Abstract
Parasitic nematodes are a major threat to global food security, particularly as the world amasses 10 billion people amid limited arable land1,2,3,4. Most traditional nematicides have been banned owing to poor nematode selectivity, leaving farmers with inadequate means of pest control4,5,6,7,8,9,10,11,12. Here we use the model nematode Caenorhabditis elegans to identify a family of selective imidazothiazole nematicides, called selectivins, that undergo cytochrome-p450-mediated bioactivation in nematodes. At low parts-per-million concentrations, selectivins perform comparably well with commercial nematicides to control root infection by Meloidogyne incognita, a highly destructive plant-parasitic nematode. Tests against numerous phylogenetically diverse non-target systems demonstrate that selectivins are more nematode-selective than most marketed nematicides. Selectivins are first-in-class bioactivated nematode controls that provide efficacy and nematode selectivity.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Code availability
The Python and R code used to generate the dose–response heatmaps and the HPLC chromatogram heatmaps have been published72.
References
Tilman, D., Balzer, C., Hill, J. & Befort, B. L. Global food demand and the sustainable intensification of agriculture. Proc. Natl Acad. Sci. USA 108, 20260–20264 (2011).
Hunter, M. C., Smith, R. G., Schipanski, M. E., Atwood, L. W. & Mortensen, D. A. Agriculture in 2050: recalibrating targets for sustainable intensification. Bioscience 67, 386–391 (2017).
Cunningham, S. A. et al. To close the yield-gap while saving biodiversity will require multiple locally relevant strategies. Agric. Ecosyst. Environ. 173, 20–27 (2013).
Popp, J., Petö, K. & Nagy, J. Pesticide productivity and food security. A review. Agron. Sustain. Dev. 33, 243–255 (2013).
Oerke, E.-C. Crop losses to pests. J. Agric. Sci. 144, 31–43 (2006).
Singh, S. K., Hodda, M. & Ash, G. J. Plant-parasitic nematodes of potential phytosanitary importance, their main hosts and reported yield losses. EPPO Bull. 43, 334–374 (2013).
Abad, P. et al. Genome sequence of the metazoan plant-parasitic nematode Meloidogyne incognita. Nat. Biotechnol. 26, 909–915 (2008).
Jones, R. K. in Nematology in South Africa: A View from the 21st Century (eds Fourie, H. et al.) 129–150 (Springer, 2017).
Desaeger, J., Wram, C. & Zasada, I. New reduced-risk agricultural nematicides—rationale and review. J. Nematol. 52, e2020-911 (2020).
EU Pesticides Database, https://ec.europa.eu/food/plant/pesticides/eu-pesticides-database/start/screen/active-substances (European Commission).
NPIC Product Research Online (NPRO), http://npic.orst.edu/NPRO/# (National Pesticide Information Center, accessed 25 July 2022).
Jordan, S., Nischwitz, C., Ramirez, R. & Gordillo, L. F. Managing the spread of alfalfa stem nematodes (Ditylenchus dipsaci): the relationship between crop rotation periods and pest reemergence. Nat. Resour. Model. 30, e12083 (2017).
d’Errico, G., Giacometti, R., Roversi, P. F., d’Errico, F. P. & Woo, S. L. Mode of action and efficacy of iprodione against the root-knot nematode Meloidogyne incognita. Ann. Appl. Biol. 171, 506–510 (2017).
Burns, A. R. et al. Caenorhabditis elegans is a useful model for anthelmintic discovery. Nat. Commun. 6, 7485 (2015).
Culetto, E. et al. The Caenorhabditis elegans unc-63 gene encodes a levamisole-sensitive nicotinic acetylcholine receptor α subunit. J. Biol. Chem. 279, 42476–42483 (2004).
Hu, Y., Xiao, S. H. & Aroian, R. V. The new anthelmintic tribendimidine is an l-type (levamisole and pyrantel) nicotinic acetylcholine receptor agonist. PLoS Negl. Trop. Dis. 3, e499 (2009).
Driscoll, M., Dean, E., Reilly, E., Bergholz, E. & Chalfie, M. Genetic and molecular analysis of a Caenorhabditis elegans β-tubulin that conveys benzimidazole sensitivity. J. Cell Biol. 109, 2993–3003 (1989).
