Abstract
Protein expression and turnover are controlled through a complex interplay of transcriptional, post-transcriptional and post-translational mechanisms to enable spatial and temporal regulation of cellular processes. To systematically elucidate such gene regulatory networks, we developed a CRISPR screening assay based on time-controlled Cas9 mutagenesis, intracellular immunostaining and fluorescence-activated cell sorting that enables the identification of regulatory factors independent of their effects on cellular fitness. We pioneered this approach by systematically probing the regulation of the transcription factor MYC, a master regulator of cell growth1,2,3. Our screens uncover a highly conserved protein, AKIRIN2, that is essentially required for nuclear protein degradation. We found that AKIRIN2 forms homodimers that directly bind to fully assembled 20S proteasomes to mediate their nuclear import. During mitosis, proteasomes are excluded from condensing chromatin and re-imported into newly formed daughter nuclei in a highly dynamic, AKIRIN2-dependent process. Cells undergoing mitosis in the absence of AKIRIN2 become devoid of nuclear proteasomes, rapidly causing accumulation of MYC and other nuclear proteins. Collectively, our study reveals a dedicated pathway controlling the nuclear import of proteasomes in vertebrates and establishes a scalable approach to decipher regulators in essential cellular processes.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Source data for Figs. 1d, e, 2a, b, 3a, b, d, 4e, f and Extended Data Figs. 2, 4a, b, g, i, 5f, 6d are included in the Supplementary Information files of the manuscript. Raw FASTQ files for RNA-seq analyses are available through the Gene Expression Omnibus (accession code GSE157663). Negative staining and cryo-EM density maps are deposited in the Electron Microscopy Data Bank with the accession codes EMD-11649 and EMD-12341, respectively. The atomic model is deposited under Protein Data Bank ID 7NHT. Raw micrographs and particle stacks are available in the EMPIAR database (EMPIAR-10752). TMT quantitative proteomics, V5 and GST co-IP/MS data have been deposited to the ProteomeXchange Consortium PRIDE repository with the accession codes PXD021898, PXD027184 and PXD027343, respectively. Human cancer cell line RNA-seq data were obtained from the ArrayExpress repository (https://www.ebi.ac.uk/arrayexpress/experiments/E-MTAB-2706). AKIRIN2 interactors identified in this study were cross-compared to the BioGRID database (https://thebiogrid.org/120430/summary/homo-sapiens/akirin2.html). The crystallographic native human 20S proteasome structure used for model building and the cryo-EM structure of the substrate-engaged human 26S proteasome shown for comparison in Extended Data Fig. 7f were obtained from the RCSB protein database (https://www.rcsb.org/structure/5LE5 and https://www.rcsb.org/structure/6MSJ, respectively).
Code availability
Custom code for screen analysis is available on GitHub (https://github.com/ZuberLab/crispr-process-nf, https://github.com/ZuberLab/crispr-mageck-nf). Custom ImageJ scripts applied for image analysis (see Methods for details) are available upon reasonable request.
References
Johnston, L. A., Prober, D. A., Edgar, B. A., Eisenman, R. N. & Gallant, P. Drosophila myc regulates cellular growth during development. Cell 98, 779–790 (1999).
Sabo, A. et al. Selective transcriptional regulation by Myc in cellular growth control and lymphomagenesis. Nature 511, 488–492 (2014).
Muhar, M. et al. SLAM-seq defines direct gene-regulatory functions of the BRD4–MYC axis. Science 360, 800–805 (2018).
Dang, C. V. MYC on the path to cancer. Cell 149, 22–35 (2012).
Hnisz, D. et al. Super-enhancers in the control of cell identity and disease. Cell 155, 934–947 (2013).
He, T. C. et al. Identification of c-MYC as a target of the APC pathway. Science 281, 1509–1512 (1998).
Dani, C. et al. Extreme instability of myc mRNA in normal and transformed human cells. Proc. Natl Acad. Sci. USA 81, 7046-7050 (1984).
Gregory, M. A. & Hann, S. R. c-Myc proteolysis by the ubiquitin-proteasome pathway: stabilization of c-Myc in Burkitt’s lymphoma cells. Mol. Cell. Biol. 20, 2423–2435 (2000).
Schwanhausser, B. et al. Global quantification of mammalian gene expression control. Nature 473, 337–342 (2011).
Meyers, R. M. et al. Computational correction of copy number effect improves specificity of CRISPR–Cas9 essentiality screens in cancer cells. Nat. Genet. 49, 1779–1784 (2017).
Dempster, J. M. et al. Extracting biological insights from the Project Achilles genome-scale CRISPR screens in cancer cell lines. Preprint at https://doi.org/10.1101/720243 (2019).
Michlits, G. et al. Multilayered VBC score predicts sgRNAs that efficiently generate loss-of-function alleles. Nat. Methods 17, 708–716 (2020).
