Abstract
Fasting initiates a multitude of adaptations to allow survival. Activation of the hypothalamic–pituitary–adrenal (HPA) axis and subsequent release of glucocorticoid hormones is a key response that mobilizes fuel stores to meet energy demands1,2,3,4,5. Despite the importance of the HPA axis response, the neural mechanisms that drive its activation during energy deficit are unknown. Here, we show that fasting-activated hypothalamic agouti-related peptide (AgRP)-expressing neurons trigger and are essential for fasting-induced HPA axis activation. AgRP neurons do so through projections to the paraventricular hypothalamus (PVH), where, in a mechanism not previously described for AgRP neurons, they presynaptically inhibit the terminals of tonically active GABAergic afferents from the bed nucleus of the stria terminalis (BNST) that otherwise restrain activity of corticotrophin-releasing hormone (CRH)-expressing neurons. This disinhibition of PVHCrh neurons requires γ-aminobutyric acid (GABA)/GABA-B receptor signalling and potently activates the HPA axis. Notably, stimulation of the HPA axis by AgRP neurons is independent of their induction of hunger, showing that these canonical ‘hunger neurons’ drive many distinctly different adaptations to the fasted state. Together, our findings identify the neural basis for fasting-induced HPA axis activation and uncover a unique means by which AgRP neurons activate downstream neurons: through presynaptic inhibition of GABAergic afferents. Given the potency of this disinhibition of tonically active BNST afferents, other activators of the HPA axis, such as psychological stress, may also work by reducing BNST inhibitory tone onto PVHCrh neurons.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Fibre photometry data reported in this study are available from: https://research.bidmc.harvard.edu/datashare/DataShareInfo.ASP?Submit=Display&ID=9. Source data are provided with this paper.
Code availability
The custom analysis code used in this study is available on GitHub (https://github.com/AMDouglass/Douglass_Resch_Madara_etal).
References
Ahima, R. S. et al. Role of leptin in the neuroendocrine response to fasting. Nature 382, 250–252 (1996).
Dallman, M. F. et al. Starvation: early signals, sensors, and sequelae. Endocrinology 140, 4015–4023 (1999).
Perry, R. J. et al. Leptin mediates a glucose–fatty acid cycle to maintain glucose homeostasis in starvation. Cell 172, 234–236.e17 (2018).
Muglia, L., Jacobson, L., Dikkest, P. & Majzoub, J. A. Corticotropin-releasing hormone deficiency reveals major fetal but not adult glucocorticoid need. Nature 373, 427–432 (1995).
Steinhauser, M. L. et al. The circulating metabolome of human starvation. JCI Insight 3, e121434 (2018).
Long, C. N. H., Katzin, B. & Fry, E. G. The adrenal cortex and carbohydrate metabolism. Endocrinology 26, 309–344 (1940).
Chihaoui, M. et al. The risk for hypoglycemia during Ramadan fasting in patients with adrenal insufficiency. Nutrition 45, 99–103 (2018).
Exton, J. Regulation of gluconeogenesis by glucocorticoids. Monogr Endocrinol 12, 535–546 (1979).
Kuo, T., McQueen, A., Chen, T. C. & Wang, J. C. Regulation of glucose homeostasis by glucocorticoids. Adv. Exp. Med. Biol. 872, 99 (2015).
Goldberg, A. L., Tischler, M. & DeMartino G, G. G. Hormonal regulation of protein degradation and synthesis in skeletal muscle. Fed. Proc. 39, 31–36 (1980).
Djurhuus, C. B. et al. Effects of cortisol on lipolysis and regional interstitial glycerol levels in humans. Am. J. Physiol. Endocrinol. Metab. 283, E172–E177 (2002).
Aponte, Y., Atasoy, D. & Sternson, S. M. AGRP neurons are sufficient to orchestrate feeding behavior rapidly and without training. Nat. Neurosci. 14, 351–355 (2011).