Dent, J. A., Smith, M. M., Vassilatis, D. K. & Avery, L. The genetics of ivermectin resistance in Caenorhabditis elegans. Proc. Natl Acad. Sci. USA 97, 2674–2679 (2000).
Kaminsky, R. et al. A new class of anthelmintics effective against drug-resistant nematodes. Nature 452, 176–180 (2008).
Guest, M. et al. The calcium-activated potassium channel, SLO-1, is required for the action of the novel cyclo-octadepsipeptide anthelmintic, emodepside, in Caenorhabditis elegans. Int. J. Parasitol. 37, 1577–1588 (2007).
Kawasaki, I., Jeong, M. H., Oh, B. K. & Shim, Y. H. Apigenin inhibits larval growth of Caenorhabditis elegans through DAF-16 activation. FEBS Lett. 584, 3587–3591 (2010).
Rand, J. B. & Russell, R. L. Choline acetyltransferase-deficient mutants of the nematode Caenorhabditis elegans. Genetics 106, 227–248 (1984).
Hu, Y., Platzer, E. G., Bellier, A. & Aroian, R. V. Discovery of a highly synergistic anthelmintic combination that shows mutual hypersusceptibility. Proc. Natl. Acad. Sci. USA 107, 5955–5960 (2010).
Burns, A. R. et al. A predictive model for drug bioaccumulation and bioactivity in Caenorhabditis elegans. Nat. Chem. Biol. 6, 549–557 (2010).
Ortiz de Montellano, P. R. Cytochrome P450-activated prodrugs. Future Med. Chem. 5, 213–228 (2013).
Leung, M. C. K., Goldstone, J. V., Boyd, W. A., Freedman, J. H. & Meyer, J. N. Caenorhabditis elegans generates biologically relevant levels of genotoxic metabolites from aflatoxin B1 but not benzo[a]pyrene in vivo. Toxicol. Sci. 118, 444–453 (2010).
Harlow, P. H., Perry, S. J., Stevens, A. J. & Flemming, A. J. Comparative metabolism of xenobiotic chemicals by cytochrome P450s in the nematode Caenorhabditis elegans. Sci Rep. 8, 13333 (2018).
Porter, T. D. New insights into the role of cytochrome P450 reductase (POR) in microsomal redox biology. Acta Pharm. Sin. B 2, 102–106 (2012).
Kalgutkar, A. S. et al. Reactive metabolite trapping studies on imidazo- and 2-methylimidazo[2,1-b]thiazole-based inverse agonists of the ghrelin receptor. Drug Metab. Dispos. 7, 1375–1388 (2013).
Ryan, E. et al. Evidence for the in vitro bioactivation of aminopyrazole derivatives: trapping reactive aminopyrazole intermediates using glutathione ethyl ester in human liver microsomes. Chem. Res. Toxicol. 28, 1747–1752 (2015).
Cooper, A. J. L. & Hanigan, M. H. Metabolism of glutathione S-conjugates: multiple pathways. Compr. Toxicol. https://doi.org/10.1016/B978-0-12-801238-3.01973-5 (2018).
Ferguson, G. D. & Bridge, W. J. The glutathione system and the related thiol network in Caenorhabditis elegans. Redox Biol. 24, 101171 (2019).
Blum, R., Meyer, K. C., Wünschmann, J., Lendzian, K. J. & Grill, E. Cytosolic action of phytochelatin synthase. Plant Physiol. 153, 159–169 (2010).
Essig, Y. J., Webb, S. M. & Stürzenbaum, S. R. Deletion of phytochelatin synthase modulates the metal accumulation pattern of cadmium exposed C. elegans. Int. J. Mol. Sci. 17, 257 (2016).
Giustarini, D., Milzani, A., Dalle-Donne, I., Tsikas, D. & Rossi, R. N-Acetylcysteine ethyl ester (NACET): a novel lipophilic cell-permeable cysteine derivative with an unusual pharmacokinetic feature and remarkable antioxidant potential. Biochem. Pharmacol. 84, 1522–1533 (2012).
