Abstract
Ecological interactions are one of the main forces that sustain Earth’s biodiversity. A major challenge for studies of ecology and evolution is to determine how these interactions affect the fitness of species when we expand from studying isolated, pairwise interactions to include networks of interacting species1,2,3,4. In networks, chains of effects caused by a range of species have an indirect effect on other species they do not interact with directly, potentially affecting the fitness outcomes of a variety of ecological interactions (such as mutualism)5,6,7. Here we apply analytical techniques and numerical simulations to 186 empirical mutualistic networks and show how both direct and indirect effects alter the fitness of species coevolving in these networks. Although the fitness of species usually increased with the number of mutualistic partners, most of the fitness variation across species was driven by indirect effects. We found that these indirect effects prevent coevolving species from adapting to their mutualistic partners and to other sources of selection pressure in the environment, thereby decreasing their fitness. Such decreases are distributed in a predictable way within networks: peripheral species receive more indirect effects and experience higher reductions in fitness than central species. This topological effect was also evident when we analysed an empirical study of an invasion of pollination networks by honeybees. As honeybees became integrated as a central species within networks, they increased the contribution of indirect effects on several other species, reducing their fitness. Our study shows how and why indirect effects can govern the adaptive landscape of species-rich mutualistic assemblages.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The dataset of empirical networks used in this study is available in a GitHub repository (https://github.com/lgcosmo/Cosmo_et_el_indirect_effects_fitness), in Zenodo (https://doi.org/10.5281/zenodo.7945239) and in the Web of Life database (www.web-of-life.es).
Code availability
All the code to perform the numerical simulations used in this study is available in a GitHub repository (https://github.com/lgcosmo/Cosmo_et_el_indirect_effects_fitness) and in Zenodo (https://doi.org/10.5281/zenodo.7945239).
References
Thompson, J. N. The coevolving web of life. Am. Nat. 173, 125–140 (2009).
Thompson, J. N. Mutualistic webs of species. Science 312, 372–373 (2006).
Bascompte, J. & Jordano, P. Plant–animal mutualistic networks: the architecture of biodiversity. Annu. Rev. Ecol. Evol. Syst. 38, 567–593 (2007).
Bascompte, J. Disentangling the web of life. Science 325, 416–419 (2009).
Guimarães, P. R. Jr, Pires, M. M., Jordano, P., Bascompte, J. & Thompson, J. N. Indirect effects drive coevolution in mutualistic networks. Nature 550, 511–514 (2017).
Wootton, J. T. The nature and consequences of indirect effects in ecological communities. Annu. Rev. Ecol. Syst. 25, 443–466 (1994).
Strauss, S. Y. Indirect effects in community ecology: their definition, study and importance. Trends Ecol. Evol. 6, 206–210 (1991).
Darwin, C. On the Origin of Species by Means of Natural Selection, or the Preservation of Favoured Races in the Struggle for Life (Murray, 1861).
Fisher, R. A. The Genetical Theory of Natural Selection (Clarendon Press, 1930).
Reed, D. H. Relationship between population size and fitness. Conserv. Biol. 19, 563–568 (2005).
Orr, H. A. Fitness and its role in evolutionary genetics. Nat. Rev. Genet. 10, 531–539 (2009).
Thompson, J. N. Relentless Evolution (Univ. Chicago Press, 2013).
Bronstein, J. L. Mutualism (Oxford Univ. Press, 2015).
Bronstein, J. L. Our current understanding of mutualism. Q. Rev. Biol. 69, 31–51 (1994).
Bronstein, J. L., Dieckmann, U. & Ferrière, R. in Evolutionary Conservation Biology (eds Ferrière, R. et al.) 305–326 (Cambridge Univ. Press, 2009).
Thompson, J. N. The Geographic Mosaic of Coevolution (Univ. Chicago Press, 2005).
Nuismer, S. L., Jordano, P. & Bascompte, J. Coevolution and the architecture of mutualistic networks. Evolution 67, 338–354 (2013).
Guimarães, P. R. Jr, Jordano, P. & Thompson, J. N. Evolution and coevolution in mutualistic networks. Ecol. Lett. 14, 877–885 (2011).
Kauffman, S. A. & Johnsen, S. Coevolution to the edge of chaos: coupled fitness landscapes, poised states, and coevolutionary avalanches. J. Theor. Biol. 149, 467–505 (1991).
Lande, R. Natural selection and random genetic drift in phenotypic evolution. Evolution 30, 314–334 (1976).