Krashes, M. J. et al. Rapid, reversible activation of AgRP neurons drives feeding behavior in mice. J. Clin. Invest. 121, 1424–1428 (2011).
Livneh, Y. et al. Homeostatic circuits selectively gate food cue responses in insular cortex. Nature 546, 611–616 (2017).
Fernandes, A. C. A. et al. Arcuate AgRP, but not POMC neurons, modulate paraventricular CRF synthesis and release in response to fasting. Cell Biosci. 12, 118 (2022).
Spencer, R. L. & Deak, T. A users guide to HPA axis research. Physiol. Behav. 178, 43–65 (2017).
Betley, J. N., Cao, Z. F. H., Ritola, K. D. & Sternson, S. M. Parallel, redundant circuit organization for homeostatic control of feeding behavior. Cell 155, 1337–1350 (2013).
Betley, J. N. et al. Neurons for hunger and thirst transmit a negative-valence teaching signal. Nature 521, 180–185 (2015).
Garfield, A. S. et al. A neural basis for melanocortin-4 receptor–regulated appetite. Nat. Neurosci. 18, 863–871 (2015).
Mahn, M. et al. Efficient optogenetic silencing of neurotransmitter release with a mosquito rhodopsin. Neuron 109, 1621–1635.e8 (2021).
Ziegler, D. R., Cullinan, W. E. & Herman, J. P. Distribution of vesicular glutamate transporter mRNA in rat hypothalamus. J. Comp. Neurol. 448, 217–229 (2002).
Cowley, M. A. et al. Integration of npy, agrp, and melanocortin signals in the hypothalamic paraventricular nucleus: evidence of a cellular basis for the adipostat. Neuron 24, 155–163 (1999).
Pronchuk, N., Beck-Sickinger, A. G. & Colmers, W. F. Multiple NPY receptors inhibit GABAA synaptic responses of rat medial parvocellular effector neurons in the hypothalamic paraventricular nucleus. Endocrinology 143, 535–543 (2002).
Mackay, J. P. et al. NPY2 receptors reduce tonic action potential-independent gabab currents in the basolateral amygdala. J. Neurosci. 39, 4909–4930 (2019).
Colmers, P. L. W. & Bains, J. S. Balancing tonic and phasic inhibition in hypothalamic corticotropin-releasing hormone neurons. J. Physiol. 596, 1919–1929 (2018).
Krashes, M. J., Shah, B. P., Koda, S. & Lowell, B. B. Rapid versus delayed stimulation of feeding by the endogenously released AgRP neuron mediators GABA, NPY, and AgRP. Cell Metab. 18, 588–595 (2013).
Wahlestedt, C. et al. Neuropeptide Y (NPY) in the area of the hypothalamic paraventricular nucleus activates the pituitary–adrenocortical axis in the rat. Brain Res. 417, 33–38 (1987).
Johnson, C. S., Bains, J. S. & Watts, A. G. Neurotransmitter diversity in pre-synaptic terminals located in the parvicellular neuroendocrine paraventricular nucleus of the rat and mouse hypothalamus. J. Comp. Neurol. 526, 1287–1306 (2018).
Cole, R. L. & Sawchenko, P. E. Neurotransmitter regulation of cellular activation and neuropeptide gene expression in the paraventricular nucleus of the hypothalamus. J. Neurosci. 22, 959–969 (2002).
Roland, B. L. & Sawchenko, P. E. Local origins of some GABAergic projections to the paraventricular and supraoptic nuclei of the hypothalamus in the rat. J. Comp. Neurol. 332, 123–143 (1993).
Cullinan, W. E., Ziegler, D. R. & Herman, J. P. Functional role of local GABAergic influences on the HPA axis. Brain Struct. Funct. 213, 63–72 (2008).
Johnson, S. B. et al. A basal forebrain site coordinates the modulation of endocrine and behavioral stress responses via divergent neural pathways. J. Neurosci. 36, 8687–8699 (2016).
