Abstract
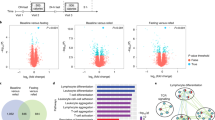
Decades of work have elucidated cytokine signalling and transcriptional pathways that control T cell differentiation and have led the way to targeted biologic therapies that are effective in a range of autoimmune, allergic and inflammatory diseases. Recent evidence indicates that obesity and metabolic disease can also influence the immune system1,2,3,4,5,6,7, although the mechanisms and effects on immunotherapy outcomes remain largely unknown. Here, using two models of atopic dermatitis, we show that lean and obese mice mount markedly different immune responses. Obesity converted the classical type 2 T helper (TH2)-predominant disease associated with atopic dermatitis to a more severe disease with prominent TH17 inflammation. We also observed divergent responses to biologic therapies targeting TH2 cytokines, which robustly protected lean mice but exacerbated disease in obese mice. Single-cell RNA sequencing coupled with genome-wide binding analyses revealed decreased activity of nuclear receptor peroxisome proliferator-activated receptor-γ (PPARγ) in TH2 cells from obese mice relative to lean mice. Conditional ablation of PPARγ in T cells revealed that PPARγ is required to focus the in vivo TH response towards a TH2-predominant state and prevent aberrant non-TH2 inflammation. Treatment of obese mice with a small-molecule PPARγ agonist limited development of TH17 pathology and unlocked therapeutic responsiveness to targeted anti-TH2 biologic therapies. These studies reveal the effects of obesity on immunological disease and suggest a precision medicine approach to target the immune dysregulation caused by obesity.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
The bulk RNA-sequencing and ChIP–seq data can be accessed in the NCBI Sequence Read Archive under the accession PRJNA553761. Additionally, the ChIP–seq data that support the findings of this study have been deposited in the Gene Expression Omnibus (GEO) under the accession code GSE189216. The single cell-RNA sequencing data used in this study are deposited in the GEO under the accession code GSE189476. Other relevant data are available from the corresponding authors upon reasonable request. Source data are provided with this paper.
Code availability
Code used in this manuscript can be provided by reasonable request to the corresponding authors.
References
Buck, M. D., Sowell, R. T., Kaech, S. M. & Pearce, E. L. Metabolic instruction of immunity. Cell 169, 570–586 (2017).
Man, K., Kutyavin, V. I. & Chawla, A. Tissue immunometabolism: development, physiology, and pathobiology. Cell Metab. 25, 11–26 (2017).
Geltink, R. I. K., Kyle, R. L. & Pearce, E. L. Unraveling the complex interplay between T cell metabolism and function. Annu. Rev. Immunol. 36, 461–488 (2018).
Michelet, X. et al. Metabolic reprogramming of natural killer cells in obesity limits antitumor responses. Nat. Immunol. 19, 1330–1640 (2018).
Nicholas, D. A. et al. Fatty acid metabolites combine with reduced β-oxidation to activate Th17 inflammation in human type 2 diabetes. Cell Metab. 30, 447–461.e5 (2019).
Ringel, A. E. et al. Obesity shapes metabolism in the tumor microenvironment to suppress anti-tumor immunity. Cell 183, 1848–1866.e26 (2020).
Zhang, C. et al. STAT3 activation-induced fatty acid oxidation in CD8+ T effector cells is critical for obesity-promoted breast tumor growth. Cell Metab. 31, 148–161.e5 (2020).
Hersoug, L. G. & Linneberg, A. The link between the epidemics of obesity and allergic diseases: does obesity induce decreased immune tolerance? Allergy 62, 1205–1213 (2007).
Wenzel, S. E. Asthma phenotypes: the evolution from clinical to molecular approaches. Nat. Med. 18, 716–725 (2012).
Zhang, A. & Silverberg, J. I. Association of atopic dermatitis with being overweight and obese: a systematic review and metaanalysis. J. Am. Acad. Dermatol. 72, 606–616.e4 (2015).
Fahy, J. V. Type 2 inflammation in asthma—present in most, absent in many. Nat. Rev. Immunol. 15, 57–65 (2015).
Bieber, T. Atopic Dermatitis. New Engl. J. Med. 358, 1483–1494 (2008).
