Abstract
The MRGPRX family of receptors (MRGPRX1–4) is a family of mas-related G-protein-coupled receptors that have evolved relatively recently1. Of these, MRGPRX2 and MRGPRX4 are key physiological and pathological mediators of itch and related mast cell-mediated hypersensitivity reactions2,3,4,5. MRGPRX2 couples to both Gi and Gq in mast cells6. Here we describe agonist-stabilized structures of MRGPRX2 coupled to Gi1 and Gq in ternary complexes with the endogenous peptide cortistatin-14 and with a synthetic agonist probe, respectively, and the development of potent antagonist probes for MRGPRX2. We also describe a specific MRGPRX4 agonist and the structure of this agonist in a complex with MRGPRX4 and Gq. Together, these findings should accelerate the structure-guided discovery of therapeutic agents for pain, itch and mast cell-mediated hypersensitivity.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The coordinate and cryo-EM map of MRGPRX2–Gq–cortistatin-14, MRGPRX2–Gi1–cortistatin-14, MRGPRX2–Gq–(R)-ZINC-3573, MRGPRX2–Gi1–(R)-ZINC-3573 and MRGPRX4–Gq–MS47134 have been deposited to PDB (EMDB) with accession codes 7S8L (EMD-24896), 7S8M (EMD-24897), 7S8N (EMD-24898), 7S8O (EMD-24899) and 7S8P (EMD-24900), respectively. The cryoEM micrographs of MRGPRX4–Gq–MS47134, MRGPRX2–Gq–cortistatin-14, MRGPRX2–Gq–(R)-ZINC-3573, MRGPRX2–Gi1–cortistatin-14 and MRGPRX2–Gi1–(R)-ZINC-3573 have been deposited in the EMPIAR database (https://www.ebi.ac.uk/empiar/) with accession numbers EMPIAR-10852, EMPIAR-10853, EMPIAR-10854, EMPIAR-10855 and EMPIAR-10856, respectively. The MRGPRX2 antagonist C9 and C9-6 and negative control C7, and the MRGPRX4 agonist MS47134 and negative control X2-2 will be made available via Sigma-Millipore.
References
Zylka, M. J., Dong, X., Southwell, A. L. & Anderson, D. J. Atypical expansion in mice of the sensory neuron-specific Mrg G protein-coupled receptor family. Proc. Natl Acad. Sci. USA 100, 10043–10048 (2003).
McNeil, B. D. et al. Identification of a mast-cell-specific receptor crucial for pseudo-allergic drug reactions. Nature 519, 237–241 (2015).
Green, D. P., Limjunyawong, N., Gour, N., Pundir, P. & Dong, X. A mast-cell-specific receptor mediates neurogenic inflammation and pain. Neuron 101, 412–420.e413 (2019).
Yu, H. et al. MRGPRX4 is a bile acid receptor for human cholestatic itch. eLife 8, e48431 (2019).
Meixiong, J., Vasavda, C., Snyder, S. H. & Dong, X. MRGPRX4 is a G protein-coupled receptor activated by bile acids that may contribute to cholestatic pruritus. Proc. Natl Acad. Sci. USA 116, 10525–10530 (2019).
Chompunud Na Ayudhya, C., Roy, S., Alkanfari, I., Ganguly, A. & Ali, H. Identification of gain and loss of function missense variants in MRGPRX2’s transmembrane and intracellular domains for mast cell activation by substance P. Int. J. Mol. Sci. 20, 5247 (2019).
Ikoma, A., Steinhoff, M., Stander, S., Yosipovitch, G. & Schmelz, M. The neurobiology of itch. Nat. Rev. Neurosci. 7, 535–547 (2006).
Greaves, M. W. & Wall, P. D. Pathophysiology of itching. Lancet 348, 938–940 (1996).
Liu, Q. et al. Sensory neuron-specific GPCR Mrgprs are itch receptors mediating chloroquine-induced pruritus. Cell 139, 1353–1365 (2009).