Radley, J. J., Gosselink, K. L. & Sawchenko, P. E. A discrete GABAergic relay mediates medial prefrontal cortical inhibition of the neuroendocrine stress response. J. Neurosci. 29, 7330–7340 (2009).
Michel, M. C. et al. XVI. International Union of Pharmacology recommendations for the nomenclature of neuropeptide Y, peptide YY, and pancreatic polypeptide receptors. Pharmacol. Rev. 50, 143–150 (1998).
Woodward, C. J. H., Hervey, G. R., Oakey, R. E. & Whitaker, E. M. The effects of fasting on plasma corticosterone kinetics in rats. Br. J. Nutr. 66, 117–127 (1991).
Chen, Y. et al. Sustained NPY signaling enables AgRP neurons to drive feeding. eLife 8, e46348 (2019).
Wahlestedt, C., Hakanson, R., Vaz, C. A. & Zukowska-Grojec, Z. Norepinephrine and neuropeptide Y: vasoconstrictor cooperation in vivo and in vitro. Am. J. Physiol. Integr. Comp. Physiol. 258, R736–R742 (1990).
Khan, A. M. et al. MAP kinases couple hindbrain-derived catecholamine signals to hypothalamic adrenocortical control mechanisms during glycemia-related challenges. J. Neurosci. 31, 18479–18491 (2011).
Atasoy, D., Betley, J. N., Su, H. H. & Sternson, S. M. Deconstruction of a neural circuit for hunger. Nature 488, 172–177 (2012).
Padilla, S. L. et al. Agouti-related peptide neural circuits mediate adaptive behaviors in the starved state. Nat. Neurosci. 19, 734–741 (2016).
Li, M. M. et al. The paraventricular hypothalamus regulates satiety and prevents obesity via two genetically distinct circuits. Neuron 102, 653–667.e6 (2019).
Baur, R. & Sigel, E. On high- and low-affinity agonist sites in GABAA receptors. J. Neurochem. 87, 325–332 (2003).
Hill, D. R. & Bowery, N. G. 3H-baclofen and 3H-GABA bind to bicuculline-insensitive GABAB sites in rat brain. Nature 290, 149–152 (1981).
Xu, S. et al. Behavioral state coding by molecularly defined paraventricular hypothalamic cell type ensembles. Science 370, eabb2494 (2020).
Atasoy, D. et al. A genetically specified connectomics approach applied to long-range feeding regulatory circuits. Nat. Neurosci. 17, 1830–1839 (2014).
Perry, R. J. et al. Leptin’s hunger-suppressing effects are mediated by the hypothalamic–pituitary–adrenocortical axis in rodents. Proc. Natl Acad. Sci. USA. 116, 13670–13679 (2019).
Tong, Q., Ye, C. P., Jones, J. E., Elmquist, J. K. & Lowell, B. B. Synaptic release of GABA by AgRP neurons is required for normal regulation of energy balance. Nat. Neurosci. 11, 998–1000 (2008).
Krashes, M. J. et al. An excitatory paraventricular nucleus to AgRP neuron circuit that drives hunger. Nature 507, 238–242 (2014).
Fenselau, H. et al. A rapidly acting glutamatergic ARC→PVH satiety circuit postsynaptically regulated by α-MSH. Nat. Neurosci. 20, 42–51 (2016).
Vong, L. et al. Leptin action on GABAergic neurons prevents obesity and reduces inhibitory tone to POMC neurons. Neuron 71, 142–154 (2011).
Madisen, L. et al. A toolbox of Cre-dependent optogenetic transgenic mice for light-induced activation and silencing. Nat. Neurosci. 15, 793–802 (2012).
Plummer, N. W. et al. Expanding the power of recombinase-based labeling to uncover cellular diversity. Dev. 142, 4385–4393 (2015).