Eckert, L. et al. The burden of atopic dermatitis in US adults: health care resource utilization data from the 2013 National Health and Wellness Survey. J. Am. Acad. Dermatol. 78, 54–61.e1 (2018).
Li, M. et al. Topical vitamin D3 and low-calcemic analogs induce thymic stromal lymphopoietin in mouse keratinocytes and trigger an atopic dermatitis. Proc. Natl Acad. Sci. USA 103, 11736–11741 (2006).
Leyva-Castillo, J. M. et al. Skin thymic stromal lymphopoietin initiates Th2 responses through an orchestrated immune cascade. Nat. Commun. 4, 2847 (2015).
Swindell, W. R. et al. Heterogeneity of inflammatory and cytokine networks in chronic plaque psoriasis. PLoS ONE 7, e34594 (2012).
Chiricozzi, A. et al. Integrative responses to IL-17 and TNF-α in human keratinocytes account for key inflammatory pathogenic circuits in psoriasis. J. Invest. Dermatol. 131, 677–687 (2011).
Hastie, A. T. et al. Baseline sputum eosinophil 1 neutrophil subgroups’ clinical characteristics and longitudinal trajectories for NHLBI Severe Asthma Research Program (SARP 3) cohort. J. Allergy Clin. Immunol. 146, 222–226 (2020).
Kuruvilla, M. E., Lee, F. E.-H. & Lee, G. B. Understanding asthma phenotypes, endotypes, and mechanisms of disease. Clin. Rev. Allerg. Immuol. 56, 219–233 (2019).
Winer, S. et al. Obesity predisposes to Th17 bias. Eur. J. Immunol. 39, 2629–2635 (2009).
Endo, Y. et al. Obesity drives Th17 cell differentiation by inducing the lipid metabolic kinase ACC1. Cell Rep. 12, 1042–1055 (2015).
Manno, G. L. et al. RNA velocity of single cells. Nature 560, 494–498 (2018).
Bergen, V., Lange, M., Peidli, S., Wolf, F. A. & Theis, F. J. Generalizing RNA velocity to transient cell states through dynamical modeling. Nat. Biotechnol. 38, 1408–1414 (2020).
Becht, E. et al. Dimensionality reduction for visualizing single-cell data using UMAP. Nat. Biotechnol. 37, 38–44 (2018).
McInnes, L., Healy, J. & Melville, J. UMAP: uniform manifold approximation and projection for dimension reduction. Preprint at https://doi.org/10.48550/arXiv.1802.03426 (2018).
Gooderham, M. J. et al. Dupilumab: A review of its use in the treatment of atopic dermatitis. J. Am. Acad. Dermatol. 78, S28–S36 (2018).
Hearn, A. P., Kent, B. D. & Jackson, D. J. Biologic treatment options for severe asthma. Curr. Opin. Immunol. 66, 151–160 (2020).
Tubau, C. & Puig, L. Therapeutic targeting of the IL-13 pathway in skin inflammation. Expert Rev. Clin. Immunol. 17, 15–25 (2021).
Laidlaw, T. M. et al. Dupilumab improves upper- and lower-airway disease control in chronic rhinosinusitis with nasal polyps and asthma. Ann. Allergy Asthma Immunol. 126, 584–592.e1 (2021).
Rosen, E. D. & Spiegelman, B. M. PPARγ: a nuclear regulator of metabolism, differentiation, and cell growth. J. Biol. Chem. 276, 37731–37734 (2001).
Tontonoz, P. & Spiegelman, B. M. Fat and beyond: the diverse biology of PPARγ. Annu. Rev. Biochem. 77, 289–312 (2008).
Ahmadian, M. et al. PPARγ signaling and metabolism: the good, the bad and the future. Nat. Med. 19, 557–566 (2013).
Lazar, M. A. Maturing of the nuclear receptor family. J. Clin. Invest. 127, 1123–1125 (2017).
Henriksson, J. et al. Genome-wide CRISPR screens in T helper cells reveal pervasive crosstalk between activation and differentiation. Cell 176, 882–896.e18 (2019).
Woerly, G. et al. Peroxisome proliferator-activated receptors α and γ down-regulate allergic inflammation and eosinophil activation. J. Exp. Med. 198, 411–421 (2003).