Sears, R., Leone, G., DeGregori, J. & Nevins, J. R. Ras enhances Myc protein stability. Mol. Cell 3, 169–179 (1999).
Welcker, M. et al. The Fbw7 tumor suppressor regulates glycogen synthase kinase 3 phosphorylation-dependent c-Myc protein degradation. Proc. Natl Acad. Sci. USA 101, 9085–9090 (2004).
Inoue, S. et al. Mule/Huwe1/Arf-BP1 suppresses Ras-driven tumorigenesis by preventing c-Myc/Miz1-mediated down-regulation of p21 and p15. Genes Dev 27, 1101–1114 (2013).
Qiao, X. et al. UBR5 is coamplified with MYC in breast tumors and encodes an ubiquitin ligase that limits MYC-dependent apoptosis. Cancer Res. 80, 1414–1427 (2020).
Goto, A. et al. Akirins are highly conserved nuclear proteins required for NF-κB-dependent gene expression in Drosophila and mice. Nat. Immunol. 9, 97–104 (2008).
Tartey, S. et al. Akirin2 is critical for inducing inflammatory genes by bridging IκB-ζ and the SWI/SNF complex. EMBO J. 33, 2332–2348 (2014).
Bonnay, F. et al. Akirin specifies NF-κB selectivity of Drosophila innate immune response via chromatin remodeling. EMBO J. 33, 2349–2362 (2014).
Fischer, M. Census and evaluation of p53 target genes. Oncogene 36, 3943–3956 (2017).
Stark, C. et al. BioGRID: a general repository for interaction datasets. Nucleic Acids Res. 34, D535–D539 (2006).
Smith, D. M. et al. Docking of the proteasomal ATPases’ carboxyl termini in the 20S proteasome’s α ring opens the gate for substrate entry. Mol. Cell 27, 731–744 (2007).
Cuylen-Haering, S. et al. Chromosome clustering by Ki-67 excludes cytoplasm during nuclear assembly. Nature 587, 285–290 (2020).
Schmitz, M. H. et al. Live-cell imaging RNAi screen identifies PP2A-B55α and importin-β1 as key mitotic exit regulators in human cells. Nat. Cell Biol. 12, 886–893 (2010).
Reits, E. A., Benham, A. M., Plougastel, B., Neefjes, J. & Trowsdale, J. Dynamics of proteasome distribution in living cells. EMBO J. 16, 6087–6094 (1997).
Pack, C. G. et al. Quantitative live-cell imaging reveals spatio-temporal dynamics and cytoplasmic assembly of the 26S proteasome. Nat. Commun. 5, 3396 (2014).
Peters, J. M., Franke, W. W. & Kleinschmidt, J. A. Distinct 19 S and 20 S subcomplexes of the 26 S proteasome and their distribution in the nucleus and the cytoplasm. J. Biol. Chem. 269, 7709–7718 (1994).
Wendler, P. & Enenkel, C. Nuclear transport of yeast proteasomes. Front. Mol. Biosci. 6, 34 (2019).
Palacios, V., Kimble, G. C., Tootle, T. L. & Buszczak, M. Importin-9 regulates chromosome segregation and packaging in Drosophila germ cells. J. Cell Sci. 134, jcs258391 (2021).
Budenholzer, L., Breckel, C., Hickey, C. M. & Hochstrasser, M. The Sts1 nuclear import adapter uses a non-canonical bipartite nuclear localization signal and is directly degraded by the proteasome. J. Cell Sci. 133, jcs236158 (2020).
Dominguez, D. et al. A high-resolution transcriptome map of cell cycle reveals novel connections between periodic genes and cancer. Cell Res. 26, 946–962 (2016).
Manasanch, E. E. & Orlowski, R. Z. Proteasome inhibitors in cancer therapy. Nat. Rev. Clin. Oncol. 14, 417–433 (2017).
Parnas, O. et al. A genome-wide CRISPR screen in primary immune cells to dissect regulatory networks. Cell 162, 675–686 (2015).
Brockmann, M. et al. Genetic wiring maps of single-cell protein states reveal an off-switch for GPCR signalling. Nature 546, 307–311 (2017).
Umkehrer, C. et al. Isolating live cell clones from barcoded populations using CRISPRa-inducible reporters. Nat. Biotechnol. 39, 174–178 (2020).
Schambach, A. et al. Lentiviral vectors pseudotyped with murine ecotropic envelope: increased biosafety and convenience in preclinical research. Exp. Hematol. 34, 588–592 (2006).
Wang, T. et al. Identification and characterization of essential genes in the human genome. Science 350, 1096–1101 (2015).
Morgens, D. W. et al. Genome-scale measurement of off-target activity using Cas9 toxicity in high-throughput screens. Nat. Commun. 8, 15178 (2017).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Li, W. et al. MAGeCK enables robust identification of essential genes from genome-scale CRISPR/Cas9 knockout screens. Genome Biol. 15, 554 (2014).