Lembo, P. M. et al. Proenkephalin A gene products activate a new family of sensory neuron–specific GPCRs. Nat. Neurosci. 5, 201–209 (2002).
Azimi, E. et al. Dual action of neurokinin-1 antagonists on Mas-related GPCRs. JCI Insight 1, e89362 (2016).
Kroeze, W. K. et al. PRESTO-Tango as an open-source resource for interrogation of the druggable human GPCRome. Nat. Struct. Mol. Biol. 22, 362–369 (2015).
Lansu, K. et al. In silico design of novel probes for the atypical opioid receptor MRGPRX2. Nat. Chem. Biol. 13, 529–536 (2017).
Subramanian, H. et al. β-Defensins activate human mast cells via Mas-related gene X2. J. Immunol. 191, 345–352 (2013).
Olsen, R. H. J. et al. TRUPATH, an open-source biosensor platform for interrogating the GPCR transducerome. Nat. Chem. Biol. 16, 841–849 (2020).
English, J. G. et al. VEGAS as a platform for facile directed evolution in mammalian cells. Cell 178, 748–761.e717 (2019).
Wacker, D. et al. Structural features for functional selectivity at serotonin receptors. Science 340, 615–619 (2013).
Lu, L., Kulka, M. & Unsworth, L. D. Peptide-mediated mast cell activation: ligand similarities for receptor recognition and protease-induced regulation. J. Leukoc. Biol. 102, 237–251 (2017).
Che, T. et al. Structure of the nanobody-stabilized active state of the kappa opioid receptor. Cell 172, 55–67.e15 (2018).
Che, T. et al. Nanobody-enabled monitoring of kappa opioid receptor states. Nat. Commun. 11, 1145 (2020).
Rasmussen, S. G. F. et al. Crystal structure of the β2 adrenergic receptor–Gs protein complex. Nature 477, 549–555 (2011).
Rosenbaum, D. M. et al. GPCR engineering yields high-resolution structural insights into β2-adrenergic receptor function. Science 318, 1266–1273 (2007).
Ogasawara, H., Furuno, M., Edamura, K. & Noguchi, M. Novel MRGPRX2 antagonists inhibit IgE-independent activation of human umbilical cord blood-derived mast cells. J. Leukoc. Biol. 106, 1069–1077 (2019).
Shinkai, H. et al. N-(cyclohexylcarbonyl)-d-phenylalanines and related compounds. A new class of oral hypoglycemic agents. 2. J. Med. Chem. 32, 1436–1441 (1989).
Irwin, J. J. & Shoichet, B. K. ZINC–a free database of commercially available compounds for virtual screening. J. Chem. Inf. Model. 45, 177–182 (2005).
Meixiong, J. et al. Identification of a bilirubin receptor that may mediate a component of cholestatic itch. eLife 8, e44116 (2019).
Azimi, E., Reddy, V. B. & Lerner, E. A. Brief communication: MRGPRX2, atopic dermatitis and red man syndrome. Itch (Phila) 2, e5 (2017).
Chen, E. et al. Inflamed ulcerative colitis regions associated to MRGPRX2-mediated mast cell degranulation and cell activation modules, defining a new therapeutic target. Gastroenterology 160, 1709–1724 (2021).
Kozlitina, J. et al. An African-specific haplotype in MRGPRX4 is associated with menthol cigarette smoking. PLoS Genet. 15, e1007916 (2019).
Li, Z. et al. Targeting human Mas-related G protein-coupled receptor X1 to inhibit persistent pain. Proc. Natl Acad. Sci. U.S.A. 114, E1996–E2005 (2017).
Thapaliya, M., Chompunud Na Ayudhya, C., Amponnawarat, A., Roy, S. & Ali, H. Mast cell-specific MRGPRX2: a key modulator of neuro-immune interaction in allergic diseases. Curr. Allergy Asthma Rep. 21, 3 (2021).
Kim, K. et al. Structure of a hallucinogen activated Gq-coupled 5-HT2a serotonin receptor. Cell 182, 1574–1588.e1519 (2020).