Santamaría, L. & Rodríguez-Gironés, M. A. Linkage rules for plant–pollinator networks: trait complementarity or exploitation barriers? PLoS Biol. 5, e31 (2007).
Rohr, R. P., Naisbit, R. E., Mazza, C. & Bersier, L.-F. Matching–centrality decomposition and the forecasting of new links in networks. Proc. R. Soc. B 283, 20152702 (2016).
Peralta, G. et al. Trait matching and phenological overlap increase the spatio-temporal stability and functionality of plant–pollinator interactions. Ecol. Lett. 23, 1107–1116 (2020).
Galen, C. High and dry: drought stress, sex-allocation trade-offs, and selection on flower size in the alpine wildflower Polemonium viscosum (Polemoniaceae). Am. Nat. 156, 72–83 (2000).
Irwin, R. E., Adler, L. S. & Brody, A. K. The dual role of floral traits: pollinator attraction and plant defense. Ecology 85, 1503–1511 (2004).
Strauss, S. Y. & Whittall, J. B. in Ecology and Evolution of Flowers (eds Harder, L. D. & Barrett, S. C. H.) 120–138 (Oxford Univ. Press, 2006).
Galetti, M. et al. Functional extinction of birds drives rapid evolutionary changes in seed size. Science 340, 1086–1090 (2013).
Yoshimura, J. & Jansen, V. A. A. Evolution and population dynamics in stochastic environments. Res. Popul. Ecol. 38, 165–182 (1996).
Bascompte, J., Possingham, H. & Roughgarden, J. Patchy populations in stochastic environments: critical number of patches for persistence. Am. Nat. 159, 128–137 (2002).
Waser, N. M., Chittka, L., Price, M. V., Williams, N. M. & Ollerton, J. Generalization in pollination systems, and why it matters. Ecology 77, 1043–1060 (1996).
Levins, R. The effect of random variations of different types on population growth. Proc. Natl Acad. Sci. USA 62, 1061–1065 (1969).
Piculell, B. J., Hoeksema, J. D. & Thompson, J. N. Interactions of biotic and abiotic environmental factors in an ectomycorrhizal symbiosis, and the potential for selection mosaics. BMC Biol. 6, 23 (2008).
Batstone, R. T., Peters, M. A. E., Simonsen, A. K., Stinchcombe, J. R. & Frederickson, M. E. Environmental variation impacts trait expression and selection in the legume–rhizobium symbiosis. Am. J. Bot. 107, 195–208 (2020).
Caruso, C. M., Eisen, K. E., Martin, R. A. & Sletvold, N. A meta-analysis of the agents of selection on floral traits. Evolution 73, 4–14 (2019).
Munguía-Rosas, M. A., Ollerton, J., Parra-Tabla, V. & De-Nova, J. A. Meta-analysis of phenotypic selection on flowering phenology suggests that early flowering plants are favoured. Ecol. Lett. 14, 511–521 (2011).
Fricke, E. C. & Svenning, J.-C. Accelerating homogenization of the global plant–frugivore meta-network. Nature 585, 74–78 (2020).
Mackin, C. R., Peña, J. F., Blanco, M. A., Balfour, N. J. & Castellanos, M. C. Rapid evolution of a floral trait following acquisition of novel pollinators. J. Ecol. 109, 2234–2246 (2021).
Stuart, Y. E. et al. Rapid evolution of a native species following invasion by a congener. Science 346, 463–466 (2014).
Valido, A., Rodríguez-Rodríguez, M. C. & Jordano, P. Honeybees disrupt the structure and functionality of plant–pollinator networks. Sci. Rep. 9, 4711 (2019).
Aizen, M. A., Morales, C. L. & Morales, J. M. Invasive mutualists erode native pollination webs. PLoS Biol. 6, e31 (2008).
Traveset, A. & Richardson, D. M. Mutualistic interactions and biological invasions. Annu. Rev. Ecol. Evol. Syst. 45, 89–113 (2014).
Traveset, A. & Richardson, D. M. Biological invasions as disruptors of plant reproductive mutualisms. Trends Ecol. Evol. 21, 208–216 (2006).
Merrel, D. J. The Adaptive Seascape: The Mechanism of Evolution (Univ. Minnesota Press, 1994).
Nuismer, S. Introduction to Coevolutionary Theory (W. H. Freeman, 2017).