Klijn, C. et al. A comprehensive transcriptional portrait of human cancer cell lines. Nat. Biotechnol. 33, 306–312 (2015).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet.journal 17, https://journal.embnet.org/index.php/embnetjournal/article/view/200 (2011).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
O’Leary, N. A. et al. Reference sequence (RefSeq) database at NCBI: current status, taxonomic expansion, and functional annotation. Nucleic Acids Res. 44, D733–D745 (2016).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Zhu, A., Ibrahim, J. G. & Love, M. I. Heavy-tailed prior distributions for sequence count data: removing the noise and preserving large differences. Bioinformatics 35, 2084–2092 (2019).
Dorfer, V. et al. MS Amanda, a universal identification algorithm optimized for high accuracy tandem mass spectra. J. Proteome Res. 13, 3679-3684 (2014).
Smyth, G. K. Linear models and empirical Bayes methods for assessing differential expression in microarray experiments. Stat. App. Gene. Mol. Biol. 3, Article3 (2004).
Mellacheruvu, D. et al. The CRAPome: a contaminant repository for affinity purification–mass spectrometry data. Nat. Methods 10, 730–736 (2013).
Schaab, C., Geiger, T., Stoehr, G., Cox, J. & Mann, M. Analysis of high accuracy, quantitative proteomics data in the MaxQB database. Mol. Cell Proteomics 11, M111. 014068 (2012).
Mi, H., Muruganujan, A., Ebert, D., Huang, X. & Thomas, P. D. PANTHER version 14: more genomes, a new PANTHER GO-slim and improvements in enrichment analysis tools. Nucleic Acids Res. 47, D419–D426 (2019).
Altschul, S. F. et al. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 25, 3389–3402 (1997).
Emms, D. M. & Kelly, S. OrthoFinder: solving fundamental biases in whole genome comparisons dramatically improves orthogroup inference accuracy. Genome Biol. 16, 157 (2015).
Crooks, G. E., Hon, G., Chandonia, J. M. & Brenner, S. E. WebLogo: a sequence logo generator. Genome Res. 14, 1188–1190 (2004).
Katoh, K., Rozewicki, J. & Yamada, K. D. MAFFT online service: multiple sequence alignment, interactive sequence choice and visualization. Briefings Bioinform. 20, 1160–1166 (2019).
Waterhouse, A. M., Procter, J. B., Martin, D. M., Clamp, M. & Barton, G. J. Jalview Version 2—a multiple sequence alignment editor and analysis workbench. Bioinformatics 25, 1189–1191 (2009).
Ishida, T. & Kinoshita, K. PrDOS: prediction of disordered protein regions from amino acid sequence. Nucleic Acids Res. 35, W460–W464 (2007).
Kosugi, S., Hasebe, M., Tomita, M. & Yanagawa, H. Systematic identification of cell cycle-dependent yeast nucleocytoplasmic shuttling proteins by prediction of composite motifs. Proc. Natl Acad. Sci. USA 106, 10171–10176 (2009).
Cuff, J. A. & Barton, G. J. Application of multiple sequence alignment profiles to improve protein secondary structure prediction. Proteins 40, 502–511 (2000).
Drozdetskiy, A., Cole, C., Procter, J. & Barton, G. J. JPred4: a protein secondary structure prediction server. Nucleic Acids Res. 43, W389–W394 (2015).
Lupas, A., Van Dyke, M. & Stock, J. Predicting coiled coils from protein sequences. Science 252, 1162–1164 (1991).
Doblmann, J. et al. apQuant: accurate label-free quantification by quality filtering. J. Proteome Res. 18, 535–541 (2019).
Studier, F. W. Protein production by auto-induction in high density shaking cultures. Protein Expr. Purif. 41, 207–234 (2005).
Sonn-Segev, A. et al. Quantifying the heterogeneity of macromolecular machines by mass photometry. Nat. Commun. 11, 1772 (2020).
Dohmen, R. J. & Scheffner, M. (eds) Ubiquitin Family Modifiers and the Proteasome (Springer, 2012).
Besche, H. C., Haas, W., Gygi, S. P. & Goldberg, A. L. Isolation of mammalian 26S proteasomes and p97/VCP complexes using the ubiquitin-like domain from HHR23B reveals novel proteasome-associated proteins. Biochemistry 48, 2538–2549 (2009).
Schorb, M., Haberbosch, I., Hagen, W. J. H., Schwab, Y. & Mastronarde, D. N. Software tools for automated transmission electron microscopy. Nat. Methods 16, 471–477 (2019).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. eLife 7, e42166 (2018).
Goddard, T. D. et al. UCSF ChimeraX: meeting modern challenges in visualization and analysis. Protein Sci. 27, 14–25 (2018).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Rohou, A. & Grigorieff, N. CTFFIND4: Fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Zivanov, J., Nakane, T. & Scheres, S. H. W. Estimation of high-order aberrations and anisotropic magnification from cryo-EM data sets in RELION-3.1. IUCrJ 7, 253–267 (2020).
