Abstract
The balance between bacterial colonization and its containment in the intestine is indispensable for the symbiotic relationship between humans and their bacteria. One component to maintain homeostasis at the mucosal surfaces is immunoglobulin A (IgA), the most abundant immunoglobulin in mammals1,2. Several studies have revealed important characteristics of poly-reactive IgA3,4, which is produced naturally without commensal bacteria. Considering the dynamic changes within the gut environment, however, it remains uncertain how the commensal-reactive IgA pool is shaped and how such IgA affects the microbial community. Here we show that acetate—one of the major gut microbial metabolites—not only increases the production of IgA in the colon, but also alters the capacity of the IgA pool to bind to specific microorganisms including Enterobacterales. Induction of commensal-reactive IgA and changes in the IgA repertoire by acetate were observed in mice monocolonized with Escherichia coli, which belongs to Enterobacterales, but not with the major commensal Bacteroides thetaiotaomicron, which suggests that acetate directs selective IgA binding to certain microorganisms. Mechanistically, acetate orchestrated the interactions between epithelial and immune cells, induced microbially stimulated CD4 T cells to support T-cell-dependent IgA production and, as a consequence, altered the localization of these bacteria within the colon. Collectively, we identified a role for gut microbial metabolites in the regulation of differential IgA production to maintain mucosal homeostasis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
The raw 16S rRNA gene sequencing and IgA repertoire sequencing data are deposited in the DNA Data Bank of Japan, under NCBI BioProject accession numbers PRJDB7422 and PRJDB7423. Taxonomic assignment of 16S rRNA gene sequencing data was determined with the Greengenes database v.13.5 (http://greengenes.secondgenome.com/)36. IgA repertoire was analysed using the IMGT database (http://www.imgt.org/)43. Source data are provided with this paper.
References
Lycke, N. Y. & Bemark, M. The regulation of gut mucosal IgA B-cell responses: recent developments. Mucosal Immunol. 10, 1361–1374 (2017).
Pabst, O., Cerovic, V. & Hornef, M. Secretory IgA in the coordination of establishment and maintenance of the microbiota. Trends Immunol. 37, 287–296 (2016).
Okai, S. et al. High-affinity monoclonal IgA regulates gut microbiota and prevents colitis in mice. Nat. Microbiol. 1, 16103 (2016).
Bunker, J. J. et al. Natural polyreactive IgA antibodies coat the intestinal microbiota. Science 358, eaan6619 (2017).
Kim, M., Qie, Y., Park, J. & Kim, C. H. Gut microbial metabolites fuel host antibody responses. Cell Host Microbe 20, 202–214 (2016).
Wu, W. et al. Microbiota metabolite short-chain fatty acid acetate promotes intestinal IgA response to microbiota which is mediated by GPR43. Mucosal Immunol. 10, 946–956 (2017).
Goverse, G. et al. Diet-derived short chain fatty acids stimulate intestinal epithelial cells to induce mucosal tolerogenic dendritic cells. J. Immunol. 198, 2172–2181 (2017).
Kato, T. et al. Multiple omics uncovers host-gut microbial mutualism during prebiotic fructooligosaccharide supplementation. DNA Res. 21, 469–480 (2014).
Hara, S. et al. Dietary antigens induce germinal center responses in Peyer’s patches and antigen-specific IgA production. Front. Immunol. 10, 2432 (2019).
Palm, N. W. et al. Immunoglobulin A coating identifies colitogenic bacteria in inflammatory bowel disease. Cell 158, 1000–1010 (2014).
Hapfelmeier, S. et al. Reversible microbial colonization of germ-free mice reveals the dynamics of IgA immune responses. Science 328, 1705–1709 (2010).
Rosner, K. et al. Third complementarity-determining region of mutated VH immunoglobulin genes contains shorter V, D, J, P, and N components than non-mutated genes. Immunology 103, 179–187 (2001).
Yasuda, K. et al. Biogeography of the intestinal mucosal and lumenal microbiome in the rhesus macaque. Cell Host Microbe 17, 385–391 (2015).
Reboldi, A. et al. IgA production requires B cell interaction with subepithelial dendritic cells in Peyer’s patches. Science 352, aaf4822 (2016).
Lin, Y. L., Ip, P. P. & Liao, F. CCR6 deficiency impairs IgA production and dysregulates antimicrobial peptide production, altering the intestinal flora. Front. Immunol. 8, 805 (2017).
Baptista, A. P. et al. Colonic patch and colonic SILT development are independent and differentially regulated events. Mucosal Immunol. 6, 511–521 (2013).