Honda, K., Marquillies, P., Capron, M. & Dombrowicz, D. Peroxisome proliferator–activated receptor γ is expressed in airways and inhibits features of airway remodeling in a mouse asthma model. J. Allergy Clin. Immunol. 113, 882–888 (2004).
Hammad, H. et al. Activation of peroxisome proliferator-activated receptor-γ in dendritic cells inhibits the development of eosinophilic airway inflammation in a mouse model of asthma. Am. J. Pathol. 164, 263–271 (2004).
Lee, K. S. et al. Modulation of airway remodeling and airway inflammation by peroxisome proliferator-activated receptor γ in a murine model of toluene diisocyanate-induced asthma. J. Immunol. 177, 5248–5257 (2006).
Chen, T. et al. PPAR-γ promotes type 2 immune responses in allergy and nematode infection. Sci. Immunol. 2, eaal5196 (2017).
Nobs, S. P. et al. PPARγ in dendritic cells and T cells drives pathogenic type-2 effector responses in lung inflammation. J. Exp. Med. 214, 3015–3035 (2017).
Stark, J. M., Tibbitt, C. A. & Coquet, J. M. The metabolic requirements of Th2 cell differentiation. Front. Immunol. 10, 2318 (2019).
Klotz, L. et al. The nuclear receptor PPARγ selectively inhibits Th17 differentiation in a T cell-intrinsic fashion and suppresses CNS autoimmunity. J. Exp. Med. 206, 2079–2089 (2009).
Bapat, S. P. et al. Depletion of fat-resident Treg cells prevents age-associated insulin resistance. Nature 528, 137–141 (2015).
NCD-RisC. Trends in adult body-mass index in 200 countries from 1975 to 2014: a pooled analysis of 1698 population-based measurement studies with 19·2 million participants. Lancet 387, 1377–1396 (2016).
The GBD 2015 Obesity Collaborators. Health effects of overweight and obesity in 195 countries over 25 years. N. Engl. J. Med. 377, 13–27 (2017).
Abarca-Gómez, L. et al. Worldwide trends in body-mass index, underweight, overweight, and obesity from 1975 to 2016: a pooled analysis of 2416 population-based measurement studies in 128.9 million children, adolescents, and adults. Lancet 390, 2627–2642 (2017).
Lucas, C. et al. Longitudinal analyses reveal immunological misfiring in severe COVID-19. Nature 584, 463–469 (2020).
Liston, A. et al. Differentiation of regulatory Foxp3+ T cells in the thymic cortex. Proc. Natl Acad. Sci. USA 105, 11903–11908 (2008).
Lee, P. P. et al. A critical role for Dnmt1 and DNA methylation in T cell development, function, and survival. Immunity 15, 763–774 (2001).
He, W. Adipose-specific peroxisome proliferator-activated receptor knockout causes insulin resistance in fat and liver but not in muscle. Proc. Natl Acad. Sci. USA 100, 15712–15717 (2003).
Rubtsov, Y. P. et al. Regulatory T cell-derived interleukin-10 limits inflammation at environmental interfaces. Immunity 28, 546–558 (2008).
Hastie, A. T. et al. Mixed sputum granulocyte longitudinal impact on lung function in the severe asthma research program. Am. J. Resp. Crit. Care 203, 882–892 (2021).
Wolf, F. A., Angerer, P. & Theis, F. J. SCANPY: large-scale single-cell gene expression data analysis. Genome Biol. 19, 15 (2018).
Traag, V. A., Waltman, L. & Eck, N. J. V. From Louvain to Leiden: guaranteeing well-connected communities. Sci. Rep. 9, 5233 (2019).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2012).
Trapnell, C. et al. Differential analysis of gene regulation at transcript resolution with RNA-seq. Nat. Biotechnol. 31, 46–53 (2012).
Roberts, A., Pimentel, H., Trapnell, C. & Pachter, L. Identification of novel transcripts in annotated genomes using RNA-seq. Bioinformatics 27, 2325–2329 (2011).
Cho, H. et al. Regulation of circadian behaviour and metabolism by REV-ERB-α and REV-ERB-β. Nature 485, 123–127 (2012).