Sanchez-Garcia, R. et al. DeepEMhancer: a deep learning solution for cryo-EM volume post-processing. Commun. Biol. 4, 874 (2021).
Schrader, J. et al. The inhibition mechanism of human 20S proteasomes enables next-generation inhibitor design. Science 353, 594–598 (2016).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Murshudov, G. N. et al. REFMAC5 for the refinement of macromolecular crystal structures. Acta Crystallogr. D Biol. Crystallogr. 67, 355–367 (2011).
Kim, D. E., Chivian, D. & Baker, D. Protein structure prediction and analysis using the Robetta server. Nucleic Acids Res. 32, W526–W531 (2004).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Croll, T. I. ISOLDE: a physically realistic environment for model building into low-resolution electron-density maps. Acta Crystallogr. D Struct. Biol. 74, 519–530 (2018).
Afonine, P. V. et al. Real-space refinement in PHENIX for cryo-EM and crystallography. Acta Crystallogr. D Struct. Biol. 74, 531–544 (2018).
Lukinavicius, G. et al. SiR-Hoechst is a far-red DNA stain for live-cell nanoscopy. Nat. Commun. 6, 8497 (2015).
Dong, Y. et al. Cryo-EM structures and dynamics of substrate-engaged human 26S proteasome. Nature 565, 49–55 (2019).
Acknowledgements
We are grateful to all members of the Zuber and Haselbach laboratories and to A. Pauli, C. Plaschka, T. Clausen, D. Gerlich, M. Petrovic, F. Grebien, N. Brown, G. Winter and M. Petronczki for experimental advice and helpful discussions. We thank members of the Obenauf, Gerlich, Clausen, Peters and Busslinger laboratories at IMP and IMBA for sharing reagents; K. Aumayr, P. Pasierbeck, the IMP BioOptics flow cytometry, microscopy and image analysis team and T. Kreslavskiy for cell sorting and imaging; G. Dürnberger and A. Lüttig at the IMP/IMBA Protein Biochemistry Core Facility for performing quantitative proteomics and immunoprecipitation mass spectrometry; A. Sommer and the VBCF-NGS team (https://www.vbcf.ac.at) for deep sequencing services; the Electron Microscopy Facility team at Vienna BioCenter Core Facilities for negative staining electron microscopy; A. Meinhart for model building advice; O. Kaya for experimental support; the IMP/IMBA Molecular Biology Service for continuous support; and G. Riddihough (Life Science Editors) for help with editing. We acknowledge Diamond Light Source for access and support of the cryo-EM facilities at the United Kingdom’s national Electron Bio-imaging Centre (under proposal EM BI25222), funded by the Wellcome Trust, the Medical Research Council and the Biotechnology and Biological Sciences Research Council. This work was funded by a Starting Grant from the European Research Council (ERC-StG-336860) to J.Z., the Austrian Science Fund (SFB grant F4710) to J.Z. and EPIC-XS (project number 823839), funded by the Horizon 2020 Programme of the European Union, to K.M. M.d.A. is the recipient of a DOC fellowship of the Austrian Academy of Sciences. Research at the IMP is generously supported by Boehringer Ingelheim and the Austrian Research Promotion Agency (Headquarter grant FFG-852936).
Author information
Authors and Affiliations
Contributions
M.d.A. and M.H. contributed equally and will be putting their name first on the citation in their CVs. M.d.A., M.H. and J.Z. conceived and planned this project. M.d.A., M.H., H.B. and I.G. designed and conducted experiments with help from M.S., S.K. and E.R. M.d.A., M.H., H.B., I.G., K.S., D.H. and J.Z. analysed and interpreted original data. A.S. performed phylogenetic analyses. T.L. established scripts for imaging analysis. D.H. and K.S. performed the 3D structure reconstruction. T.N. and R.I. performed deep sequencing and mass spectrometry data analyses, respectively. J.J., S.D., R.K. and M.V. established critical reagents and methodology. K.M. and G.V. provided critical input on experimental designs and data analyses. M.d.A., M.H., D.H. and J.Z. co-wrote the manuscript with input from all co-authors.
Corresponding authors
Ethics declarations
Competing interests
J.Z. is a founder, shareholder and scientific advisor of Quantro Therapeutics. J.Z., D.H. and the Zuber and Haselbach laboratories receive research support and funding from Boehringer Ingelheim. J.J. is now an employee of Twist Bioscience and T.N. is now an employee of Quantro Therapeutics. Other authors declare no competing interests.
