Abstract
Gastrulation is the fundamental process in all multicellular animals through which the basic body plan is first laid down1,2,3,4. It is pivotal in generating cellular diversity coordinated with spatial patterning. In humans, gastrulation occurs in the third week after fertilization. Our understanding of this process in humans is relatively limited and based primarily on historical specimens5,6,7,8, experimental models9,10,11,12 or, more recently, in vitro cultured samples13,14,15,16. Here we characterize in a spatially resolved manner the single-cell transcriptional profile of an entire gastrulating human embryo, staged to be between 16 and 19 days after fertilization. We use these data to analyse the cell types present and to make comparisons with other model systems. In addition to pluripotent epiblast, we identified primordial germ cells, red blood cells and various mesodermal and endodermal cell types. This dataset offers a unique glimpse into a central but inaccessible stage of our development. This characterization provides new context for interpreting experiments in other model systems and represents a valuable resource for guiding directed differentiation of human cells in vitro.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The raw data from our study can be downloaded from ArrayExpress under accession code E-MTAB-9388. The processed data can be downloaded from http://www.human-gastrula.net. Datasets used as references include mouse gastrula data (E-MTAB-6967); pre-implantation embryo data: E-MTAB-3929. Source data are provided with this paper.
Code availability
All data were analysed with standard programs and packages, as detailed in Methods. The code used to create the human gastrula shiny app is available at https://github.com/ScialdoneLab/human-gastrula-shiny.
References
Stern, C. D. Gastrulation: From Cells to Embryo (CSHL Press, 2004).
Tam, P. P. L. & Loebel, D. A. F. Gene function in mouse embryogenesis: get set for gastrulation. Nat. Rev. Genet. 8, 368–381 (2007).
Bardot, E. S. & Hadjantonakis, A. K. Mouse gastrulation: coordination of tissue patterning, specification and diversification of cell fate. Mech. Dev. 163, 103617 (2020).
Arnold, S. J. & Robertson, E. J. Making a commitment: cell lineage allocation and axis patterning in the early mouse embryo. Nat. Rev. Mol. Cell Biol. 10, 91–103 (2009).
O’Rahilly, R. & Müller, F. Developmental stages in human embryos: Revised and new measurements. Cells Tissues Organs 192, 73–84 (2010).
Yamaguchi, Y. & Yamada, S. The Kyoto collection of human embryos and fetuses: History and recent advancements in modern methods. Cells Tissues Organs 205, 314–319 (2019).
Florian, J. & Hill, J. P. An early human embryo (no. 1285, Manchester Collection), with capsular attachment of the connecting stalk. J. Anat. 69, 399–411 (1935).
De Bakker, B. S. et al. An interactive three-dimensional digital atlas and quantitative database of human development. Science 354, aag0053 (2016).
Warmflash, A., Sorre, B., Etoc, F., Siggia, E. D. & Brivanlou, A. H. A method to recapitulate early embryonic spatial patterning in human embryonic stem cells. Nat. Methods 11, 847–854 (2014).
Martyn, I., Kanno, T. Y., Ruzo, A., Siggia, E. D. & Brivanlou, A. H. Self-organization of a human organizer by combined Wnt and Nodal signaling. Nature 558, 132–135 (2018).
Simunovic, M. et al. A 3D model of a human epiblast reveals BMP4-driven symmetry breaking. Nat. Cell Biol. 21, 900–910 (2019).
Moris, N. et al. An in vitro model of early anteroposterior organization during human development. Nature 582, 410–415 (2020).
Chen, D. et al. Human primordial germ cells are specified from lineage-primed progenitors. Cell Rep. 29, 4568–4582.e5 (2019).
Molè, M. A. et al. A single cell characterisation of human embryogenesis identifies pluripotency transitions and putative anterior hypoblast centre. Nat. Commun. 12, 3769 (2021).
Xiang, L. et al. A developmental landscape of 3D-cultured human pre-gastrulation embryos. Nature 577, 537–542 (2020).
Zhou, F. et al. Reconstituting the transcriptome and DNA methylome landscapes of human implantation. Nature 572, 660–664 (2019).
O’Rahilly, R. & Müller, F. eds. Developmental Stages in Human Embryos. (Carnegie Institute of Washington, 1987).
Pijuan-Sala, B. et al. A single-cell molecular map of mouse gastrulation and early organogenesis. Nature 566, 490–495 (2019).