Trudgill, D. L. & Blok, V. C. Apomictic, polyphagus root-knot nematodes: exceptionally successful and damaging biotrophic root pathogens. Annu. Rev. Phytopathol. 39, 53–77 (2001).
Oka, Y. From old-generation to next-generation nematicides. Agronomy 10, 1387 (2020).
Devguard 500SC Label,https://za.uplonline.com/download_links/awyXPTiWs8OT3dGCErYdw3nnA3nLnwxpCpUn7jzb.pdf (deVGen/UPL).
Asif, M., Rehman, B., Parihar, K., Ganai, M. A. & Siddiqui, M. A. Effect of various physico-chemical factors on the incidence of root knot nematode Meloidogyne spp. infesting tomato in District Aligarh (Uttar Pradesh) India. J. Plant Sci. 10, 234–243 (2015).
Barker, K. R., Schmitt, D. P. & Imbriani, J. L. in An Advanced Treatise on Meloidogyne Volume II: Methodology (eds Barker, K. R. et al.) 135–148 (North Carolina State University Department of Plant Pathology and the United States Agency for International Development, 1985).
Marion, M. J., Hantz, O. & Durantel, D. in Hepatocytes: Methods and Protocols (ed. Maurel, P.) 261–272 (Humana Press, 2010).
Rocher, F. et al. Salicylic acid transport in Ricinus communis involves a pH-dependent carrier system in addition to diffusion. Plant Physiol. 150, 2081–2091 (2009).
Wram, C. L., Hesse, C. N. & Zasada, I. A. Transcriptional response of Meloidogyne incognita to non-fumigant nematicides. Sci Rep. 12, 9814 (2022).
Dara, S. K. The new integrated pest management paradigm for the modern age. J. Integr. Pest. Manag. 10, 12 (2019).
Cardarelli, M., Woo, S. L., Rouphael, Y. & Colla, G. Seed treatments with microorganisms can have a biostimulant effect by influencing germination and seedling growth of crops. Plants 11, 259 (2022).
Migunova, V. D. & Sasanelli, N. Bacteria as biocontrol tool against phytoparasitic nematodes. Plants 10, 389 (2021).
Slomczynska, U. et al. in Discovery and Synthesis of Crop Protection Products (eds. Maienfisch, P. & Stevenson, T.) 129–147 (American Chemical Society, 2015).
Jensen, J. P., Kalwa, U., Pandey, S. & Tylka, G. L. Avicta and clariva affect the biology of the soybean cyst nematode, Heterodera glycines. Plant Dis. 102, 2480–2486 (2018).
Ahmed, S., Zhou, Z., Zhou, J. & Chen, S.-Q. Pharmacogenomics of drug metabolizing enzymes and transporters: relevance to precision medicine. Genomics Proteomics Bioinformatics 14, 298–313 (2016).
Nixon, S. A. et al. Where are all the anthelmintics? Challenges and opportunities on the path to new anthelmintics. Int. J. Parasitol. Drugs Drug Resist. 14, 8–16 (2020).
Lewis, J. A. & Fleming, J. T. in Caenorhabditis elegans: Modern Biological Analysis of an Organism (eds Epstein, H. F. & Shakes, D. C.) 3–29 (Academic Press, 1995).
Burns, A. R. et al. High-throughput screening of small molecules for bioactivity and target identification in Caenorhabditis elegans. Nat. Protoc. 1, 1906–1914 (2006).
Demeler, J., Kuttler, U. & von Samson-Himmelstjerna, G. Veterinary parasitology adaptation and evaluation of three different in vitro tests for the detection of resistance to anthelmintics in gastro intestinal nematodes of cattle. Vet. Parasitol. 170, 61–70 (2010).
Bartley, D. J. et al. A survey of anthelmintic resistant nematode parasites in Scottish sheep flocks. Vet. Parasitol. 117, 61–71 (2003).
Jindapunnapat, K., Reetz, N. D., Macdonald, M. H., Bhagavathy, G. & Meyer, S. L. F. Activity of vetiver extracts and essential oil against Meloidogyne incognita. J. Nematol. 50, 147–162 (2018).
Wram, C. L. & Zasada, I. A. Short-term effects of sublethal doses of nematicides on Meloidogyne incognita. Phytopathology 109, 1605–1613 (2019).