Erickson, C., Clegg, K. E. & Palmiter, R. D. Sensitivity to leptin and susceptibility to seizures of mice lacking neuropeptide Y. Nature 381, 415–418 (1996).
Harms, K. J., Tovar, K. R. & Craig, A. M. Synapse-specific regulation of AMPA receptor subunit composition by activity. J. Neurosci. 25, 6379–6388 (2005).
Chen, T. W. et al. Ultrasensitive fluorescent proteins for imaging neuronal activity. Nature 499, 295–300 (2013).
Atasoy, D., Aponte, Y., Su, H. H. & Sternson, S. M. A FLEX switch targets channelrhodopsin-2 to multiple cell types for imaging and long-range circuit mapping. J. Neurosci. 28, 7025–7030 (2008).
Akam, T. & Walton, M. E. pyPhotometry: open source Python based hardware and software for fiber photometry data acquisition. Sci Rep. 9, 3521 (2019).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587.e29 (2021).
Acknowledgements
We thank M. Andermann, J. Majzoub and the Lowell laboratory for helpful discussions; J. Yu, C. Wu and Y. Li for technical assistance; S. Liberles for providing the NPY-KO mice; and the Brigham Assay Core for running the ACTH radioimmunoassay. The confocal imaging was performed at the BIDMC Confocal Imaging Core. This work was supported by the National Institutes of Health (NIH) (P30DK046200, R01DK122976, R01DK075632, R01DK089044, R01DK111401 and R01DK096010 to B.B.L. and R00HL144923 to J.M.R.). A.M.D. was supported by the Charles A. King Trust Postdoctoral Research Fellowship Program. H.K. was supported by a Brain & Behavior Research Foundation (BBRF) Young Investigator Grant.
Author information
Authors and Affiliations
Contributions
A.M.D., J.M.R. and B.B.L. conceived the study and designed experiments. J.C.M. performed and analysed slice electrophysiology experiments. A.M.D. and H.K. performed and analysed fibre photometry experiments. O.Y. provided the AAV-eOPN3 and guidance on its use. A.G. and M.Y. made the AAV-synaptophysin-GCaMP6f. A.M.D. and J.M.R. performed and analysed all other experiments and prepared the figures. Z.Y. assisted with stereotaxic surgeries. A.M.D., J.M.R. and B.B.L. wrote the manuscript with input from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks William Colmers, Alan Watts and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Fasting activation of the HPA axis is regulated by leptin but not by the arcuate → PVH melanocortin system.
a, Scheme of the HPA axis. b, Timeline for leptin injection. c, Corticosterone levels were elevated in mice fasted for 24 h compared to fed mice (n = 8 per group. Two-tailed unpaired t-test- ***P = 0.0006). d, Intraperitoneal administration of leptin reduced corticosterone levels in fasted mice (n = 8 vehicle, n = 9 leptin. Two-tailed unpaired t-test- *P = 0.0130). e, Schematic of optogenetic stimulation of AgRP neurons. f, One hour food intake by Agrp-IRES-Cre mice expressing ChR2 or GFP in the arcuate (n = 4 AgRPGFP, n = 6 AgRPChR2. Two-way ANOVA; Sidak’s multiple comparisons test. AgRPGFP ON vs. AgRPChR2 ON- ***P = 0.0008; AgRPChR2 OFF vs. AgRPChR2 ON- ***P = 0.0001). g, Schematic of Cre-dependent AAV-hM4Di-mCherry stereotaxic injection into the PVH of Mc4r-t2A-Cre mice. h, Three hour food intake by Mc4r-t2A-Cre mice expressing hM4Di or mCherry in the PVH (n = 9 MC4RmCherry, n = 6 MC4RhM4Di. Two-way ANOVA; Sidak’s multiple comparisons test. MC4RmCherry CNO vs. MC4RhM4Di CNO- ****P = <0.0001; MC4RhM4Di Vehicle vs. MC4RhM4Di CNO- **P = 0.0015). i, Timeline for chemogenetic inhibition of PVHMc4r neurons. j, Inhibition of PVHMc4r neurons did not affect corticosterone in fed mice (n = 9 MC4RmCherry, n = 6 MC4RhM4Di. Two-tailed unpaired t-test- P = 0.5933). k, Schematic of Cre-dependent AAV-hM3 Dq-mCherry stereotaxic injection into arcuate nucleus of Pomc-IRES-Cre mice. l, Timeline for chemogenetic activation of POMC neurons. m, Activation of POMC neurons did not affect corticosterone in fasted mice (n = 9 POMCmCherry, n = 7 POMChM3Dq. Two-tailed unpaired t-test- P = 0.6736). Scale bar = 200 µm. Data represent mean ± sem.