Ma, H. et al. In vitro culture of cynomolgus monkey embryos beyond early gastrulation. Science 366, eaax7890 (2019).
Petropoulos, S. et al. Single-cell RNA-seq reveals lineage and X chromosome dynamics in human preimplantation embryos. Cell 165, 1012–1026 (2016).
Messmer, T. et al. Transcriptional heterogeneity in naive and primed human pluripotent stem cells at single-cell resolution. Cell Rep. 26, 815–824.e4 (2019).
Haghverdi, L., Büttner, M., Wolf, F. A., Buettner, F. & Theis, F. J. Diffusion pseudotime robustly reconstructs lineage branching. Nat. Methods 13, 845–848 (2016).
La Manno, G. et al. RNA velocity of single cells. Nature 560, 494–498 (2018).
Streit, A. The preplacodal region: an ectodermal domain with multipotential progenitors that contribute to sense organs and cranial sensory ganglia. Int. J. Dev. Biol. 51, 447–461 (2007).
Trevers, K. E. et al. Neural induction by the node and placode induction by head mesoderm share an initial state resembling neural plate border and ES cells. Proc. Natl Acad. Sci. USA 115, 355–360 (2017).
Delile, J. et al. Single cell transcriptomics reveals spatial and temporal dynamics of gene expression in the developing mouse spinal cord. Development 146, dev1738078 (2019).
Yang, L. et al. An early phase of embryonic Dlx5 expression defines the rostral boundary of the neural plate. J. Neurosci. 18, 8322–8330 (1998).
Roost, M. S. et al. KeyGenes, a tool to probe tissue differentiation using a human fetal transcriptional atlas. Stem Cell Rep. 4, 1112–1124 (2015).
Chiquoine, A. D. The identification, origin, and migration of the primordial germ cells in the mouse embryo. Anat. Rec. 118, 135–146 (1954).
Magnúsdóttir, E. & Surani, A. M. How to make a primordial germ cell. Development 141, 245–252 (2014).
Sasaki, K. et al. The germ cell fate of cynomolgus monkeys is specified in the nascent amnion. Dev. Cell 39, 169–185 (2016).
Picelli, S. et al. Full-length RNA-seq from single cells using Smart-seq2. Nat. Protoc. 9, 171–181 (2014).
Patro, R., Duggal, G., Love, M. I., Irizarry, R. A. & Kingsford, C. Salmon provides fast and bias-aware quantification of transcript expression. Nat. Methods 14, 417–419 (2017).
Lun, A. T. L., Bach, K. & Marioni, J. C. Pooling across cells to normalize single-cell RNA sequencing data with many zero counts. Genome Biol. 17, 75 (2016).
Wolf, F. A., Angerer, P. & Theis, F. J. SCANPY: Large-scale single-cell gene expression data analysis. Genome Biol. 19, 15 (2018).
McInnes, L., Healy, J., Saul, N. & Großberger, L. UMAP: uniform manifold approximation and projection. J. Open Source Softw. 3, 861 (2018).
Traag, V. A., Waltman, L. & van Eck, N. J. From Louvain to Leiden: guaranteeing well-connected communities. Sci. Rep. 9, 5233 (2019).
Patrick, E. A. Clustering using a similarity measure based on shared near neighbors. IEEE Trans. C-22, 1025–1034 (1973).
Froussios, K., Mourão, K., Simpson, G., Barton, G. & Schurch, N. Relative abundance of transcripts (RATs): identifying differential isoform abundance from RNA-seq. F1000Research 8, 213 (2019).
Dobin, A. et al. STAR: Ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Bergen, V., Lange, M., Peidli, S., Wolf, F. A. & Theis, F. J. Generalizing RNA velocity to transient cell states through dynamical modeling. Nat. Biotechnol. 38, 1408–1414 (2020).
Durinck, S., Spellman, P. T., Birney, E. & Huber, W. Mapping identifiers for the integration of genomic datasets with the R/ Bioconductor package biomaRt. Nat. Protoc. 4, 1184–1191 (2009).
Efremova, M., Vento-Tormo, M., Teichmann, S. A. & Vento-Tormo, R. CellPhoneDB: inferring cell–cell communication from combined expression of multi-subunit ligand–receptor complexes. Nat. Protoc. 15, 1484–1506 (2020).