Draper-Joyce, C. J. et al. Structure of the adenosine-bound human adenosine A1 receptor-Gi complex. Nature 558, 559–563 (2018).
Koehl, A. et al. Structure of the µ-opioid receptor–Gi protein complex. Nature 558, 547–552 (2018).
Mastronarde, D. N. Automated electron microscope tomography using robust prediction of specimen movements. J. Struct. Biol. 152, 36–51 (2005).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Punjani, A., Zhang, H. & Fleet, D. J. Non-uniform refinement: adaptive regularization improves single-particle cryo-EM reconstruction. Nat. Methods 17, 1214–1221 (2020).
Bepler, T., Kelley, K., Noble, A. J. & Berger, B. Topaz-Denoise: general deep denoising models for cryoEM and cryoET. Nat. Commun. 11, 5208 (2020).
Rosenthal, P. B. & Henderson, R. Optimal determination of particle orientation, absolute hand, and contrast loss in single-particle electron cryomicroscopy. J. Mol. Biol. 333, 721–745 (2003).
Heymann, J. B. & Belnap, D. M. Bsoft: image processing and molecular modelling for electron microscopy. J. Struct. Biol. 157, 3–18 (2007).
Sanchez-Garcia, R. et al. DeepEMhacer: a deep learning solution for cryo-EM volume post-processing. Commun. Biol. 4, 874 (2021).
Grant, T., Rohou, A. & Grigorieff, N. cisTEM, user-friendly software for single-particle image processing. eLife 7, e35383 (2018).
Xing, C. et al. Cryo-EM structure of the human cannabinoid receptor CB2–Gi signaling complex. Cell 180, 645–654.e613 (2020).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
Robertson, M. J., van Zundert, G. C. P., Borrelli, K. & Skiniotis, G. GemSpot: a pipeline for robust modeling of ligands into cryo-EM maps. Structure 28, 707–716.e703 (2020).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D 66, 12–21 (2010).
Besnard, J. et al. Automated design of ligands to polypharmacological profiles. Nature 492, 215–220 (2012).
Kroeze, W. K. et al. PRESTO-Tango as an open-source resource for interrogation of the druggable human GPCRome. Nat. Struct. Mol. Biol. 22, 362–369 (2015).
Longo, P. A., Kavran, J. M., Kim, M. S. & Leahy, D. J. Transient mammalian cell transfection with polyethylenimine (PEI). Methods Enzymol. 529, 227–240 (2013).
Acknowledgements
This work was supported by NIH grants U24DA116195 (to B.L.R., B.K.S. and J.J.) and R35GM122481 (to B.K.S.), and by the Michael Hooker Distinguished Professorship to B.L.R. and NIH grant R01-DK121969, R01-DK121032 and R56-AI139620 to S.N.A. We thank J. Peck and J. Strauss of the UNC Cryo-EM Core Facility for technical assistance in this project. W.Y. is partly supported by Program for Breakthrough Biomedical Research funded by the Sandler Foundation, University of California, San Francisco. L.Y.J. is a Howard Hughes Medical Institute investigator. We thank S.-L. Shyng for the plasmids encoding human Kir6.2 and SUR1. The Titan X Pascal used for this research was kindly donated to J.F.F. by the NVIDIA Corporation.