Acknowledgements
We thank A. Levine for advice and critical review of the manuscript; E. Marsh, B. Shy, S. Dodgson and B. Schaar for review of the manuscript; S. Pyle for scientific graphic illustration; Y. Dai, J. Alvarez, A. Cheng and Y. Zhang for technical assistance; and L. Ong, C. Brondos, J. Woo, J. Okano and J. Sawin for administrative assistance. We thank the Severe Asthma Research Program (SARP) for access to their patient data. S.P.B. was supported by U.S. National Institutes of Health (NIH) grants F30 DK096828, T32 GM007198, R38 HL143581 and K38 HL154202. J.L.S. and B.S.N. were supported by U.S. NIH grant P30GM127211. J.M.S. was supported by KAIST (N11210257) and the National Research Foundation of Korea (2017K1A1A2013124 and 2021R1A2C200757311). I.-K.L was supported by the Basic Science Research Program through the National Research Foundation of Korea (NRF) funded by the Ministry of Science and ICT (NRF-2017R1A2B3006406). R.L.G. is supported by NIH grants U01AI52038, R01AI53185, R01AR076082, R01DK121760 and R37AI052453. L.E.C.A. was supported by an American Heart Association grant 16BGIA27790079 and a VA BLR&D Career Development Award 1IK2BX001313. R.M.E. holds the March of Dimes Chair in Molecular and Developmental Biology at the Salk Institute, a SWCRF Investigator Award, and is supported by the NOMIS Foundation–Science of Health, and by U.S. NIH grants HL147835, HL105278, and CA014195. This research was made possible by funding from HHMI, the Foundation Leducq, the Don and Lorraine Freeberg Foundation, the Larry L. Hillblom foundation, and David C. Copley Foundation. Y.Z. was supported by the NOMIS Foundation, the Crohn’s and Colitis Foundation, the Leona M. and Harry B. Helmsley Charitable Trust and the National Institute of Health (R01-AI107027, R01-AI1511123, R21-AI154919, S10-OD023689 and NCI CCSG P30-014195). A.M. holds a Career Award for Medical Scientists from the Burroughs Wellcome Fund, is an investigator at the Chan Zuckerberg Biohub, and is a recipient of the Cancer Research Institute (CRI) Lloyd J. Old STAR grant. The Marson laboratory has received funds from the Innovative Genomics Institute (IGI), the Simons Foundation, and the Parker Institute for Cancer Immunotherapy (PICI). The Marson laboratory has received gifts from the Byers family, B. Bakar, K. Jordan, and E. Radutzky. This work was also supported by National Cancer Institute funded Salk Institute Cancer Center core facilities (CA014195) and the James B. Pendleton Charitable Trust as well as the UCSF Parnassus Flow Cytometry Core funded in part by the Diabetes Research Center grant from the National Institutes of Health (P30 DK063720). Research reported in this publication was supported by the National Institute of Environmental Health Sciences of the National Institutes of Health under Award Number P42ES010337 and by the National Institute of Diabetes and Digestive and Kidney Diseases of the National Institutes of Health under Award Numbers R01DK057978 and R01DK120480 (all to R.M.E.). The content of this manuscript is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
Author information
Authors and Affiliations
Contributions
S.P.B. conceived of the research. S.P.B., J.M.S., B.S.N., I.-K.L., E.G.-Y., K.M.A., P.G.W., J.V.F., D.S., A.T.H., R.L.G., C.J.Y., R.M.E., Y.Z. and A.M. designed and supervised the research. S.P.B., C.W., C.T.M., Y.L., A.Y., Z.J., L.-J.Z., I.V., C.Z., V.Q.N., Z.L., C.C., W.S.Z., X.R., W.Q., S.G.G., C. Liu, E.J.C., M.F., J.L.S., L.E.C.A. and Y.Z. performed experiments. S.P.B., C.W., C.T.M., Y.L., M.C.P., H.H., X.R., J.N.C., L.E.C.A., C. Liddle, S.L., R.M.E., Y.Z. and A.M. analysed data. S.P.B. wrote the manuscript. S.P.B., A.R.A., R.T.Y., M.D., C.J.Y., R.M.E., Y.Z. and A.M. reviewed and edited the manuscript.