Meyer, S. L. F., Chauhan, K. R. & MacDonald, M. H. Evaluation of roselle (Hibiscus sabdariffa) leaf and pomegranate (Punica granatum) fruit rind for activity against Meloidogyne incognita. Nematropica 46, 85–96 (2016).
Meyer, S. L. F. et al. Plantago lanceolata and Plantago rugelii extracts are toxic to Meloidogyne incognita but not to certain microbes. J. Nematol. 38, 333–338 (2006).
Sherman, F. Getting started with yeast. Methods Enzymol. 350, 3–41 (2002).
Howe, K. L. et al. WormBase ParaSite—a comprehensive resource for helminth genomics. Mol. Biochem. Parasitol. 215, 2–10 (2016).
Blanc-Mathieu, R. et al. Hybridization and polyploidy enable genomic plasticity without sex in the most devastating plant-parasitic nematodes. PLoS Genet. 13, e1006777 (2017).
Sievers, F. et al. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol. Syst. Biol. 7, 539 (2011).
WormBase release WS283, http://www.wormbase.org (Alliance of Genome Resources, accessed 16 December 2021).
Wram, C. L. & Zasada, I. Differential response of Meloidogyne, Pratylenchus, Globodera, and Xiphinema species to the nematicide fluazaindolizine. Phytopathology. 110, 2003–2009 (2020).
Chundawat, T. S., Kumari, P., Sharma, N. & Bhagat, S. Strategic synthesis and in vitro antimicrobial evaluation of novel difluoromethylated 1-(1,3-diphenyl-1H-pyrazol-4-yl)-3,3-difluoro-1,3-dihydro-indol-2-ones. Med. Chem. Res. 25, 2335–2348 (2016).
Pyl, T., Giebelmann, R. & Beyer, H. Über bicyclische heterocyclen mit gemeinsamem stickstoffatom, I. Zur kenntnis der imidazo[2.1-b]thiazole. Justus Liebigs Ann. Chem. 643, 145–153 (1961).
Huang, G. et al. Engineering broad root-knot resistance in transgenic plants by RNAi silencing of a conserved and essential root-knot nematode parasitism gene. Proc. Natl Acad. Sci. USA 103, 14302–14306 (2006).
Mota, F. C. et al. New sources of resistance to Meloidogyne incognita race 3 in wild cotton accessions and histological characterization of the defence mechanisms. Plant Pathol. 62, 1173–1183 (2013).
Basso, M. F. et al. MiDaf16-like and MiSkn1-like gene families are reliable targets to develop biotechnological tools for the control and management of Meloidogyne incognita. Sci Rep. 10, 6991 (2020).
Hajihassani, A., Rutter, W. B. & Luo, X. Resistant pepper carrying N, Me1, and Me3 have different effects on penetration and reproduction of four major Meloidogyne species. J. Nematol. 51, 1–9 (2019).
de Souza, J. D. A. et al. Knocking-down Meloidogyne incognita proteases by plant-delivered dsRNA has negative pleiotropic effect on nematode vigor. PLoS ONE 8, e85364 (2013).
Burns, A. R. and Roy, P. J. Python and R scripts for the generation of dose–response heatmaps and heatmaps of HPLC chromatograms. Zenodo https://doi.org/10.5281/zenodo.7731172 (2023).