Extended Data Fig. 2 AgRP neuron collaterals and characterization of GABAergic afferents to PVHCrh neurons.
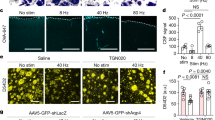
a, Schematic of collateral mapping via injection of retro-AAV-FlpO into Agrp-IRES-Cre::FLTG mice. b, AgRP neuron terminals detected upon injection of retro-AAV-FlpO into individual AgRP neuron efferent sites (denoted by row headings) in Agrp-IRES-Cre::FLTG mice. a.c: anterior commissure, 3V: third ventricle, a.q: aqueduct, s.c.p: superior cerebellar peduncle. c, Schematic of rabies-based collateral mapping. EnvA pseudotyped rabies-∆G-GFP was injected into AgRP neuron terminal areas (PVH example) in Agrp-IRES-Cre mice that were previously injected with AAV-FLEX-TVA-mCherry in the arcuate. d, AgRP neuron terminals detected upon injection of EnvA pseudotyped rabies-∆G-GFP into individual AgRP neuron efferent sites (denoted by row headings) in Agrp-IRES-Cre mice. e, Schematic of monosynaptic rabies mapping from PVHCrh neurons. f-j, Representative images of starter cells and putative afferents containing rabies-GFP labeled neurons in the PVH (f), BNST and mPOA (g), PV (h), LH and anterior arcuate nucleus (i) and DMH and posterior arcuate nucleus (j). PVH: paraventricular hypothalamus, a.c: anterior commissure, BNST: bed nucleus of the stria terminalis, mPOA: medial preoptic area, 3V: Third ventricle, PV: periventricular hypothalamus, LH: lateral hypothalamus, Arc: arcuate nucleus, DMH: dorsomedial hypothalamus. k, Schematic of recordings of BNST→PVH terminals. These mice are those used in Fig. 5j–o. l, BNST→PVH axons reduced activity in response to a looming hand motion stimulus over the animal (n = 3 mice). m, Expression of NPYRs in major PVH neuron cell types derived from single-cell RNA sequencing of PVH neurons43, plotted as average normalized expression level and fraction of cells in each cluster that express the receptor. Scale bar = 200 µm. Data represent mean ± sem.
Extended Data Fig. 3 Schematic of AAV spread in Agrp-IRES-Cre mice, AAV spread and fiber placements in Agrp-IRES-Cre::Crh-IRES-Cre mice and fiber placements in Agrp-IRES-Cre::LSL-ChR2-eYFP mice.