Corresponding authors
Ethics declarations
Competing interests
E.G.Y. has received institutional research funds from and is a consultant for Abbvie, Almirall, Amgen, Asana Biosciences, AstraZeneca, Boerhinger-Ingelhiem, Cara Therapeutics, Celgene, Elli Lilly, Galderma, Glenmark/Ichnos Sciences, Janssen, Kyowa Kirin, Leo Pharma, Pfizer, Regeneron, and UCB. E.G.Y. has received institutional research funds from AnaptysBio, Innovaderm, KAO, Kiniksa, Novan, Novartis, and Ralexar. E.G.Y. is a consultant for Arena, Aslan Pharmaceuticals, Bristol-Myers Squibb, Connect Pharma, EMD Serono, Evidera, Incyte, Pandion Therapeutics, RAPT Therapeutics, Sanofi, SATO Pharmaceutical, Siolta Therapeutics, Target Pharma Solutions, and Ventyx Biosciences. R.L.G. is a consultant and has equity interest in MatriSys Biosciences and Sente Inc. C.J.Y. is a Scientific Advisory Board member for and holds equity in Related Sciences and ImmunAI, a consultant for and holds equity in Maze Therapeutics, and a consultant for TReX Bio. C.J.Y. has received research support from Chan Zuckerberg Initiative, Chan Zuckerberg Biohub, and Genentech. A.M. is a compensated cofounder, member of the boards of directors and a member of the scientific advisory boards of Spotlight Therapeutics and Arsenal Biosciences. A.M. is a cofounder, member of the boards of directors and a member of the scientific advisory board of Survey Genomics. A.M. is a compensated member of the scientific advisory board of NewLimit. A.M. owns stock in Arsenal Biosciences, Spotlight Therapeutics, NewLimit, Survey Genomics, PACT Pharma, and Merck. A.M. has received fees from PACT Pharma, Juno Therapeutics, Trizell, Vertex, Merck, Amgen, Genentech, AlphaSights, 23andMe, ALDA, Rupert Case Management and Bernstein. A.M. is an investor in and informal advisor to Offline Ventures and a client of EPIQ. The Marson lab has received research support from Juno Therapeutics, Epinomics, Sanofi, GlaxoSmithKline, Gilead and Anthem.
Peer review
Peer review information
Nature thanks Percy Knolle, Ruslan Medzhitov and Sarah Teichmann for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Diet-induced and monogenic models of obesity provoke increased disease severity upon MC903-AD challenge.
a, Scheme of MC903-induced atopic dermatitis disease model where animals are fed either high fat diet (HFD) or normal diet (ND) for two weeks before initiating MC903 treatments. Black arrows indicate MC903 or EtOH administration to ear. b, Change in ear thickness during AD development as in (a). c, Representative H&E ear histology, Day 11. Scale bars, 100 μm. Dashed line, leukocytic expansion of the dermis. d, Total Tconv, Treg, and CD8+ T cell number from whole ear, Day 11. e–h, Body mass (e), change in ear thickness during AD challenge (f), representative images at Day 10 of challenge (g), and representative H&E ear histology, Day 10 (h) of ND-fed mice, HFD-fed obese mice, or ND-fed ob/ob or db/db mice. i, j, Body mass at Day 0 (i) and change in ear thickness upon AD challenge (j) of 9 week old ob/ob mice fed ND or HFD for three weeks. k–m, Timeline of body mass (k), change in ear thickness (l), and representative images at Day 10 of challenge (m) of always obese (AO), once obese/now lean (OO), and never obese (NO) mice. Red arrow (k) indicates replacement of HFD with control lean diet for OO mice. n = 5 for all groups in (b–m) except 4 never obese (NO) mice were used in (l) and (m). Data are mean ± s.e.m. Peak values were tested with Welch’s t-test in (b, j) and ordinary one-way ANOVA with all groups tested against ND in (f). Ordinary one-way ANOVA with all groups tested against ND in (e) and with all groups tested against all groups in (l). P values were adjusted for multiple comparisons in (d) using Holm-Šídák method and Šídák’s multiple comparisons test in (e, f, l). ns – not significant, #P < 0.07, *P < 0.05, **P < 0.01, ***P < 0.001.