Acknowledgements
Free-living nematode strains were provided by the CGC (University of Minnesota), M.-A. Félix (Institute of Biology of the Ecole Normale Supérieure, Paris, France) and R. Rae (Liverpool John Moores University, Liverpool, UK). M. hapla was provided by B. Mimee at Agriculture and Agri-food Canada in Saint-Jean-sur-Richelieu, Quebec, Canada. MS analyses were performed by staff at the Advanced Instrumentation for Molecular Structure Mass Spectrometry Laboratory in the Department of Chemistry at the University of Toronto, Canada. NMR analyses were performed by staff at the CSICOMP NMR Facility in the Department of Chemistry at the University of Toronto, Canada. Many thanks to D. Colwell and D. Gray at the Lethbridge and Agri-Food Canada Research Centre for supplying faeces from infected calves for C. oncophora egg collection; K. Yoshioka at the University of Toronto, Canada, for P. simiae WSC417; J. Goldstone from the Woods Hole Oceanographic Institution for providing the M. incognita cyp gene names; B. Ciruna for the gift of NACET; G. Brown for the culture of S. cerevisiae; B. Derry for helpful comments on the manuscript; and N. Robbins from L. Cowen’s laboratory for comments on the work. The following grants supported this work: CIHR grants (313296 and 173448), an NSERC i2i grant (555963-20), a David Dime Grant, and a Canada Research Chair (Tier 1) to P.J.R.; a NSERC grant (RGPIN-2020-04168) to M.L.; a CIHR Foundation grant (FDN-154288) and a Canada Research Chair (Tier 1) to L.E.C.; an EvoFunPath fellowship from the NSERC CREATE program (555337-2021) to E.P.; a CIHR project grant (303157) to I.S. C.A.M.F. is supported by the Alberta Innovates Technology Fund (G2016000681). J.S.G. is supported by a NSERC discovery grant (RGPIN-2021-02489), and by the NSERC-CREATE Host–Parasite Interactions Program graduate training programme at the University of Calgary.
Author information
Authors and Affiliations
Contributions
Unless otherwise noted, all experiments were carried out in the laboratory of P.J.R. All free-living nematode dose–response experiments were carried out by A.R.B. A.R.B. performed the HPLC-based metabolism experiments and analysed all related MS data. A.R.B. performed the NACET experiments. B.M.P., E.M.R. and A.R.B. performed the C. oncophora dose–response experiments in J.S.G.’s laboratory. S. cerevisiae work was done by A.R.B., J.K. and B.C. M. incognita bioinformatics was done by J.K. B.C. performed the yeast-based M. incognita CYP expression screen, and data were analysed by A.R.B. J.S. performed the HEK293 cell dose–response assays in I.S.’s laboratory. J.T. (in the laboratory of H.M.K.) and J.R.V. (in the laboratory of J.J.D.) performed the zebrafish dose–response assays. S.M. and S.W.C. performed the HepaRG cell dose–response experiments in S.A.M.’s laboratory. E.P. performed the C. albicans assays in L.E.C.’s laboratory. In the laboratory of S.R.C., A.S.V. performed the Arabidopsis greening assays. A.R.B performed the P. simiae dose–response experiments. C.A.M.F. and Q.X. carried out the mouse experiments. A.R.B. performed the metabolite purification for NMR analyses. R.J.B. carried out the selectivin syntheses and characterization in M.L.’s laboratory, and helped with metabolite NMR data analyses. A.R.B. performed the C. elegans cyp RNAi screen. M.H.M. carried out the in vitro M. incognita assays in S.L.F.M.’s laboratory. M.K. carried out the in vitro M. chitwoodi assays and the M. incognita soil-based reproduction assays in tomato plants in the laboratory of I.Z. A.R.B. and J.K. carried out the M. hapla assays. D. melanogaster experiments were carried out by C.H. and A.R.B. in the laboratories of P.J.R. and H.M.K. The genetics screens for resistant mutants were performed by A.R.B. The project was conceived by A.R.B. and P.J.R. The manuscript was written by A.R.B. and P.J.R.
Corresponding authors
Ethics declarations
Competing interests
L.E.C. is a co-founder and shareholder in Bright Angel Therapeutics, a platform company for development of new antifungal therapeutics. L.E.C. is a consultant for Boragen, a small-molecule development company focused on leveraging the specific chemical properties of boron chemistry for crop protection and animal health. L.E.C. is a Science Advisor for Kapoose Creek, a company that harnesses the therapeutic potential of fungi. Mention of trade names or commercial products in this publication is solely for the purpose of providing specific information and does not imply recommendation or endorsement by the US Department of Agriculture. USDA is an equal opportunity provider and employer. The University of Toronto is pursuing patent protection related to the use of selectivions as nematacides for which two national phase applications, US Patent Application 17/624,629, and Canadian Patent Application 3,146,085, are currently under prosecution and list A.R.B., M.L., R.B. and P.J.R. as inventors. A patent application, listing A.R.B., J.K., B.C., and P.J.R. as inventors, is under preparation for the use of yeast-based expression systems for the identification of bioactivated nematacides, insecticides, pesticides and drug leads. The patent application will be soon filed listing the inventors and the University of Toronto as applicants.