Additional information
Peer review information Nature thanks Roderick Beijersbergen, Cordula Enenkel, Edmond Watson and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Engineering and validation of clonal iCas9 cell lines.
a, Schematic of clonal iCas9 cell line engineering and validation. Roman numbers indicate the vectors used for each cell line. In K562, generation of tightly regulatable iCas9 clones required additional suppression of leaky transcription from Tet-responsive promoters via a TetR-KRAB fusion protein. b–d, Evaluation of iCas9 function. b, Flow cytometric evaluation of inducible Cas9-P2A-GFP (RKO, K562) or -BFP (MIA-PaCa-2) expression in the presence or absence of DOX. c, Competitive proliferation assays in iCas9 cells transduced with sgRNAs targeting the core-essential genes PSMA3 (RKO), PLK1 (MIA-PaCa-2), or RPL23 (K562). Percentage of sgRNA+ cells was monitored by flow cytometry in the presence or absence of DOX for 10 days. Values are normalized to Day 0. d, Evaluation of surface marker editing in iCas9 cells transduced with sgRNAs targeting the surface markers CD151 (RKO, MIA-PaCa-2) or CD46 (K562). Editing was evaluated by immunostaining and flow cytometry 48 h after Cas9 induction. e, Analysis of MYC/MYCN mRNA levels based on42. A MYC/MYCN specific antibody was used for the detection of MYC. rtTA3, reverse tetracycline transactivator; DOX, doxycycline; PuroR, puromycin resistance gene; HygroR, hygromycin resistance gene; TRE3G, tetracycline response element; 2A, P2A self-cleaving peptide; GFP, green fluorescent protein; BFP, blue fluorescent protein; APC, allophycocyanin; RPKM, reads per kilo base per million mapped reads.
Extended Data Fig. 2 Identification of MYC regulators in iCas9 screens depends on their turnover and essentiality.
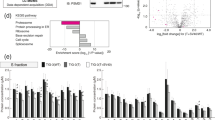
a–b, First timepoints of FACS-based CRISPR MYC-regulator (a) and dropout (b) screens. a, Gene-level sgRNA enrichment in MYClo (left panels) or MYChi cells (right panels) over MYCmid (RKO, K562) or unsorted (MIA-PaCa-2) cells and MAGeCK41 one-sided P values. Dashed lines indicate 95th percentile of enrichment and significance (P < 0.01). Essential genes based on12,37,38 within the scoring window are highlighted in red, 20S-CP subunits in blue. b, Gene-level sgRNA depletion in unsorted populations at the first screen timepoint compared to day 0. SgRNAs targeting highly turned-over essential proteins deplete already 2.5 days after Cas9-induction. c, d, Second screen timepoints as in a and b. The identification of MYC regulators at each timepoint depends on their turnover and essentiality. Short-lived essential proteins scored at the first timepoint in FACS-based (a) and dropout screens (b); however, due to rapid effects on cell viability (d), sgRNAs targeting these genes were undetectable at the second timepoint in FACS-based screens (c). Conversely, knockout of more stable proteins or protein complexes such as the 20S proteasome had only limited effects in MYC-regulation- (a) or dropout- screens after 2.5 days (b), while effects were readily detectable 4–5 days post Cas9 induction (c, d). The kinetics of gene editing, protein turnover and depletion of sgRNA-expressing cells are further determined by the cellular context, with rapid effects in diploid RKO cells, and delayed effects in hypertriploid K562 and tetraploid MIA-PaCa-2 cells.
Extended Data Fig. 3 AKIRIN2 is an essential regulator of MYC expression.
a–c, Flow cytometric quantification of endogenous MYC protein levels after inducible knockout of MYC regulators in RKO (a), MIA-PaCa2 (b), and K562 cells (c) 1-3 days after Cas9 induction. For MIA-PaCa2 and K562 the timepoint with the maximal effect on MYC protein abundance is shown. RKO maximal timepoints as in Fig. 2c. d–e, Percentage of cleaved-caspase-3+ (d) and dead cells (e) quantified by flow-cytometry before and 1-3 days after induction of AKIRIN2 knockout. f, Competitive proliferation assays. Percentage of sgRNA+ iCas9-RKO cells was monitored by flow cytometry in 24 h intervals after Cas9 induction. Values are normalized to Day 0. Data in a is representative of three, in b–c of two independent experiments. Data in d–f is shown as mean ± s.d. (n = 3 biological replicates). PE, phycoerythrin.
Extended Data Fig. 4 AKIRIN2 regulates nuclear protein turnover.