Extended Data Fig. 2 Increased inflammation in obesity in multiple models of allergic inflammatory disease.
a, Scheme of TOP (Tape stripping followed by Ova-Papain exposure)-induced atopic dermatitis disease model with either high fat diet (HFD) or normal diet (ND). Black arrows indicate tape stripping followed by OVA-papain administration. b, Change in ear thickness during AD development. c, Representative pictures of ears at Day 8 of challenge. d, Representative H&E ear histology, Day 8. Scale bars, 100 μm. e–g, Timeline of body mass (e), change in ear thickness (f), and representative pictures of ears (g) at Day 8 of challenge of always obese (AO), once obese/now lean (OO), and never obese (NO) mice. Red arrow (e) indicates replacement of HFD with control lean diet for OO mice. h, Scheme of OVA-Alum allergic asthma model. Purple arrows indicate OVA-Alum i.p. injection. i, Total immune cell numbers from bronchoalveolar lavage fluid upon sacrifice on Day 24. j, Total CD4+ and CD8+ T cell numbers from lung draining lymph node, Day 24. k, Total Tconv, Treg, and CD8+ T cell numbers from whole ear of lean and obese mice challenged with MC903-AD, Day 11. Asthma model was conducted once. Macro, macrophages; Eos, eosinophils; Lymph, lymphocytes; PMNs, polymorphonuclear leukocytes; n = 5 for (b–g); n = 10 for Lean-OVA and Obese-OVA, n = 5 for Lean-Control, and n = 4 for Obese-Control in (i); n = 10 for Lean-OVA, n = 6 for Obese-OVA, n = 4 for Lean-Control, and n = 3 for Obese-Control in (j). n = 5 for all groups in (k). Data are mean ± s.e.m. Peak values were tested with Welch’s t-test in (b) and ordinary one-way ANOVA with preselected followup tests as indicated in (f). P values were adjusted for multiple comparisons using Šídák’s multiple comparisons test in (f) and the Holm-Šídák method in (i–k) where only the Lean-OVA and Obese-OVA groups were compared in (i, j). #P < 0.12, *P < 0.05, **P < 0.01.
Extended Data Fig. 3 Selected flow cytometry gating strategies for evaluating cytokine-competence.
a, b, Different T cell subsets were identified through the use of distinct antibody cocktails. Here, using obese mice, we show how the skin-resident/infiltrating hematopoietic cells can be analyzed by flow cytometry to identify CD4+ T cells (a) and selected cytokine competence of those CD4+ T cells (b).
Extended Data Fig. 4 Multiple models of allergic inflammatory disease demonstrate increased TH17-driven inflammation in obesity.
a, b, Total IL4+, IL13+, IFNγ+, IL17A+, and IL17F+ cell numbers of CD4+ T cells from whole ear (a) or ear skin draining lymph node (b) at Day 8 of mice challenged with TOP-induced AD. c, Total IL4+, IL13+, IFNγ+, IL17A+, and IL17F+ cell numbers of CD4+ T cells from bronchoalveolar lavage fluid upon sacrifice on Day 24 of mice challenged with experimental allergic airway disease (ovalbumin sensitization and challenge). n = 5 for all groups in (a, b); n = 10 for Lean and n = 6 for Obese in (c). Mann-Whitney tests were conducted in (a, b) and P values were adjusted for multiple comparisons using Holm-Šídák method in (a–c). *P < 0.05, **P < 0.01.
Extended Data Fig. 5 Evidence of increased TH17 inflammation or decreased TH2-associated pathology in obese patients with allergic disease.
a, Scheme demonstrating workflow and integration of human AD patient serum proteomics with cytokine-induced gene expression studies of human keratinocytes (KCs) correlated with patient BMI. b, c, Scatterplots depicting the serum protein levels from AD patients of genes specifically induced by IL17A (b) or IL13 (c) in human KCs versus patient BMI. n = 59 moderate-to-severe AD patients for scatter plots in (b, c). r and p values for scatter plots in (b, c) were obtained using Spearman rank correlation. d, e, Sputum eosinophil percentage from human severe asthma patients across a range of BMIs, on their first clinical baseline visit as part of SARP, represented irrespective of age of onset (d) or broken down by age of onset (e). n = 272, 211, and 95 for patients with BMI <30, 30-40, and >40, respectively in (d). n = 113, 83, and 52 (pediatric-onset patients) and n = 70, 66, and 23 (adult-onset patients) with BMI <30, 30–40, and >40, respectively in (e). P value from test of linear trend (post-test after statistically significant ordinary one-way ANOVA) in (d, e).