a, Schematics representing AAV spread (shaded regions) for every animal related to experiments in Fig. 1a–c. Each animal is represented by a different colour. b, Representative image of hM3Dq-mCherry expression in AgRP neuron somas in the arcuate nucleus. c, Schematics representing AAV spread (shaded regions) for every animal related to experiments in Fig. 1a,d–e. Each animal is represented by a different colour. d, Representative image of hM4Di-mCherry expression in AgRP neuron somas in the arcuate nucleus. e, Schematics representing AAV spread (shaded regions) and fiber placement (rectangles) for every animal related to experiments in Fig. 1f–i. Each animal is represented by a different colour. See Fig. 1g for a representative histological image. f, Schematics representing AAV spread (shaded regions) and fiber placement (rectangles) for every animal related to experiments in Fig. 1f,l–m. Each animal is represented by a different colour. g, Representative images of GCaMP6s expression in PVHCrh neurons, optic fiber placement in the PVH and hM4Di-mCherry expression in AgRP neuron somas in the arcuate nucleus. h, Schematics representing fiber placements (rectangles) for every animal related to experiments in Fig. 1n–q and Extended Data Fig. 1e,f. Each animal is represented by a different colour. i, Representative image of ChR2-eYFP expression in AgRP neuron somas in the arcuate nucleus and fiber placement. Scale bar = 200 µm. 3V = Third ventricle. The schematics were created using The Mouse Brain in Stereotaxic Coordinates Second Edition (Paxinos and Franklin).
Extended Data Fig. 4 Schematic of AAV spread and fiber placements in Agrp-IRES-Cre mice.
a, Schematics representing AAV spread (shaded regions) and fiber placements (rectangles) for every animal related to experiments in Fig. 2a–c. Each animal is represented by a different colour. b, Representative images of ChR2-mCherry expression in AgRP neuron somas in the arcuate nucleus and fiber placement in the BNST. c, Schematics representing AAV spread (shaded regions) and fiber placements (rectangles) for every animal related to experiments in Fig. 2a–c. Each animal is represented by a different colour. d, Representative images of ChR2-mCherry expression in AgRP neuron somas in the arcuate nucleus and fiber placement in the PVH. Scale bar = 200 µm. 3V = Third ventricle. a.c. = anterior commisure. The schematics were created using The Mouse Brain in Stereotaxic Coordinates Second Edition (Paxinos and Franklin).
Extended Data Fig. 5 Schematic of AAV spread and fiber placements in Agrp-IRES-Cre mice.
a, Schematics representing AAV spread (shaded regions) and fiber placements (rectangles) for every animal related to experiments in Fig. 2a–c. Each animal is represented by a different colour. b, Representative image of ChR2-mCherry expression in AgRP neuron somas in the arcuate nucleus and fiber placement in the LH. Scale bar = 500 µm. c, Schematics representing AAV spread (shaded regions) and fiber placements (rectangles) for every animal related to experiments in Fig. 2a–c. Each animal is represented by a different colour. d, Representative images of ChR2-mCherry expression in AgRP neuron somas in the arcuate nucleus and fiber placement in the PBN. Left scale bar = 200 µm, Right scale bar = 500 µm. 3V = Third ventricle. The schematics were created using The Mouse Brain in Stereotaxic Coordinates Second Edition (Paxinos and Franklin).
Extended Data Fig. 6 Schematic of AAV spread and fiber placements in Agrp-IRES-Cre mice.
a, Schematics representing AAV spread (shaded regions) and fiber placements (rectangles) for every animal related to experiments in Fig. 2f–h. Each animal is represented by a different colour. b, Representative images of eOpn3-mScarlet expression in AgRP neuron somas in the arcuate nucleus and fiber placement in the BNST. c, Schematics representing AAV spread (shaded regions) and fiber placements (rectangles) for every animal related to experiments in Fig. 2f–h. Each animal is represented by a different colour. d, Representative images of eOpn3-mScarlet expression in AgRP neuron somas in the arcuate nucleus and fiber placement in the PVH. e, Schematics representing AAV spread (shaded regions) and fiber placements (rectangles) for every animal related to experiments in Fig. 2f–h. Each animal is represented by a different colour. f, Representative images of eOpn3-mScarlet expression in AgRP neuron somas in the arcuate nucleus and fiber placement in the LH. Scale bar = 200 µm. 3V = Third ventricle. a.c. = anterior commisure. The schematics were created using The Mouse Brain in Stereotaxic Coordinates Second Edition (Paxinos and Franklin).