Author information
Authors and Affiliations
Contributions
C.C. designed the experiments, performed the cloning, expression and purification of all the signalling complexes for cryo-EM study, built the models, refined the structures, performed BRET assays, participated in MRGPRX4 drug screening, analysed the data, and prepared the figures, tables and manuscript. H.J.K. designed experiments, performed drug screening and functional assays, analysed the data and assisted in preparing the manuscript, and reviewed and edited the manuscript. J.F.F. made the grids, and collected and processed the cryo-EM data. I.S. performed the SAR study for MRGPRX2. H.C., C.Z. and J.L. performed the SAR study for MRGPRX4, and designed, synthesized and characterized the MRGPRX4 agonists. W.Y. performed electrophysiology for Kir6.2/SUR1. R.H.G. assisted in the modelling and validation of structures. B.W.H. performed the LAD2 mast cell degranulation assay. B.J.B. assisted in the structure modelling. J.K. supported agonist discovery and optimization for MRGPRX4. S.T.S. performed the GPCRome assay. B.E.K. designed the Gq protein construct and assisted in the reviewing and editing of the manuscript. K.L., J.D.M., W.K.K., J.F.D. and R.H.J.O. performed the initial screening for MRGPRX4 compounds. J.G.E., X.-P.H. and Y.L. helped with the initial functional assays. S.Z. and K.K. assisted in the protein expression. S.N.A. guided the LAD2 mast cell degranulation assay. J.J. supervised the medicinal chemistry experiments. B.K.S. supervised the docking and compound design and edited the manuscript. L.Y.J. contributed to funding application. B.L.R. supervised the entire project, guided the structural and functional work and prepared the manuscript.
Corresponding authors
Ethics declarations
Competing interests
A patent describing the MRGPRX2 antagonists has been filed by UCSF listing B.L.R., B.K.S., C.C., I.S. and H.J.K. as inventors.
Additional information
Peer review information Nature thanks the anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 MRGPRX2 transducerome screening using TRUPATH.
a, MRGPRX2 effectively couples to 14 distinct G proteins upon stimulation of agonists (R)-ZINC-3573 and cortistatin-14 (C-14) in HEK 293T cells. Net BRET values of MRGPRX2 together with positive controls of either neurotensin receptor 1 (NTSR1, agonist NT1-13) or β2AR (agonist isoproterenol) are shown in each panel. Data represent mean ± s.e.m. of n = 3 biological replicates. b, Heatmap of the relative log potency (logEC50) of (R)-ZINC-3573 and cortistatin-14 for 14 distinct G proteins. c, Heatmap of the relative efficacy (Emax) of (R)-ZINC-3573 and cortistatin-14 for 14 distinct G proteins.
Extended Data Fig. 2 CryoEM images and data-processing of MRGPRX2-Gq-(R)-ZINC-3573, MRGPRX2-Gi1-(R)-ZINC-3573 and MRGPRX4-Gq-MS47134 complex.
a–c, Representative motion corrected cryo-EM micrographs (scale bar, 100 nm) of respective ligand bound GPCR heterotrimeric complex particles imaged at a nominal 45k x magnification and representative two-dimensional class averages. The experiment was repeated three times with similar result. The exact number of movies and particles used for each complex are shown in the flow chart. d–f, Flow chart of cryo-EM data processing, GSFSC plot of auto-masked final map (black) and map-to-model real-space cross correlation (red) as calculated form phenix.mtriage. g–i, Respective polar plots of particle angular distributions and local resolution estimations heat maps. j–l, Local cryo-EM density maps of TM1-7, respective ligands, and α5 and αN helix of respective G-protein.
Extended Data Fig. 3 CryoEM images and data-processing of MRGPRX2-Gq-Cortistatin-14 and MRGPRX2-Gi1-Cortistatin-14 complex.
a, b, Representative motion corrected cryo-EM micrograph (scale bar, 100 nm) of MRGPRX2 G-protein cortistatin-14 (C14) particles imaged at a nominal 45k x magnification and representative two-dimensional class averages. The experiment was repeated three times with similar result. The exact number of movies and particles used for each complex are shown in the flow chart. c, d, Flow chart of cryo-EM data processing. GSFSC plot of auto-masked final map (black) and map-to-model real-space cross correlation (red) as calculated form phenix.mtriage. e, f, Viewing direction distribution and local resolution estimation heat maps. g, h, Local cryo-EM density maps of TM1-7, Cortistatin-14 ligand, α5 and α N helix of respective G-protein. Also shown inset are residues W151 and F82 of the b-subunit (blue).