Extended Data Fig. 6 Gene heatmaps from scRNA-Seq data used in Fig. 1 and Fig. 2.
Heatmaps of transcription factors (a), cytokines (b), and markers of activation, quiescence, and memory (c) overlain on UMAP plot from Fig. 1f to assign names to clusters. Grayscale indicates gene expression, with the highest expressing cells in black.
Extended Data Fig. 7 Blocking lymphocyte egress from secondary lymphoid organs during AD challenge reduces disease severity and TH17 inflammation in the lesions of obese mice.
a, Change in ear thickness during development of MC903-AD of lean or obese mice treated with vehicle (water) or FTY720. b, Representative pictures of ears at Day 10 of challenge. c, Total IL4+, IL13+, IFNγ+, IL17A+, and IL17F+ CD4+ T cell numbers assessed by flow cytometry from draining lymph node of lean (left) and obese (right) mice treated with vehicle or FTY720. n = 5 for all groups in (a–c). Data are mean ± s.e.m. Peak values were tested with Welch’s t-test in (a). P values were adjusted for multiple comparisons using Holm-Šídák method in (c). *P < 0.05, **P < 0.01, ****P < 0.0001.
Extended Data Fig. 8 Targeted anti-IL-4/IL-13 blockade is ineffective in obese mice challenged with TOP.
a, Change in ear thickness during development of TOP-AD of lean or obese mice treated with anti-IL-4/IL-13 or IgG1 isotype control. b, Representative pictures of ears at Day 8 of challenge. c, Representative images of H&E-stained histology of ears at Day 8. Scale bars, 100 μm. d, Lesional number of CD4+ T cells with the indicated cytokine competence as measured by intracellular cytokine staining via flow cytometry from lean and obese mice treated with anti-IL-4/IL-13 or isotype control. n = 5 for all groups in (a–d). Data are mean ± s.e.m. Peak values were tested with Welch’s t-test in (a). P values were adjusted for multiple comparisons using Holm-Šídák method in (d). #P < 0.06, *P < 0.05, **P < 0.01.
Extended Data Fig. 9 PPARγ is differentially expressed in in vitro differentiated TH2 cells.
a, Fragments per kilobase of transcripts per million mapped reads (FPKM) values of nuclear hormone receptor (NHR) superfamily genes differentially expressed in in vitro differentiated TH1, TH2, and TH17 cells. NHR genes that are differentially expressed in TH2 cells are encircled (cells pooled from 4 mice before inducing differentiation in triplicate, same data set used in (e)). b, c, Relative expression (using Hprt expression as housekeeping gene) of indicated genes in in vitro differentiated TH1, TH2, and TH17 cells (cells pooled from 4 mice before inducing differentiation in triplicate). b, Gene expression determined at Hour 120 post induction of differentiation. c, Time course of gene expression from Hours 0–96 post induction of differentiation. d, Western blot of PPARγ and tubulin at Hours 72 and 96 in in vitro differentiated TH1, TH2, and TH17 cells (cells pooled from 3 mice before inducing differentiation). e, FPKM values of genes that are differentially expressed in TH2 cells and involved in transcriptional regulation. Position of Pparg is marked with a red dot. f, Top scoring DNA motif of PPARγ ChIP-Seq peaks in in vitro differentiated TH2 cells via de novo analysis (cells pooled from 4 mice). g, Visualization of PPARγ ChIP-Seq experiment utilizing UCSC genome browser across following genomic loci: Pdk4, Cpt1a, Plin2. Data are mean ± s.e.m. *P < 0.05, **P < 0.01, ***P < 0.001, Student’s t-test.