a–c, Transcriptional changes after acute knockout of AKIRIN2 (a) or PSMA3 (b). RNA-seq of iCas9-RKO cells was performed 2 (sgAKIRIN2, sgAAVS1) or 3 days (sgPSMA3) after Cas9 induction (n = 3 biological replicates). Genes significantly up- or downregulated (P ≤ 0.01; Benjamini-Hochberg corrected two-sided Wald-test) at least two-fold are highlighted in orange, TP53 target genes according to20 in red. c, Principal component (PC) analysis of the 1000 most highly expressed genes. d, e, AKIRIN2 and MYC protein half-life quantification. Immunoblot time-series of iCas9-RKO cells treated with cycloheximide (CHX) 2 days after Cas9 induction (d) and half-life quantification (e) of AKIRIN2 (half-life = 46 min, 95% CI = 37–58 min) and MYC (half-life = 25 and 178 min with 95% CI = 21–31 and 51–238 min in sgAAVS1 control and sgAKIRIN2 cells, respectively). Data is shown as mean ± s.d. (n ≥ 3 independent experiments). Dashed lines indicate halflives. f, g, Quantitative proteomics following induced AKIRIN2 or PSMA3 knockout. Samples were obtained as described for a-c (n = 2 biological replicates). f, Principal component analysis of the 1000 most highly expressed proteins. g, Scatter plot of transcriptome- versus proteome-changes upon acute AKIRIN2 knockout compared to sgAAVS1 control. AKIRIN2 targets as in Fig. 3a, b (orange; n = 124) are upregulated only on protein-, but not on mRNA-level. TP53 target genes as in a, b are shown in red. h, Western blot of selected AKIRIN2 targets after AKIRIN2 or proteasome knockout. iCas9-RKO cells expressing the indicated sgRNAs were harvested before, and 2 and 3 days after Cas9 induction. i, Euler diagram of proteasome targets and AKIRIN2 targets as defined in Fig. 3a, b. CI, confidence interval; FC, fold change.
Extended Data Fig. 5 AKIRIN2 is a highly conserved nuclear protein.
a, Alignment of Akirin orthologs from 11 representative model organisms. AKIRIN2 and AKIRIN1 are highly conserved in vertebrates. In invertebrate metazoans, only one Akirin ortholog per species was found, which is more closely related to AKIRIN2 than to AKIRIN1. Colors denote clustal amino acid identity. b–d, Knockout-rescue studies evaluating essentiality of AKIRIN2 protein features as in Fig. 4c, d. b, Schematic setup of knockout-rescue experiments and AKIRIN2 cDNA variants, in which sgAKIRIN2 seed and PAM sequences were removed through synonymous mutations. Roman numbers are continued from Fig. 4c. c, Competitive proliferation assays of iCas9-RKO cells co-expressing sgAKIRIN2 and the indicated AKIRIN2 cDNA variant. Cells were monitored using flow cytometry for 10 days after Cas9 induction. Values are normalized to Day 0. Data is shown as mean ± s.d. (n = 3 independent experiments). d, Immunoblotting of V5 and AKIRIN2 in iCas9-RKO cells expressing the indicated AKIRIN2 cDNA variants. e, Immunofluorescence localization of AKIRIN2. V5-AKIRIN2 knockout-rescue RKO cells were stained with α-V5 antibody. Images are representative of 3 independent experiments. Scale bar, 15 µm. f, Co-IP/MS of full-length V5-AKIRIN1 purified with α-V5 antibody. Enrichment was calculated over V5-NLS-GFP control (Benjamini-Hochberg corrected limma moderated two-sided t-test, n = 6 biological replicates from 2 independent experiments).
Extended Data Fig. 6 AKIRIN2 purification and negative staining EM.
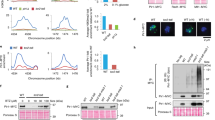
a, Schematic four-step purification of recombinant His-GST-AKIRIN2 from E. coli (left) and size-exclusion chromatogram (right). Blue bars indicate GST-AKIRIN2 containing fractions analyzed by SDS-PAGE. b, SDS-PAGE of eluted fractions. Lower 26 kDa band corresponds to free GST. c, d, Co-IP/MS of sucrose fractionated proteins co-purified with GST-AKIRIN2 from cytosolic HeLa cell lysate. Experimental setup (c) and normalized protein abundance of selected AKIRIN2-interactors across sucrose fractions (d). 20S-CP and 19S-RP datapoints represent the mean of all 20S-CP and 19S-ATPase and non-ATPase subunits as defined in Fig. 2b, respectively. e–h, Negative staining electron microscopy of AKIRIN2-proteasome complexes. Experimental setup (e) and SDS-PAGE (f) of sucrose gradient fractionated proteins co-purified with GST-AKIRIN2 (left) or GST-UBL (right) from PEG-concentrated HeLa cell lysate. Red boxes indicate fractions used for negative staining electron microscopy (g, h). 3D reconstructions (g) and representative negative stain electron micrograph and 2D class averages (h) of 26S proteasome complexes co-purified with GST-AKIRIN2 (left) or GST-UBL (right). Arrows indicate AKIRIN2-specific densities. Data in a–b and f–h are representative of three independent experiments. Scale bar, 50 nm. SEC, size-exclusion chromatography; EM, electron microscopy; CP, core (proteasome) particle; RP, regulatory particle.