Extended Data Fig. 7 Schematic of AAV spread and fiber placements in Agrp-IRES-Cre::NPY-KO, Agrp-IRES-Cre::vGatlox/lox and Agrp-IRES-Cre::NPY-KO::vGatlox/lox mice.
a, Schematics representing AAV spread (shaded regions) and fiber placements (rectangles) for every animal related to experiments in Fig. 4a–c. Each animal is represented by a different colour. b–d, Representative images of ChR2-mCherry expression in AgRP neuron somas and fiber placement in the arcuate nucleus. Scale bar = 200 µm. 3V = Third ventricle. The schematics were created using The Mouse Brain in Stereotaxic Coordinates Second Edition (Paxinos and Franklin).
Extended Data Fig. 8 Schematic of infusion cannula placements in WT mice.
a, Schematics representing infusion cannula placements (rectangles) for every animal related to experiments in Fig. 4d–f. Each animal is represented by a different colour. b–d, Representative images of cannula placement above the PVH. Scale bar = 200 µm. The schematics were created using The Mouse Brain in Stereotaxic Coordinates Second Edition (Paxinos and Franklin).
Extended Data Fig. 9 Schematic of AAV spread and fiber placements in vGAT-IRES-Cre mice.
a, Schematics representing AAV spread (shaded regions) for every animal related to experiments in Fig. 5c–e. Each animal is represented by a different colour. b, Representative image of hM4Di-mCherry expression in BNSTvGAT neuron somas. c, Schematics representing AAV spread (shaded regions) for every animal related to experiments in Fig. 5c–e. Each animal is represented by a different colour. d, Representative image of hM4Di-mCherry expression in LHvGAT neuron somas. e, Schematics representing AAV spread (shaded regions) and fiber placements (rectangles) for every animal related to experiments in Fig. 5f–i. Each animal is represented by a different colour. See Fig. 5g for a representative histological image. Scale bar = 200 µm. a.c. = anterior commisure, i.c. = internal capsule. The schematics were created using The Mouse Brain in Stereotaxic Coordinates Second Edition (Paxinos and Franklin).
Extended Data Fig. 10 Schematic of AAV spread and fiber placements in Agrp-IRES-Cre mice, Mc4r-IRES-Cre and Pomc-IRES-Cre mice.
a, Schematics representing AAV spread (shaded regions) and fiber placements (rectangles) for every animal related to experiments in Fig. 5j–n and Extended Data Fig. 2k,l. Each animal is represented by a different colour. See Fig. 5k for a representative histological image. b, Schematics representing AAV spread (shaded regions) for every animal related to experiments in Extended Data Fig. 1g-j. Each animal is represented by a different colour. c, Representative image of hM4Di-mCherry expression in PVHMc4r neuron somas. d, Schematics representing AAV spread (shaded regions) for every animal related to experiments in Extended Data Fig. 1k-m. Each animal is represented by a different colour. e, Representative image of hM3Dq-mCherry expression in POMC neuron somas in the arcuate nucleus. Scale bar = 200 µm. 3V = Third ventricle. The schematics were created using The Mouse Brain in Stereotaxic Coordinates Second Edition (Paxinos and Franklin).
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Douglass, A.M., Resch, J.M., Madara, J.C. et al. Neural basis for fasting activation of the hypothalamic–pituitary–adrenal axis. Nature 620, 154–162 (2023). https://doi.org/10.1038/s41586-023-06358-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-023-06358-0
This article is cited by
-
Reciprocal activity of AgRP and POMC neurons governs coordinated control of feeding and metabolism
Nature Metabolism (2024)
-
Effects of lifestyle factors on leukocytes in cardiovascular health and disease
Nature Reviews Cardiology (2024)
-
Gut microbiota connects the brain and the heart: potential mechanisms and clinical implications
Psychopharmacology (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.