Extended Data Fig. 10 PPARγ TKO mice display neither overt systemic inflammation nor altered T cell development, and PPARγ Treg cKO (Foxp3Cre Ppargfl/fl) mice do not display an overt difference in AD severity relative to controls.
a–e, Comparison between PPARγ TKO and Control (Cd4Cre Pparg+/+) mice. a, Picture of control and PPARγ TKO spleens and lymph nodes. b, Spleen mass. c, d, Treg (c) and activated Tconv (d) cell frequency in spleen and LN. e, Developing T cell subsets in thymus. f–j, Comparison between PPARγ Treg cKO and Control (Foxp3Cre Pparg+/+) mice. f, Change in ear thickness during development of atopic dermatitis. g, Representative images of H&E-stained histology of ears at Day 10. Scale bars, 100μm. Dashed line, leukocytic expansion of the dermis. h, Absolute lesional leukocyte number assessed by flow cytometry at Day 13. i, Lesional activated Tconv cells as a percentage of Tconv cells assessed by flow cytometry at Day 13. j, Lesional Treg cells as a percentage of CD4+ T cells assessed by flow cytometry at Day 13. Control, Foxp3Cre Pparg+/+. LN, lymph node. For (a–e), n = 3 mice per group. For (f–i), n = 4 mice per group. Data are mean ± s.e.m.
Extended Data Fig. 11 PPARγ in T cells is dispensable for the insulin-sensitizing action of TZDs.
a–d, Glucose tolerance tests (GTTs) of Control (a, b) or PPARγ TKO (c, d) mice that have been treated with or without Rosi. Glucose and insulin traces (a, c) and corresponding area under the curves (AUCs) (b, d) are presented. Control, Cd4Cre; PPARγ TKO, Cd4Cre Ppargfl/fl; For (a–d) n = 10 for all Control mice with exception in (a, b) where n = 9 for the glucose measurements for mice treated with Rosi; n = 8 for PPARγ TKO mice with exception in (c, d) where n = 7 for the PPARγ TKO Rosi-treated mice that were sampled for the serum glucose measurements and n = 5 for the PPARγ TKO Rosi-treated mice that were sampled for the serum insulin measurements. Gluc., Glucose; Ins., Insulin; Rosi, Rosiglitazone. This experiment was conducted once with a weight- and age-matched cohort. Data are mean ± s.e.m. *P < 0.05, **P < 0.01.
Extended Data Fig. 12 Gene heatmaps from scRNA-Seq data used in Fig. 3.
Heatmaps of transcription factors (a), cytokines/cytokine receptors (b), and markers of activation, quiescence, and memory (c) overlain on UMAP plot from Fig. 3c to assign names to clusters. Grayscale indicates gene expression, with the highest expressing cells in black.
Extended Data Fig. 13 Pictures of the ears of obese DMSO- or Rosi-HFD fed mice challenged with AD while treated with anti-IL-4/IL-13 or IgG1 isotype control (a), and initiating Rosi treatment at AD-challenge onset prevents worsening of disease upon treatment with anti-IL-4/IL-13 in obese mice (b, c).
a, Representative pictures of ears at Day 10. b, Change in ear thickness during development of MC903-AD of obese mice treated with Rosi or DMSO with anti-IL-4/IL-13 or IgG1 isotype control. c, Representative images of H&E-stained histology of ears at Day 10. Scale bars, 100 μm. n = 5 for all groups in (a–c). Rosi-mixed HFD introduced four weeks prior to initiation of experimental AD in (a) and upon AD-challenge onset in (b, c). Data are mean ± s.e.m. Peak values were tested with Welch’s t-test in (b). *P < 0.05.
Supplementary information
Supplementary Information
This file contains Supplementary Table 1.
Source data
Rights and permissions
About this article
Cite this article
Bapat, S.P., Whitty, C., Mowery, C.T. et al. Obesity alters pathology and treatment response in inflammatory disease. Nature 604, 337–342 (2022). https://doi.org/10.1038/s41586-022-04536-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-022-04536-0
This article is cited by
-
CRISPR–Cas9 applications in T cells and adoptive T cell therapies
Cellular & Molecular Biology Letters (2024)
-
Piperine reduces hair oiliness by inhibiting adipogenesis of hair stem cells
Applied Biological Chemistry (2024)
-
Adipokines in atopic dermatitis: the link between obesity and atopic dermatitis
Lipids in Health and Disease (2024)
-
Maternal circadian rhythm disruption affects neonatal inflammation via metabolic reprograming of myeloid cells
Nature Metabolism (2024)
-
Bile acid signaling in the regulation of whole body metabolic and immunological homeostasis
Science China Life Sciences (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.