Kiselev, V. Y., Yiu, A. & Hemberg, M. scmap: Projection of single-cell RNA-seq data across data sets. Nat. Methods 15, 359–362 (2018).
Grün, D. et al. Single-cell messenger RNA sequencing reveals rare intestinal cell types. Nature 525, 251–255 (2015).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Scialdone, A. et al. Computational assignment of cell-cycle stage from single-cell transcriptome data. Methods 85, 54–61 (2015).
Leng, N. et al. Oscope identifies oscillatory genes in unsynchronized single-cell RNA-seq experiments. Nat. Methods 12, 947–950 (2015).
Segal, J. M. et al. Single cell analysis of human foetal liver captures the transcriptional profile of hepatobiliary hybrid progenitors. Nat. Commun. 10, 3350 (2019).
Li, H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. Preprint at https://arxiv.org/abs/1303.3997 (2013).
Yang, R., Van Etten, J. L. & Dehm, S. M. Indel detection from DNA and RNA sequencing data with transIndel. BMC Genomics 19, 270 (2018).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902.e21 (2019).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Chen, G. et al. Chemically defined conditions for human iPSC derivation and culture. Nat. Methods 8, 424–429 (2011).
Johansson, B. M. & Wiles, M. V. Evidence for involvement of activin A and bone morphogenetic protein 4 in mammalian mesoderm and hematopoietic development. Mol. Cell. Biol. 15, 141–151 (1995).
Choi, H. M. T. et al. Third-generation in situ hybridization chain reaction: multiplexed, quantitative, sensitive, versatile, robust. Develpoment 145, dev165753 (2018).
Tyser, R. C. V. et al. Characterization of a common progenitor pool of the epicardium and myocardium. Science 371, eabb2986 (2021).
Acknowledgements
Human embryonic material was provided by the MRC–Wellcome Trust funded (grant no. 099175/Z/12/Z and MR/R006237/1) Human Developmental Biology Resource (www.hdbr.org). We thank N. Ashley (Oxford MRC single-cell facility) for help with sequencing, M. De Bruijn, B. Gottgens, J. Palis, L. Robertson, T. Rodriguez and M.-E. Torres-Padilla for helpful comments. This work was funded by British Heart Foundation Immediate Postdoctoral Basic Science Research Fellowship no. FS/18/24/33424 to R.C.V.T., JSPS Overseas Research Fellowship to S.N., European Research Council advanced grant ERC: 741707 to L.V., funding from the Helmholtz Association to A.S. and Wellcome Awards 105031/C/14/Z, 108438/Z/15/Z, 215116/Z/18/Z and 103788/Z/14/Z to S.S.
Author information
Authors and Affiliations
Contributions
Human gastrula processing: R.C.V.T. and S.S. Human ES cell in vitro experiments: S.N. Computational analyses of sequence data: E.M. and A.S. Gastrula single-cell annotation and analyses: R.C.V.T., E.M., A.S. and S.S. Human ES cell in vitro data analysis: R.C.V.T. and S.N. Preparation of illustrations and figures: R.C.V.T. Preparation of manuscript draft: R.C.V.T., E.M., S.N., A.S. and S.S. Editing and review of final manuscript: R.C.V.T., E.M., S.N., L.V., A.S. and S.S. Study coordination: A.S. and S.S.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks Joshua Welch and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Quality control of scRNA-seq dataset.