Peer review
Peer review information
Nature thanks Johan Desaeger, Timothy Geary, Fredd Vergara and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 The selectivin-A imidazole-thiol metabolite is not nematicidal.
a, Mass spectrometry (MS) data for the M4 metabolite HPLC fraction and untreated control fraction, as well as MS/MS data for the 418.99 mass. The proposed M4 disulfide structure is shown, along with the protonated imidazole-thiol monomer structure resulting from fragmentation. b, A proposed metabolic pathway producing the imidazole-thiol metabolite (ITM) which oxidizes to form M4/ITM disulfide (ITM-DiS). Imidazothiazole ring opening is likely CYP-dependent. c, Biomass-normalized and background-corrected HPLC chromatograms for lysates of selectivin-A-treated L1 worms lysed ± iodoacetamide (IAA). Peaks corresponding to selectivin-A (sel-A), metabolites M1-M4, and alkylated ITM (ITM-alk) are indicated. d, MS data for the ITM-alk HPLC fraction and untreated control fraction. e, HPLC-DAD quantification of selectivin-A, M1-M4, and ITM-alk in the lysates of selectivin-A-treated L1 worms lysed ± IAA (n = 3 biological replicates with 200,000 worms per replicate). Area under the curve (AUC) for metabolites M1-M3 and ITM-alk was calculated at 260 nm, and at 300 nm for M4. Data are presented as mean ± SEM. For each analyte, unpaired one-tailed Student’s t-tests were performed comparing the means of the ± IAA conditions. P-values are shown. f, Dose-response for C. elegans L1s treated with selectivins or commercially-sourced ITM. g, Biomass-normalized and background-corrected HPLC chromatograms for lysates of L1 worms incubated in commercially-sourced ITM, lysed ± IAA. The top chromatogram is the ITM standard injected directly onto the column, and is not normalized or corrected. ITM oxidizes to ITM-DiS prior to HPLC analysis. ITM-DiS and ITM-alk peaks are indicated. h, HPLC-DAD quantification of M4/ITM-DiS and ITM-alk in the lysates of 100 µM selectivin-A-treated or 100 µM ITM-treated L1 worms, lysed ± IAA (n = 3 biological replicates with 200,000 worms per replicate). AUC was calculated at 260 nm. Data are presented as mean ± SEM.
Extended Data Fig. 2 NMR spectra for HPLC-purified selectivin-A metabolite M2.
a, 1H NMR spectra. The structure of M2 with all carbon and heteroatom assignments is given. The top number in each multiplet box corresponds to the carbon that the proton is connected to. 1H NMR (700 MHz, cd3od) δ 8.02 (s, 1H), 7.82 – 7.78 (m, 2H), 7.42 – 7.37 (m, 2H), 5.98 (dd, J = 7.9, 2.2 Hz, 1H), 4.74 (dd, J = 7.6, 5.3 Hz, 1H), 4.66 (dd, J = 14.9, 8.0 Hz, 1H), 3.86 (s, 2H), 3.81 (dd, J = 14.9, 2.2 Hz, 1H), 3.62 (t, J = 6.2 Hz, 1H), 3.23 (dd, J = 14.2, 5.3 Hz, 1H), 3.08 (dd, J = 14.1, 7.6 Hz, 1H), 2.54 (ddd, J = 15.9, 7.9, 6.6 Hz, 1H), 2.48 (dt, J = 15.8, 6.6 Hz, 1H), 2.16 – 2.05 (m, 2H), 1.97 (s, 3H). b, 1H-1H COSY NMR spectra. c, 1H-13C HSQC NMR spectra. d, 1H-13C HMBC NMR spectra.