Birskis-Barros, I., Freitas, A. V. L. & Guimarães, P. R. Jr. Habitat generalist species constrain the diversity of mimicry rings in heterogeneous habitats. Sci. Rep. 11, 5072 (2021).
Bezanson, J., Edelman, A., Karpinski, S. & Shah, V. B. Julia: a fresh approach to numerical computing. SIAM Rev. 59, 65–98 (2017).
R Core Team. R: A Language and Environment for Statistical Computing. https://www.R-project.org/ (R Foundation for Statistical Computing, 2022).
Cosmo, L. G. Indirect effects shape species fitness in coevolved mutualistic networks (version 1.0.0). https://doi.org/10.5281/zenodo.7945239 (2023).
Acknowledgements
This study was financed in part by the Coordenação de Aperfeiçoamento de Pessoal de Nível Superior – Brasil (CAPES), finance code 001. L.G.C. is funded by a São Paulo Research Foundation PhD scholarship (FAPESP; grants 2019/22146-3 and 2022/07939-0). P.R.G. is funded by Brazil’s Council for Scientific and Technological Development (CNPq; grant 307134/2017-2), FAPESP (grant 2018/14809-0) and the Royal Society (CHL/R1/180156). A.P.A.A. was supported by CAPES, finance code 001, and FAPESP (grant 2016/14277-2). M.M.P. is funded by FAPESP (grant 2019/25478-7) and CNPq (grant 313059/2022-5). M.A.M.A. is funded by FAPESP (grant 2021/14335-0-ICTP‐SAIFR), and by CNPq (grant 301082/2019‐7). A.V. is supported by the Spanish Ministry of Science, Innovation, and Universities (PGC2018-099772-B-100). P.J. is funded by the Spanish Ministry of Science, Innovation, and Universities (CGL2017-82847-P), LifeWatch ERIC-SUMHAL (LIFEWATCH-2019-09-CSIC-13)/FEDER-EU funding), and the VI and VII Research Funding Program from the Universidad de Sevilla (2021/00000826). J.B. is supported by the Swiss National Science Foundation (grant 310030_197201). We thank R. Cogni, M. A. R. Mello, C. R. Montoya and M. C. Vidal for comments on previous versions of this manuscript.
Author information
Authors and Affiliations
Contributions
All authors designed the study. L.G.C., P.R.G. and M.A.M.A. derived the fitness function. L.G.C. and P.R.G. developed the analytical approximations. L.G.C. performed simulations and conducted analyses. A.V. and P.J. contributed to data acquisition and analysis of the field experiment. L.G.C., P.R.G. and J.B. wrote the first draft of the manuscript, and all authors contributed substantially to the final draft.
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing financial interests.
Peer review
Peer review information
Nature thanks Thilo Gross and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Species traits and fitness quickly reach equilibrium values after coevolving in mutualistic networks.
a–c, Example for an ant-plant mutualistic network (panel a) of how species traits (panel b) and fitness (panel c) quickly reach a coevolutionary equilibrium. d, The coevolutionary equilibrium is reached even if not all species survive throughout the dynamics, as illustrated by three species that were randomly extinct from the network (for illustrative purposes, species whose trait values reach zero). Each point and line correspond to the values for each species in the network (represented by different colors). The diamond-shaped points on the right of panel b represent the environmental optima of each species \(({\theta }_{i}\)). The dashed lines in panel d represent the trait values at equilibrium predicted by equation (12) using the matrix of the interactions among surviving species. Parameter values are as follows: \({\sigma }_{G{z}_{i}}^{2}=1.0\), \({\varrho }_{i}=0.2\), \(\alpha =0.2\), \({m}_{i}=0.5\). Initial trait values and environmental optima were sampled from a uniform distribution U [0, 10].
Extended Data Fig. 2 Coevolution in mutualistic networks increases the variability in species fitness when a certain percentage of the species with the lowest fitness become extinct, and the surviving species coevolve to a new equilibrium.