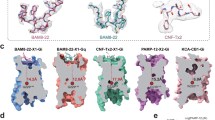
Extended Data Fig. 4 Structural comparison of Gq- and Gi1-coupled MRGPRX2 complex.
a, b, Structural comparison of the MRGPRX2-Gi1-cortistatin-14 complex (blue) with MRGPRX2-Gq-cortistatin-14 complex (cyan). Top view for the key interactions in sub-pocket 1 (a). Side view to show the overall conformational of cortistatin-14 (b). c–e, structural comparison of MRGPRX2-Gi1-(R)-ZINC-3573 complex with MRGPRX2-Gq-(R)-ZINC-3573 complex. Gi1 and Gq are shown in green and salmon, respectively. Gi1-coupled MRGPRX2 and Gq-coupled MRGPRX2 are shown in blue and cyan, respectively. Side view of the whole complex (c), top view (d) and bottom view (e) of MRGPRX2. f, ICL3 of Gq is not clearly resolved in the Gq-coupled MRGPRX2 complex. g, Close-up view of the ICL3 in the Gi1-coupled MRGPRX2 structure with surrounding EM map at a threshold of 0.14. h, i, MRGPRX2 ICL3 mutations R214ICL3A and L216ICL3A impair cortistatin-14 (h) and (R)-ZINC-3573 (i) stimulated Gi1 activation. Data represent mean ± s.e.m. of n = 3 biological replicates. j, k, BRET2 Gi1 assays reveal that I135ICL2A mutation of MRGPRX2 attenuates cortistatin-14 (j) and (R)-ZINC-3573 (k) stimulated Gi1 activation. Data represent mean ± s.e.m. of n = 3 biological replicates. l–m, BRET2 Gq assays reveal that I135ICL2A mutation of MRGPRX2 greatly reduced cortistatin-14 (l) and (R)-ZINC-3573 (m) stimulated Gq activation. Data represent mean ± s.e.m. of n = 3 biological replicates.
Extended Data Fig. 5 Non-conserved motifs in Mas-related GPCRs and the critical role of acidic residues E1644.60 and D1845.38 in MRGPRX2 activation.
a, Sequence alignment of the key residues in sodium site, DRY motif, PIF motif and CWxP motif, as well as residues involved in disulfide bond formation in Mas-related GPCRs. Class A conserved residues are highlighted in green. b, cryoEM map of the TM4-TM5 disulfide bond in MRGPRX2-Gq-(R)-ZINC-3573 complex. c, d, Break of the TM4-TM5 disulfide bond by C1684.64A and C1805.34A mutations abolishes the cortistatin-14 stimulated Gq activation (c) and reduces the Emax of (R)-ZINC-3573 stimulated Gq activation by 60% (d). Data represent mean ± s.e.m. of n = 3 biological replicates. e–g, Compared with WT (e), E1644.60A (f) and D1845.38A (g) totally abolish the peptide stimulated Gq activation of MRGPRX2. Data represent mean ± s.e.m. of n = 3 biological replicates.
Extended Data Fig. 6 Unique structural features of MRGPRX2 and MRGPRX4.
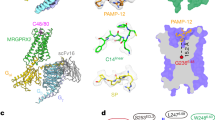
a, MRGPRX2 and MRGPRX4 have a unique structural arrangement at the PIF motif compared with the G protein coupled active structures of 5-HT2AR (PDB ID 6WHA), A2AR (PDB ID 5G53) and β2AR (PDB ID 3SN6). Residue 5.50 shifts away from the TM3-TM6 interface and does not engage L3.40 and F6.44 in MRGPRX2 and MRGPRX4. b, With G6.48, TM6 of both MRGPRX2 and MRGPRX4 packs closer to TM3 compared with the G protein coupled active structures of 5-HT2AR (PDB ID 6WHA), A2AR (PDB ID 5G53) and β2AR (PDB ID 3SN6), leading to an occluded canonical agonist binding pocket. c, (R)-ZINC-3573, cortistatin-14 and MS47134 bind to MRGPRX2 and MRGPRX4 at a position that is far away from residue 6.48, respectively. Cortistatin-14 is shown as cartoon. Small molecule compounds of receptors are shown as spheres.
