DA (500 nM). Designated PA-specific targets in red (competed by active competitor > 5-fold; enriched by PA-DA > 2-fold; and > 4-fold competition difference between PA and ePA). Dotted lines indicate competition (x-axis) and enrichment (y-axis) thresholds. Data presented as mean of biological replicates (n = 2). See Supplementary Tables 7–9 for source data. (b-d) Immunoblot of NMD3 engagement by PA-DA (500 nM) co-treated with PA or ePA (8×) as well as by ePA-DA (500 nM) in HeLa (b), HCC1806 (c), and MC38 cells (d). (n = 2 biologically independent samples). (e) NMD3 is engaged by PA-DA in a dose-dependent manner in Jurkat cells. Results are representative of three independent experiments. (f-i) CETSA validation of NMD3 as a target of PA in Jurkat and MC38 cells. (f-g) Immunoblotting and quantitation of NMD3 thermal aggregation curves (mean ± s.d.) in MC38 cells treated with PA. (n = 3 biologically independent samples). (h-i) Dose-response (ITDR) fingerprint of NMD3 stabilization by PA in Jurkat cells and corresponding quantitation. Data presented as mean ± s.d. of biological replicated experiments (n = 3). Statistical analysis performed using multiple unpaired two-tailed Student t-test; P-values are shown. (j) Isothermal dose-response (ITDR) fingerprint of NMD3 stabilization by PA in MC38 cells (n = 3 biologically independent samples). For uncropped immunoblot images, see Supplementary Fig. 3.
Extended Data Fig. 6 Portimine A activity dependent on NMD3 and impairs polysome formation.
(a) Immunoblot of NMD3 in HeLa and MC38 cells expressing control or NMD3-specific shRNAs. (b-c) PA has reduced viability effects in Jurkat cells transduced with shRNA targeting NMD3 compared to shCtrl cells in HeLa (b, 1 nM, 24 h) and MC38 (c, 1 nM, 48 h). Data presented as mean ± s.d. of biologically replicated experiments (n = 3 for HeLa, n = 6 for MC38). Statistical analysis was performed using unpaired two-tailed Student t-test. P-values are shown. (d) Quantitation of 60 S:80 S ratio and 80 S:polysome ratio from data shown in Fig. 4f. Presented as mean ± s.d. of biological replicates (n = 3). Statistical analysis performed using unpaired two-tailed Student t-test; P-values are shown. (e) Polysome profiling of MC38 cells treated with PA (50 nM, 6 h). Results are representative of two independent experiments. (f) PA, but not ePA, inhibits new protein synthesis in HeLa, HCC1806, and MC38 cells as determined by O-propargyl puromycin incorporation. Results normalized to vehicle and presented as the mean ± s.d. across biologically replicated experiments (n = 3). Statistical analysis was performed using multiple unpaired two-tailed Student t-test. P-values are shown. (g, h) Immunoblot of Mcl-1 and c-Myc after PA and ePA treatment in g) Jurkat and h) MC38 cells. Results are representative of two independent experiments. (i) Quantitation of protein abundances from Fig. 4j. Results presented as mean ± s.d. of biologically replicates (n = 3). Statistical analysis was performed using multiple unpaired two-tailed Student t-test; P-values are shown. (j) MCL1 and MYC mRNA expression assessed by quantitative PCR in Jurkat cells treated with PA (10 nM). Results normalized to vehicle and values indicate mean ± s.d. (n = 3 biologically independent samples). Statistical analysis performed using multiple unpaired two-tailed Student t-test. P-values are shown. For uncropped immunoblot images, see Supplementary Fig. 3.
Supplementary information
Supplementary Information
This file contains Supplementary Materials and Methods; Synthetic Procedures, Supplementary Tables 1–6, NMR spectra and Supplementary References.
Supplementary Tables
This file contains Supplementary Tables 7–9.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Tang, J., Li, W., Chiu, TY. et al. Synthesis of portimines reveals the basis of their anti-cancer activity. Nature 622, 507–513 (2023). https://doi.org/10.1038/s41586-023-06535-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-023-06535-1
This article is cited by
-
Functionalizing tandem mass tags for streamlining click-based quantitative chemoproteomics
Communications Chemistry (2024)
-
Chemoproteomic development of SLC15A4 inhibitors with anti-inflammatory activity
Nature Chemical Biology (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.