Extended Data Fig. 7 Cryo-EM data processing and analysis of the AKIRIN2-proteasome complex.
a, Representative micrograph and 2D class averages of proteasome complexes co-purified with AKIRIN2. Data is representative of three independent experiments. Scale bar, 50 nm. b, Angular distributions of individual particles. c, Fourier Shell Correlation (FSC) curve of the refined EM map at 3.2 Å resolution. d, Cryo-EM density map of AKIRIN2 bound to the 20S proteasome as in Fig. 4g, side view. e, f, Cryo-EM model of AKIRIN2 bound to the 20S proteasome (e) and the substrate-engaged 26S proteasome complex (f). Detail views show the binding sites of AKIRIN2. In the active proteasome conformation, the α3/α4 pocket (top) and α2/α3 pocket (bottom) are occupied by PSMC1 (Rpt2) and PSMC5 (Rpt6), respectively. At a low threshold, the α1/α2 pocket also contains a density that may be attributed to the AKIRIN2 C-terminal motif. 20S subunits are shown in blue, AKIRIN2 in red, PSMC1 and PSMC5 in orange, other 19S subunits in transparent white. 26S structure83 was accessed via PDB ID: 6MSJ. FSC, Fourier shell correlation.
Extended Data Fig. 8 AKIRIN2 is a critical mediator of nuclear proteasome import.
a–c, Engineering of 20S proteasome reporter cells. a, Schematic of vectors (left) used for PSMB4 knockout-rescue (right). iCas9-RKO cells were co-transduced with an sgRNA targeting PSMB4 and a cDNA encoding a FLAG-tagged sgRNA-resistant PSMB4-mCherry fusion protein. sgRNA and cDNA double positive cells were monitored by flow cytometry for 7 days after Cas9 induction. Percentage is normalized to Day 0. b, Immunoblot analysis of single cell-derived clonal PSMB4-mCherry reporter cells and WT iCas9-RKO cells. c, SDS-PAGE and immunoblot of PSMB4 in WT iCas9-RKO (top) and clonal PSMB4-mCherry reporter cells (bottom) after sucrose gradient fractionation. d, e, Representative IF images (d) and quantification (e) of endogenous PSMA5 (blue) levels in iCas9-RKO cells after induced knockout of AKIRIN2 (n = 2,729 cells) or AAVS1 (n = 3,625 cells). f–h, mCherry-PSMD3 19S proteasome reporter as in a–c. i–j, Representative confocal images (i) and quantification (j) of mCherry-PSMD3 signal after induced knockout of AKIRIN2 (n = 6,858 cells) or AAVS1 (n = 7,309 cells). k-n, Nuclear proteasome import (k, l) and MYC protein levels (m, n) after induced IPO9 knockout. Representative confocal images (k) and quantification (l) of PSMB4-mCherry signal 7 days after induced IPO9 (n = 8,055 cells) or AAVS1 (n = 8,336 cells) knockout. Representative flow cytometry histogram of endogenous MYC protein levels 7 days after induced IPO9 knockout (m) and fold change compared to sgRNA- cells (one-sided Welch’s t-test) (n). Bars and whiskers represent mean and standard deviation of 3 biological replicates. Nuclei in e, j, l were segmented based on DNA channel and signal was normalized to the mean nuclear signal in sgAAVS1 cells. Data in e, j, l is shown as mean ± s.d. (n = 9 biological replicates from 3 independent experiments; two-sided Welch’s -test). Data in c, h is representative of 2 independent experiments. Scale bars, 15 µm. FC, fold change.
Extended Data Fig. 9 AKIRIN2 controls the nuclear re-import of proteasomes following mitosis.
a, Quantification of nuclear PSMB4-mCherry signal in individual dividing iCas9-RKO reporter cells after AKIRIN2 (n = 38 cells) or AAVS1 knockout (n = 27 cells) as shown in Fig. 5d over time. b–d, Time-lapse imaging of mitotic PSMB4-mCherry RKO reporter cells co-expressing IBB-GFP. b, Quantification of nuclear PSMB4-mCherry and IBB-GFP signal in dividing reporter cells shown as averaged data (b, mean ± 95% CI, n = 27 cells from two independent experiments), individual cell-tracks (c), and representative time-lapse images (d). Dashed line indicates the onset of IBB-GFP import. DNA was visualized with SiR-Hoechst. Signal intensity is normalized to t = −100 min. Scale bar, 10 µm. e, Quantification of MYC-mCherry signal as in a after AKIRIN2 (n = 40 cells) or AAVS1 knockout (n = 38 cells) as shown in Fig. 5f. f, Model of post-mitotic nuclear proteasome import mediated by AKIRIN2. Cells undergoing mitotic cell division upon loss of AKIRIN2 give rise to daughter cells that are devoid of nuclear proteasomes. Signal in a, e is normalized to mean nuclear signal in sgAAVS1 cells. Pseudotime is normalized to the timepoint of maximal chromatin condensation (a, e) or to the onset of nuclear PSMB4-mCherry import (b, c).
Supplementary information
Supplementary Figs.
This file contains Supplementary Figs. 1–3. Supplementary Fig. 1 contains uncropped images of western blots and SDS gels. Supplementary Fig. 2 shows representative gating strategies for FACS-based MYC regulator screens and flow cytometric analyses. Supplementary Fig. 3 schematically shows cryo electron microscopy image processing and model building.