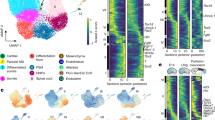
a, Dorsal view of the dissected embryonic disk showing the primitive streak and node (Scale bar = 500μm; n = 1). b, Brightfield images showing embryo dissection with schematic diagrams highlighting the three anatomical regions collected (yolk sac, rostral and caudal regions of embryonic disk; Scale bar = 500μm; n = 1). c, Metrics used to assess the quality of the scRNA-seq libraries. From top to bottom the scatter plots show the number of detected genes, the fraction of reads mapped to the human genome, the fraction of reads mapped to mitochondrial genes and the fraction of reads mapped to ERCC spike-ins, all as a function of the total number of reads. Cells that passed quality control are marked by green circles, while black circles indicate cells that failed the quality control and were excluded from downstream analyses. d, The boxplots show the total log expression of normalized counts for XIST and Y-genes across all clusters. While XIST was mostly not detected, Y-chromosome genes had always non-zero counts; this suggests that there is no contamination from maternal tissues in any of the clusters. n = 1195 cells were examined from a single embryo. Horizontal black lines denote median values and boxes cover the 25th and 75th percentiles range; whiskers extend to 1.5 × IQR. e, The stacked barplots indicate the percentages of cells from each cluster in the phase G1, S or G2/M of the cell cycle, as predicted from their transcriptomic profiles. f, Insertion-deletion length and size distribution of gastrula and fetal liver data. Y axis represents total number of indels on merged cells, while x axis represents indel length in base pairs. Hemato-Endothelial Progenitors (HEP), Endoderm (End), Advanced Mesoderm (AM), Primitive Streak (PS), Extraembryonic Mesoderm (ExM), Axial Mesoderm (AxM), Erythroblasts (Ery), Emergent Mesoderm (EM), Epiblast (Epi), Nascent Mesoderm (NM), Ectoderm (Amniotic/Embryonic (EAE)).
Extended Data Fig. 2 Characterisation and comparison of a CS7 human gastrula with Non-human primate and Mouse.
a, Heatmap with the normalized log expression of well characterized marker genes for the identified cell types: Epiblast (Epi), Ectoderm (Amniotic/Embryonic (EAE)), Primitive Streak (PS), Nascent Mesoderm (NM), Emergent Mesoderm (EM), Advanced Mesoderm (AM), Extraembryonic Mesoderm (ExM), Axial Mesoderm (AxM), Endoderm (Endo), Hemato-Endothelial Progenitors (HEP), Erythroblasts (Ery). b, Stacked bar plots highlighting the anatomical region that cells were collected from and the percentage breakdown of each cluster. Numbers in brackets represent the total number of cells per cluster. c, Heatmap showing the fraction of human gastrula cells allocated to mouse cell types at E7.25 (data from18). d, Dendrogram showing hierarchical clustering of the transcriptomes of cell types from human gastrula and cultured cynomolgus macaque embryos at 16-day post-fertilization (from19). e, Top, UMAP plots showing the log expression of MEST and GCNT2. Bottom, violin plots showing the log expression of total transcripts (top row) and selected isoforms scaled by the maximum value in different cell types. Isoform names refer to Ensembl nomenclature.
Extended Data Fig. 3 In Vitro vs In Vivo comparisons.
a, Dendrogram representation built on corrected expression values obtained with Seurat showing comparison of an in vitro model of pluripotency with in vivo data. b, Log-fold changes of expression levels of the genes between primed vs naïve hESC (y axis) and CS7 epiblast vs E6 data (x axis). Selected genes are highlighted in red; the blue line is obtained through a linear regression. A statistically significant positive correlation is found (Pearson’s correlation coefficient ~0.63, p-value = 3e-107), indicating that the hESC resemble the in vivo primed and naïve states at the transcriptome-wide level. c, Heatmaps showing the correlations between the transcriptomic profiles of the human gastrula cell types (rows) and sections of human gastruloids taken at different positions along the rostral-caudal axis (columns) in two different replicates (Gastruloid 1 and Gastruloid 2). Only the values of the statistically significant correlations (p-value < 0.01; 2-tailed Pearson’s correlation, see Methods) are reported, while all the non-significant correlations were set to 0. d, UMAP representation of the human gastrula data with the PGCs highlighted. d, Diffusion map of cells from all 11 clusters. The first three diffusion components (DC1, 2, 3) are plotted in different combinations. In the top panels, cells are coloured by the clusters they belong to, while in the bottom panels the colours indicate the region each cell was dissected from. Ectoderm (amniotic/embryonic) (EAE), Epiblast (Epi), Primitive Streak (PS), Axial Mesoderm (AxM), Nascent Mesoderm (NM), Emergent Mesoderm (EM), Advanced Mesoderm (AM), Erythroblasts (Ery), Hemato-Endothelial Progenitors (HEP), Endoderm (Endo), Extraembryonic Mesoderm (ExM).