Extended Data Fig. 7 Analog screening and functional characterization of MRGPRX2 antagonists.
a, b, Dose-response curves of initial 14 analogs of ‘1592 (a) and 8 analogs of C9 (b) in the presence of EC80 concentration of (R)-ZINC-3573 using MRGPRX2 FLIPR Ca2+ assay. Data represent mean ± s.e.m. of n = 3 biological replicates. c, Dose-response curves of two potent MRGPRX2 antagonists C9 and C9-6 and an inactive compound C7 in the presence of EC80 of each MRGPRX2 peptide using MRGPRX2 FLIPR Ca2+ assay. Data represent mean ± s.e.m. of n = 3 biological replicates.
Extended Data Fig. 8 Functional characterization of optimized MRGPRX4 agonists.
a, Dose-response curves of Kir6.2/SUR1 current inhibition by indicated chemicals. Data represent mean ± s.e.m. from n=4 biological replicates. b, d, f, h. Current-voltage relationships of whole-cell traces recorded in 150 mM KCl with the supplements of indicated chemicals of the labeled concentrations. c, e, g, Time courses showing the whole-cell-current responses to the indicated chemicals of the labeled concentrations. i, MRGPRX4 agonists X4-4 and MS47134 have a higher selectivity over Kir6.2/SUR1 channel compared with nateglinide. EC50 (nM) of each tested compound is shown. j, Screening of MS47134 across the GPCRome (at 320 receptors) using the PRESTO-Tango platform with 3 µM MS47134. Red dashed line indicated threefold of basal levels. Data represent mean ± s.e.m. of fold over basal for each receptor (n=4 technical replicates).
Extended Data Fig. 9 Structural comparison of Gq-coupled MRGPRX2 and MRGPRX4.
a–d, Structural comparison of the MRGPRX4-Gq-MS47134 complex with the MRGPRX2-Gq-(R)-ZINC-3573 complex. The receptor and Gq protein of MRGPRX4-Gq complex are colored by green and blue, respectively. The receptor and Gq protein of MRGPRX2-Gq complex are colored by cyan and salmon, respectively. Side view (a), Close-up view of αN-ICL2 interaction region (b), α5 helix region (c), and the cytoplasmic side of receptors (d). e, The acidic residues E1574.60 and D1775.38 of MRGPRX4 are shielded by the inserted ECL2. Side chain of D177 is not resolved but modeled here for a better visual interpretation. f, Residues E1644.60 and D1845.38 of MRGPRX2 extend to the cationic agonists accessible pocket. g, Due to the variance in residue 2.39, Y243H5.23 of Gq adopts different side-chain conformations to interact with Y130ICL2 of MRGPRX4 and Y137ICL2 of MRGPRX2. h, i, BRET2 Gq assays for Y130ICL2A of MRGPRX4 (h) and Y137ICL2A of MRGPRX2 (i). Data represent mean ± s.e.m. of n = 3 biological replicates.
Supplementary information
Supplementary Information
See Supplementary Information page 1 for contents
Rights and permissions
About this article
Cite this article
Cao, C., Kang, H.J., Singh, I. et al. Structure, function and pharmacology of human itch GPCRs. Nature 600, 170–175 (2021). https://doi.org/10.1038/s41586-021-04126-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-021-04126-6
This article is cited by
-
Proinflammatory chemokine CXCL14 activates MAS-related G protein-coupled receptor MRGPRX2 and its putative mouse ortholog MRGPRB2
Communications Biology (2024)
-
Promiscuous G-protein activation by the calcium-sensing receptor
Nature (2024)
-
Synovial microenvironment-influenced mast cells promote the progression of rheumatoid arthritis
Nature Communications (2024)
-
Bitter taste receptor activation by cholesterol and an intracellular tastant
Nature (2024)
-
Structural basis of amine odorant perception by a mammal olfactory receptor
Nature (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.