Supplementary Table 1
Raw sgRNA counts of MYC regulator screens. For each screen condition, raw sequencing reads for individual sgRNAs are provided. Reads are not normalized to sequencing depth. Column names are provided in the format cellline_FACSgate_timepoint_replicate.
Supplementary Table 2
MAGeCK analysis of MYC regulator screens. Gene-level average, median-normalized log2 fold changes, one-tailed logarithmized P values and FDRs calculated by MAGeCK v0.5.9. The table contains the merged dataset representing the minimum P value of both time points, as well as individual datasets for each time point. GO terms are annotated, and genes highlighted and categorized in Fig. 1 and Fig. 2 are indicated. Column names are provided in the format variable_cellline_FACSgate_timepoint.
Supplementary Table 3
Quantitative proteomics and RNA-seq results. Combined table of TMT mass spectrometry and RNA-seq results including normalized protein abundance and mRNA TPM averaged over two or three replicates, respectively, as well as log2 fold changes and logarithmized P values (Benjamini–Hochberg-corrected two-tailed limma moderated t-test and Wald test, respectively). Classification into AKIRIN2-responsive and -independent proteasome targets according to Fig. 3 is annotated.
Supplementary Table 4
GO term enrichment analysis of proteasome targets. Results of GO term enrichment analysis of AKIRIN2-reponsive and -independent proteasome targets using PANTHER two-tailed Fisher’s exact over-representation test with Benjamini–Hochberg multiple testing correction on GO database (release 2020-06-01). Ratio of FDRs (ΔFDR) for AKIRIN2-responsive and -independent genes for each GO term is provided.
Supplementary Table 5
Akirin orthologues. Gene name, species name, primary accession number and protein length in amino acids for 103 Akirin orthologues from 77 different species used for phylogenetic analyses. Consideration of individual orthologues for the different analyses and figures are annotated.
Supplementary Table 6
AKIRIN2 co-immunoprecipitation mass spectrometry results. Normalized, imputed protein abundances identified by co-immunoprecipitation mass spectrometry analysis of V5-AKIRIN2, V5-AKIRIN1, V5-AKIRIN2ΔYVS and V5-GFP in RKO cells averaged over six replicates, including log2 fold changes and logarithmized P values (Benjamini–Hochberg-corrected two-tailed limma moderated t-test) as well as mass spectrometric quantification of proteins co-purified with GSTAKIRIN2 from HeLa cell extract after fractionation on a 10–30% sucrose gradient.
Supplementary Table 7
Proteasome subunit nomenclature. Human gene name, common protein nomenclature and chain ID and protein nomenclature used in cryo-EM model of AKIRIN2 bound to the 20S proteasome.
Supplementary Table 8
EM model building statistics. EM data collection and processing statistics for negative stain EM structure of AKIRIN2 bound to the 26S proteasome and data collection, processing, refinement and validation statistics for cryo-EM structure of AKIRIN2 bound to the 20S proteasome.
Supplementary Table 9
Glossary. Abbreviations used in the manuscript and their definitions.
Supplementary Video 1
Live-cell imaging of mCherry–MYC reporter cells. Movie highlights kinetics of MYC stabilization after inducible knockout of AKIRIN2, FBXW7 or the 20S proteasome subunit PSMA1. Cells were imaged every 2 h for 96 h. Scale bar, 100 μm.
Supplementary Video 2
Confocal live-cell imaging of mitotic RKO reporter cells after inducible AKIRIN2/AAVS1 knockout. mCherryMYC or PSMB4mCherry reporter cells were imaged 15–24 h after induction of AKIRIN2/AAVS1 knockout and imaged at 5- or 10-min intervals as indicated. mCherry–MYC is shown in red, PSMB4mCherry in blue and DNA in grey. Scale bar, 10 μm (single cell) and 50 μm (overview, 10 h).
Rights and permissions
About this article
Cite this article
de Almeida, M., Hinterndorfer, M., Brunner, H. et al. AKIRIN2 controls the nuclear import of proteasomes in vertebrates. Nature 599, 491–496 (2021). https://doi.org/10.1038/s41586-021-04035-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-021-04035-8
This article is cited by
-
Functionalizing tandem mass tags for streamlining click-based quantitative chemoproteomics
Communications Chemistry (2024)
-
Discovery of a Drug-like, Natural Product-Inspired DCAF11 Ligand Chemotype
Nature Communications (2023)
-
ETV6 dependency in Ewing sarcoma by antagonism of EWS-FLI1-mediated enhancer activation
Nature Cell Biology (2023)
-
Gene Modulation with CRISPR-based Tools in Human iPSC-Cardiomyocytes
Stem Cell Reviews and Reports (2023)
-
Direct control of lysosomal catabolic activity by mTORC1 through regulation of V-ATPase assembly
Nature Communications (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.