Extended Data Fig. 4 Rostral and Caudal differences in diversification of mesodermal subtypes.
a, UMAP highlighting combinatorial gene expression. Individual gene expression (left) is reported as the log expression whilst combinatorial plots (right) show scaled log expression values. b, Diffusion map of cells from the 6 mesoderm related clusters (Primitive Streak, PS; Nascent Mesoderm, NM; Emergent Mesoderm, EM; Mesoderm, Meso; Axial Mesoderm, AxM; Extraembryonic Mesoderm, ExM), with the first and the second diffusion components plotted. c, Diffusion map of mesodermal showing the log expression levels of mesodermal markers genes. d, Differential gene expression between rostral and caudal advanced mesoderm cells. Significantly upregulated in rostral (*) or caudal (#) cells. e-j, Diffusion map of mesodermal clusters showing log expression levels of mesoderm subtype markers.
Extended Data Fig. 5 Differentiation of the epiblast.
a, Diffusion map of cells from the Epiblast, Primitive Streak, Nascent Mesoderm and Ectoderm (amniotic/embryonic). The first two diffusion components are plotted (DC1 and DC2) and cells are colored by their cluster (top) or the anatomical region they were isolated from (bottom). b and c, Normalized log gene expression changes along a pseudotime coordinate (see Fig. 4a) running from 0 to 1 and spanning the Ectoderm (amniotic/embryonic) (EAE), the Epiblast (EPI), the Primitive Streak (PS) and the Nascent Mesoderm (NM), as depicted by the arrow on top. The selected genes highlight Primitive Streak and mesoderm formation (panel b) as well as ectoderm differentiation (panel c).
Extended Data Fig. 6 Mesoderm formation in human and mouse.
a, Diffusion map with cells from the human (top two plots) or mouse (bottom two plots) Epiblast, Primitive Streak and Nascent Mesoderm clusters. Cells are colored based on their cluster of origin or on their diffusion pseudotime coordinate. b, Upset plot for the number of differentially expressed (DE) genes as a function of the diffusion pseudotime (dpt) shown in panel a in mouse (m) or human (h). Here, only genes that are differentially expressed in both species and with a log-fold change > 1 along the trajectory are included. Genes are split according to their increasing (up) or decreasing (down) trend as a function of dpt. c, Comparison of pseudotime analysis during primitive streak and nascent mesoderm formation in human and mouse (data from18). Cells in epiblast (Epi), Primitive Streak (PS) and Nascent Mesoderm (NM) clusters from human and mouse embryos at matching stages (see Methods) were independently aligned along a differentiation trajectory and a diffusion pseudotime coordinate (dpt) was calculated for each (top). The expression pattern and standard error of the mean of selected genes along pseudotime is plotted for human (left, continuous lines) and mouse (right, dashed lines). Both SNAI1 and CDH1 showed comparable expression profiles during mesoderm formation in mouse and human whilst MSGN1was differently expressed between species.
Extended Data Fig. 7 Characterization of EMT during hESC mesoderm formation.
a, Bright-field microscopy images of D0 hESC (left), D1 Meso (center) and D1 MEK Inhibition (right) ESC colonies (top panels). Fluorescence microscopy images of E-Cadherin staining (bottom panels). b, Quantification of transcript levels for selected pluripotent, EMT and mesendoderm genes across the three conditions PLU, ME, ME+PD. c, Quantification of transcript levels for selected non-neural ectoderm genes across the three conditions PLU, ME, ME+PD. (n = 6 from three different experiments. Center line, median; box limits, upper and lower quartiles; whiskers, minimum and maximum; dots, mean value per experiement. ns = p-value ≥ 0.05; *** = p-value < 0.001; **** = p-value < 0.0001 (Ordinary one-way ANOVA after passing a Shapiro-Wilk normality test. Kruskal-Wallis multiple comparison test used if Shapiro-Wilk normality test failed (MSGN1, TDGF1, HAND1, DLX5). House-keeping genes, HKGs. See SI Table 17 for source data and exact p-values
Extended Data Fig. 8 Comparison of signaling during mesoderm formation in the human and mouse.
Heatmap comparison of the z-score-normalized log expression values of components of FGF, TGF-β and Wnt signaling pathways in the human gastrula, mouse embryos (E7.25 stage) and cultured cynomolgus macaque embryos (16 d.p.f stage). From human and mouse we considered the Epiblast (Epi), Primitive Streak (PS) and Nascent Mesoderm (NM) clusters; in the macaque, we used the clusters annotated as postL-Epi, L-Gast1 and L-Gast2.