Each set of panels represents a specific scenario where a certain percentage of the species in the network experience extinction after reaching the initial coevolutionary equilibrium. In all scenarios extinctions occurred in a specific order, starting with the species possessing the lowest fitness until a desired percentage of extinctions was reached. The corresponding extinction percentages for each scenario are as follows: a–b, 10%; c–d, 20%; e–f, 30%; g–h, 40%; and i–j, 50%. In all panels the red histogram bars depict the distribution of fitness of the surviving species in the new coevolutionary equilibrium for 10³ numerical simulations parameterized with the initial structure of empirical networks (n = 186 empirical networks). Green histogram bars correspond to the scenario in which species coevolve as isolated pairs and there are no extinctions. In the boxplots each point corresponds to the mean value for 10³ numerical simulations for a given species coevolving in the empirical mutualistic networks (n = 186 empirical networks). Fitness values are rescaled relative to the average of the scenario in which species coevolve in networks or as isolated pairs. Other parameter values are as follows: \({\sigma }_{G{z}_{i}}^{2}=1.0\), \({\varrho }_{i}\) = 0.2, \(\alpha =0.2\), and \({m}_{i}=0.5\). \({\theta }_{i}\) and initial trait values were sampled from a uniform distribution U [0, 10].
Extended Data Fig. 3 Coevolution in mutualistic networks increases the variability in species fitness for different levels of strength of mutualistic selection.
a, Histogram showing the distribution of mean equilibrium fitness of species for 10³ numerical simulations of a pair of coevolving species (green histogram bars), or of species within the 186 empirical networks used to parameterize the model (red histogram bars), for different values of \({m}_{i}\) (values above each panel). b, Boxplot showing how species fitness vary with the number of mutualistic partners for different values of \({m}_{i}\) (the intensity of mutualistic selection, values above each panel). Each point corresponds to the mean value for 10³ numerical simulations for a given species. In all panels fitness values are rescaled relative to the average of each scenario and \({m}_{i}\) (coevolution in pairs or in networks). Other parameter values are as follows: \({\sigma }_{G{z}_{i}}^{2}=1.0\), \({\varrho }_{i}\) = 0.2, \(\alpha =0.2\), and \({m}_{i}\) as indicated on top of each panel. \({\theta }_{i}\) and initial trait values were sampled from a uniform distribution U [0, 10].
Extended Data Fig. 4 Indirect effects drive species fitness for different parameterizations of the model.
a, Examples of how indirect evolutionary effects drive the fitness of species in numerical simulations across all empirical networks (n=186 empirical networks) for different values of \({m}_{i}\) (values above each panel), for species with five mutualistic partners. b, Examples of how indirect evolutionary effects drive the fitness of species in numerical simulations across all empirical networks (n = 186 empirical networks) for different intervals of \({\theta }_{i}\) (values above each panel), and sensitivity of species adaptive landscapes (\({\varrho }_{i}\), diferente colors) for species with five mutualistic partners. Points in all panels represent average results for 10³ numerical simulations of each combination of empirical network and parameter values. Other parameter values are as follows: \({\sigma }_{G{z}_{i}}^{2}=1.0\) and \(\alpha =0.2\). Values of \({m}_{i}\) and \({\varrho }_{i}\) as indicated on each panel. In a, \({\theta }_{i}\) and initial trait values were sampled from a uniform distribution U [0, 10], while in b the upper bound of the uniform distribution is indicated in the values above each panel.
Extended Data Fig. 5 Peripheral species are more affected by indirect effects drive for different networks and levels of mutualistic selection.
Results from numerical simulations parameterized with the structure of empirical networks (n = 186 empirical networks), showing how the contribution of indirect evolutionary effects is smaller for core than peripheral species within the same network. This result holds for all values of \({m}_{i}\), the intensity of mutualistic selection (values above each panel). Each point corresponds to the average for 10³ numerical simulations for each combination of species position (core or peripheral), empirical network and \({m}_{i}\). Points of different colors correspond to species that were classified either as core species (red points) or peripheral species (blue points). Parameter values are as follows: \({m}_{i}={variable}\), \({\sigma }_{G{z}_{i}}^{2}=1.0\), \({\varrho }_{i}\) = 0.2, \(\alpha =0.2\). \({\theta }_{i}\) and initial trait values were sampled from a uniform statistical distribution U [0, 10].
Extended Data Fig. 6 Indirect effects drive the fitness of surviving species when the least fit species become extinct, and the surviving ones coevolve to a new equilibrium.
Each panel corresponds to scenarios in which a certain percentage of the species in the network underwent extinction after reaching a first coevolutionary equilibrium. For all scenarios extinctions occurred in a specific order, starting with the species possessing the lowest fitness, until a given percentage of extinctions was reached. The corresponding percentage of species extinct are as follows: a, scenario without extinctions; b, 10%; c, 20%; d, 30%; e, 40%; and f, 50%. Points in each panel represent average results for species with three mutualistic partners across 10³ numerical simulations parameterized with the initial structure of 186 empirical networks. In panels b–f, indirect evolutionary effects were computed from the matrix of evolutionary effects (Q-matrix) among the surviving species (equation 14). Parameter values are as follows: \({m}_{i}=0.5\), \({\sigma }_{G{z}_{i}}^{2}=1.0\), \({\varrho }_{i}\) = 0.2, \(\alpha =0.2\). \({\theta }_{i}\) and initial trait values were sampled from a uniform statistical distribution U [0, 10].