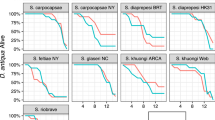
Extended Data Fig. 3 Selectivin bioactivation is nematode-specific.
a and b, Biomass-normalized and background-corrected HPLC chromatograms of lysates (a) and incubation buffers (b) from C. elegans (Ce), C. briggsae (Cb), P. pacificus (Pp), Rhabditophanes sp. KR3021 (KR), M. incognita (Mi), D. melanogaster (Dm), and D. rerio (Dr) larvae, as well as S. cerevisiae (Sc) cells, incubated in 100 µM selectivin-A. Selectivin-A and metabolite peaks are indicated. c, Total detectable production of M1 + M2 + M3 in lysate and buffer for each of the seven species tested. d, M2 tissue accumulation in each species tested. e, Unmodified selectivin-A tissue accumulation in each of the test species. AUC is area under the curve at 260 nm. Data are presented as mean ± SEM. For c and d, unpaired one-tailed Student’s t-tests were performed for all pairwise comparisons of means. The means sharing a letter are statistically indistinguishable (p > 0.01). The p-values for each comparison in c and d can be found in the Statistical Information subsection of the Methods. For e, unpaired one-tailed Student’s t-tests were performed relative to the C. elegans mean. P-values are shown. For c-e, n = 4 biological replicates for Ce, Cb, Pp, and Dr, and n = 3 biological replicates for KR, Mi, Sc, and Dm.
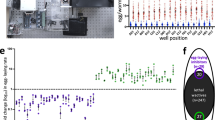
Extended Data Fig. 4 Selectivin treatment generally increases the root weights of tomato plants challenged with nematode infection.
The effects of 11 selectivin analogs and 4 commercial nematicides on the root weights of tomato plants challenged with M. incognita infection is shown. Root weight values are percent of untreated control. For each analog, aqueous molar concentration, parts-per-million soil concentration, and kilograms-per-hectare values are shown. The R-groups for each selectivin analog are indicated (see Fig. 1a for the selectivin core scaffold and R-group positions). Data are presented as mean ± SEM.
Extended Data Fig. 5 Free-living nematodes likely detoxify selectivin-E via hydroxylation and subsequent phosphoglucosidation.
a, Structure of selectivin-E (sel-E) with exact mass. b-c, Raw HPLC chromatograms for lysates of C. elegans and P. pacificus L1 worms incubated in 100 µM selectivin-E or DMSO alone (untreated). The top chromatogram is the selectivin-E standard injected directly onto the column. The unmodified selectivin-E parent compound and selectivin-E metabolites E-M1, E-M2, and E-M3 are indicated. c, Mass spectrometry (MS) data for the E-M1, E-M2, and E-M3 HPLC fractions and corresponding untreated control fractions. Raw MS counts are shown for the most abundant masses identified in the mass spectra that are not detectable in the corresponding untreated control fractions. Raw counts were taken from centroid plots of the raw mass spectra. Counts below 1 × 103 were considered not detectable (nd). Based on the presumptive masses of the three metabolites we predict that E-M1 is a hydroxylated metabolite of selectivin-E, E-M2 is an O-linked glucoside of selectivin-E, and E-M3 is an O-linked phosphoglucoside. Molecular formulas are indicated for the predicted metabolite structures, as well as exact masses, P. pacificus experimental accurate masses, and the ppm difference between the two. A ppm difference of less than 5 ppm suggests that the molecular formulae are correct. Ce, C. elegans; Pp, P. pacificus.
Supplementary information
Supplementary Table 1
Accurate mass and exact mass values for the selectivin-A metabolites.
Supplementary Table 2
C. elegans cyp RNAi screen data.
Supplementary Table 3
Identity matrix for the 31 M. incognita CYP protein sequences Identified from the v3 genome.
Supplementary Table 4
C. elegans CYP-35C1, C. elegans EMB-8 and M. incognita CYP cDNA sequences codon-optimized for yeast expression.
Supplementary Table 5
Yeast-based M. incognita cyp expression screen data.
Supplementary Table 6
M. incognita gene read counts and expression ranks obtained from RNA sequencing analysis from ref. 43.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Burns, A.R., Baker, R.J., Kitner, M. et al. Selective control of parasitic nematodes using bioactivated nematicides. Nature 618, 102–109 (2023). https://doi.org/10.1038/s41586-023-06105-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-023-06105-5
This article is cited by
-
Screening of indigenous entomopathogenic fungal isolates on plant parasitic nematodes in China
European Journal of Plant Pathology (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.