Extended Data Fig. 9 Endoderm subcluster identification.
a, Heatmap showing the scaled log expression levels of marker genes of the four endodermal subclusters. b, Percentage of cells dissected from the Caudal, Rostral or Yolk Sac portion of the embryo in the four endodermal subclusters. c, Percentage of cells based on their predicted cell-cycle phase of the four endodermal subclusters. d, Diffusion map of cells from the Endoderm cluster. The first two diffusion components (DC1 and DC2) are plotted and cells are coloured by the sub clusters (left), anatomical origin (central) or the predicted cell-cycle phase (right). Yolk Sac, YS; Definitive Endoderm (DE) 1 and 2. e, Diffusion map of cells from the Endoderm cluster with DC1 and DC3 plotted, showing log expression levels of Pan-endoderm, Yolk-sac endoderm and definitive endoderm markers. f, Log expression levels of Anterior Definitive Endoderm markers. These genes are more highly expressed in DE2. g, Log expression levels of Gut Endoderm markers, showing limited expression. h, Maximum intensity projection and mid-sagittal section (h’) of an E7.0 mouse embryo showing expression of Gjb1 (yolk sac endoderm marker) as well as Cer1 and Hhex (anterior definitive endoderm markers) using Hybridization Chain Reaction (n = 4). Cer1 and Hhex show greater expression in the anterior embryonic endoderm. Anterior, Ant; Posterior, Pos; Yolk-sac Endoderm, YSE. i, Violin plots showing the scaled log expression of total transcripts (top row) and individual isoforms in different endodermal subclusters. Isoform lables refer to Ensembl transcript numbers.
Extended Data Fig. 10 Hemato-Endothelial Progenitors subclusters.
a, Boxplots showing the total log expression of normalized counts for XIST and Y-genes in Erythroblasts (Ery) and Hemato-Endothelial Progenitors (HEP), indicating no contamination from maternal tissue. n = 143 cells were examined from a single embryo. Horizontal black lines denote median values and boxes cover the 25th and 75th percentiles range; whiskers extend to 1.5 × IQR.b, UMAP of HEP and Erythroblast clusters showing log expression of blood related marker genes. c, Heatmap showing the scaled log expression of well-characterized marker genes for both the Hemato-Endothelial Progenitors subclusters and Erythroblast cluster. d, Heatmap showing the normalized log expression levels of the top 5 marker genes of the four Hemato-Endothelial Progenitors subclusters. e, Diffusion maps of HEP subclusters and Erythroblasts showing diffusion components (DC) 1, 2 and 3. f, Violin plots showing the scaled log expression of Globin genes in the five blood related clusters: Erythroblasts (Ery), Myeloid Progenitors (MP), Endothelium, Megakaryocyte-Erythroid Progenitors (MEP) and Erythro-Myeloid progenitors (EMP). Each grey dot represents a single cell. g, Heatmap showing the estimated mapping of human Erythroid and HEP subclusters to mouse blood-related clusters. Scalebar represents the fraction of human cells mapped to each category. h, Bar graph showing the number of cells present in the mouse scRNA-seq dataset18 at different development timepoints.
Supplementary information
Supplementary Information
This file contains Supplementary Notes 1 – 4 and Supplementary References.
Supplementary Table 1
Human gastrula cluster marker genes. Table showing the marker genes ranked by statistical significance for the 11 different clusters identified, including: Epiblast (Epi), ectoderm (amniotic/embryonic) (EAE), primitive streak (PS), nascent mesoderm (NM), emergent mesoderm (EM), advanced mesoderm (AM), extraembryonic mesoderm (ExM), axial mesoderm (AxM), endoderm (Endo), HEP and erythroblasts (Ery).
Supplementary Table 2
Cell origin per cluster. Table showing the percentage of cells from a specific anatomical region for each cluster.
Supplementary Table 3
Transcript isoform differences for all clusters comparisons. Tables showing transcript isoform comparisons between clusters. Each worksheet refers to the comparison of a single cluster with every other cluster of cell types, and includes the names of the genes whose isoforms are differentially expressed with the relative P value. One-sided chi-square test. extraembryonic mesoderm (ExM), HEP, ectoderm (amniotic/embryonic) (EAE).