Extended Data Fig. 7 Invasion of a network by a supergeneralist changes the fitness of native species via coevolution for different levels of mutualistic selection.
Histograms showing the average change in native species fitness (n = 10³ numerical simulations for each of the 73 empirical networks) after coevolving with the invasive species for different values of \({m}_{i}\) (the intensity of mutualistic selection, values above each panel). The frequency in the y-axis represents log(Counts). Other parameter values are as follows: \({\sigma }_{G{z}_{i}}^{2}=1.0\), \({\varrho }_{i}\) = 0.2, \(\alpha =0.2\), and \({m}_{i}\) as indicated on top of each panel. \({\theta }_{i}\) and initial trait values were sampled from a uniform distribution U [0, 10].
Extended Data Fig. 8 Direct and indirect evolutionary effects drive the change in fitness of native species directly interacting with a supergeneralist invader.
Relationship between the average change in species fitness (n = 10³ numerical simulations for each of the 73 empirical networks) after the invasion and the change in the contribution of indirect evolutionary effects for direct partners of A. mellifera and for different values of \({m}_{i}\) (the intensity of mutualistic selection, values above each panel). Parameter values are as follows: \({\sigma }_{G{z}_{i}}^{2}=1.0\), \({\varrho }_{i}\) = 0.2, \(\alpha =0.2\), and \({m}_{i}\) as indicated on top of each panel. \({\theta }_{i}\) and initial trait values were sampled from a uniform distribution U [0, 10].
Extended Data Fig. 9 Indirect evolutionary effects drive the change in fitness of native species only indirectly interacting with a supergeneralist invader.
Relationship between the average change in species fitness (n = 103 numerical simulations for each of the 73 empirical networks) after the invasion and the change in the contribution of indirect evolutionary effects for indirect partners of A. mellifera and for different values of \({m}_{i}\) (the intensity of mutualistic selection, values above each panel). Other parameter values are as follows: \({\sigma }_{G{z}_{i}}^{2}=1.0\), \({\varrho }_{i}\) = 0.2, \(\alpha =0.2\), and \({m}_{i}\) as indicated on top of each panel. \({\theta }_{i}\) and initial trait values were sampled from a uniform distribution U [0, 10].
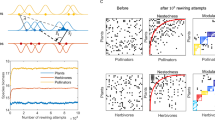
Extended Data Fig. 10 Indirect evolutionary effects and rewiring of interactions shape the fitness consequences of the invasion of a network by the supergeneralist A. mellifera.
a–b, Representations of the (a) pre- (before beekeeping activity) and (b) post-Apis (after beekeeping activity) network structures, showing how the invasion by A. mellifera (in red) reorganizes interactions. c–d, Histograms showing (c) the change in the number of partners and (d) the change in fitness that native species experienced after coevolving with A. mellifera. e, Relationship between the change in indirect evolutionary effects caused by A. mellifera and the change in the fitness of native species. The results in panels d and e correspond to the average results for the native species of 10³ numerical simulations of the coevolutionary dynamics in the pre- and post-Apis networks. Parameter values are as follows: \({m}_{i}=0.5\), \({\sigma }_{G{z}_{i}}^{2}=1.0\), \({\varrho }_{i}\) = 0.2, \(\alpha =0.2\). \({\theta }_{i}\) and initial trait values were sampled from a uniform statistical distribution U [0, 10].
Supplementary information
Supplementary Methods
The analytical study of the coevolutionary model, derivation of the fitness function, analytical approximations, sensitivity analysis and additional analysis using empirical networks.
Supplementary Table 1
The references of the empirical networks used to perform numerical simulations.
Supplementary Table 2
The references of the subset of the empirical networks used to perform numerical simulations of the Apis mellifera invasion scenario.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Cosmo, L.G., Assis, A.P.A., de Aguiar, M.A.M. et al. Indirect effects shape species fitness in coevolved mutualistic networks. Nature 619, 788–792 (2023). https://doi.org/10.1038/s41586-023-06319-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-023-06319-7
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.