Supplementary Table 4
Top 30 genes with highest log-fold change in each quadrant of CS7 vs E6 embryos and naive vs primed human ES cell correlation. Table showing the top 30 genes with the highest log2-fold changes in each quadrant of the CS7 vs E6 embryos and naive vs primed human ES cell comparison (Fig. 3b). The numeric values in the table represent log2-fold changes in shown comparisons.
Supplementary Table 5
Differentially expressed genes along the epiblast to nascent mesoderm trajectory. List of differentially expressed genes and their trends during epiblast to nascent mesoderm transition in the human gastrula. The trends are ‘up’ or ‘down’ when there is an increasing or decreasing trend with a log2-fold change greater than 1 between the expression values at the beginning and at the end of the trajectory; ‘flat’ genes are those having a log fold change less than 1 between initial and final expression value.
Supplementary Table 6
Genes correlating with TBXT along the epiblast to nascent mesoderm trajectory. List of genes that correlate with TBXT along the epiblast to nascent mesoderm trajectory. Values represent correlation coefficient (coef), P value (p-val) and FDR. Two-sided Spearman’s rho test.
Supplementary Table 7
Differentially expressed genes in human and mouse during epiblast to nascent mesoderm differentiation. Comparison of differentially expressed genes in mouse and human along the epiblast to nascent mesoderm differentiation trajectory. Genes are marked by whether they are differentially expressed in mouse (DE mouse) or human (DE human), their expression trends in mouse and human and FDR in these species. The trends are ‘up’ or ‘down’ when there is an increasing or decreasing trend with a log2 fold change greater than 1 between the expression values at the beginning and at the end of the trajectory; ‘flat’ genes are those having a log fold change less than 1 between initial and final expression value.
Supplementary Table 8
Ectoderm (amniotic/embryonic) subcluster genes. Table of the top 50 marker genes for the ectoderm (amniotic/embryonic) subclusters (amnion and non-neural ectoderm (NNE)).
Supplementary Table 9
Primordial germ cell primitive streak differentially expressed genes. List of top differentially expressed genes between PGC and primitive streak with FDR. Two-sided Wilcoxon rank-sum test.
Supplementary Table 10
PGC cross-species gene-expression analysis. Table showing 50 shared and disparate genes when comparing human primordial germ cells to cynomolgus macaque and mouse. Only genes differentially expressed for PGCs in each species are shown.
Supplementary Table 11
Endoderm subcluster marker genes. List of endoderm subcluster marker genes. Definitive endoderm, DE; yolk sac, YS.
Supplementary Table 12
Transcript isoform differences for endoderm subclusters. Tables showing transcript isoform comparisons between endoderm subclusters. Each worksheet refers to the comparison of a single subcluster with every other endoderm subclusters, and includes the names of the genes whose isoforms are differentially expressed with the relative P value. Definitive endoderm, DE; yolk sac, YS. One-sided chi-square test.
Supplementary Table 13
HEP subcluster genes. List of HEP subcluster marker genes and associated FDR.
Supplementary Table 14
Transcript Isoform differences in HEP subclusters. Tables showing transcript isoform comparisons between HEP subclusters. Each worksheet refers to the comparison of a single subcluster with every other HEP subclusters, and includes the names of the genes whose isoforms are differentially expressed with the relative p-value. One-sided chi-square test.
Supplementary Table 15
Real-time PCR primer details. List of real-time PCR primer sequences.
Supplementary Table 16
Extraembryonic and advanced mesoderm differentially expressed genes. List of top 100 most differentially expressed genes between advanced mesoderm and extraembryonic (Exe) mesoderm.
Supplementary Table 17
Source data for real-time PCR analysis.
Rights and permissions
About this article
Cite this article
Tyser, R.C.V., Mahammadov, E., Nakanoh, S. et al. Single-cell transcriptomic characterization of a gastrulating human embryo. Nature 600, 285–289 (2021). https://doi.org/10.1038/s41586-021-04158-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-021-04158-y
This article is cited by
-
HAND factors regulate cardiac lineage commitment and differentiation from human pluripotent stem cells
Stem Cell Research & Therapy (2024)
-
Transcriptional signals of transformation in human cancer
Genome Medicine (2024)
-
Derivation of human primordial germ cell-like cells in an embryonic-like culture
Nature Communications (2024)
-
DELVE: feature selection for preserving biological trajectories in single-cell data
Nature Communications (2024)
-
The ethical and legal challenges of human foetal brain tissue-derived organoids
EMBO Reports (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.