Lee, A. Y. S. et al. Expression of membrane-bound CC Chemokine ligand 20 on follicular T helper cells in T–B-cell conjugates. Front. Immunol. 8, 1871 (2017).
Kubinak, J. L. et al. MyD88 signaling in T cells directs IgA-mediated control of the microbiota to promote health. Cell Host Microbe 17, 153–163 (2015).
Vatanen, T. et al. Variation in microbiome LPS immunogenicity contributes to autoimmunity in humans. Cell 165, 842–853 (2016).
Preite, S. et al. Somatic mutations and affinity maturation are impaired by excessive numbers of T follicular helper cells and restored by Treg cells or memory T cells. Eur. J. Immunol. 45, 3010–3021 (2015).
Kau, A. L. et al. Functional characterization of IgA-targeted bacterial taxa from undernourished Malawian children that produce diet-dependent enteropathy. Sci. Transl. Med. 7, 276ra24 (2015).
Sugahara, H. et al. Decreased taxon-specific IgA response in relation to the changes of gut microbiota composition in the elderly. Front. Microbiol. 8, 1757 (2017).
Ragonnaud, E. & Biragyn, A. Gut microbiota as the key controllers of “healthy” aging of elderly people. Immun. Ageing 18, 2 (2021).
Kono, H., Hashimoto, H. & Shimizu, Y. NMR characterization of cellulose acetate: chemical shift assignments, substituent effects, and chemical shift additivity. Carbohydr. Polym. 118, 91–100 (2015).
Kamide, K., Saito, M. & Abe, T. Dilute solution properties of water-solube incompletely substituted cellulose acetate. Polym. J. 13, 421–431 (1981).
Shimamoto, S., Kohmoto, T. & Shibata, T. in Cellulose Derivatives: Modification, Characterization, and Nanostructures ACS Symposium Series Vol. 688, Ch. 14, 194–200 (ASC Publications, 1998).
Tezuka, Y. & Tsuchiya, Y. Determination of substituent distribution in cellulose acetate by means of a 13C NMR study on its propanoated derivative. Carbohydr. Res. 273, 83–91 (1995).
Clausen, M. R., Bonnén, H., Tvede, M. & Mortensen, P. B. Colonic fermentation to short-chain fatty acids is decreased in antibiotic-associated diarrhea. Gastroenterology 101, 1497–1504 (1991).
Kucharzik, T., Hudson, J. T. III, Waikel, R. L., Martin, W. D. & Williams, I. R. CCR6 expression distinguishes mouse myeloid and lymphoid dendritic cell subsets: demonstration using a CCR6 EGFP knock-in mouse. Eur. J. Immunol. 32, 104–112 (2002).
Kimura, I. et al. The gut microbiota suppresses insulin-mediated fat accumulation via the short-chain fatty acid receptor GPR43. Nat. Commun. 4, 1829 (2013).
Malissen, M. et al. Altered T cell development in mice with a targeted mutation of the CD3-epsilon gene. EMBO J. 14, 4641–4653 (1995).
Muramatsu, M. et al. Class switch recombination and hypermutation require activation-induced cytidine deaminase (AID), a potential RNA editing enzyme. Cell 102, 553–563 (2000).
Hase, K. et al. Uptake through glycoprotein 2 of FimH+ bacteria by M cells initiates mucosal immune response. Nature 462, 226–230 (2009).
Kozich, J. J., Westcott, S. L., Baxter, N. T., Highlander, S. K. & Schloss, P. D. Development of a dual-index sequencing strategy and curation pipeline for analyzing amplicon sequence data on the MiSeq Illumina sequencing platform. Appl. Environ. Microbiol. 79, 5112–5120 (2013).
Caporaso, J. G. et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 7, 335–336 (2010).
DeSantis, T. Z. et al. Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. Appl. Environ. Microbiol. 72, 5069–5072 (2006).
Osman, N., Adawi, D., Ahrné, S., Jeppsson, B. & Molin, G. Probiotics and blueberry attenuate the severity of dextran sulfate sodium (DSS)-induced colitis. Dig. Dis. Sci. 53, 2464–2473 (2008).
Furusawa, Y. et al. Commensal microbe-derived butyrate induces the differentiation of colonic regulatory T cells. Nature 504, 446–450 (2013).
Gong, J. et al. Diversity and phylogenetic analysis of bacteria in the mucosa of chicken ceca and comparison with bacteria in the cecal lumen. FEMS Microbiol. Lett. 208, 1–7 (2002).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Vicente-Suarez, I. et al. Unique lamina propria stromal cells imprint the functional phenotype of mucosal dendritic cells. Mucosal Immunol. 8, 141–151 (2015).
Kanaya, T. et al. Development of intestinal M cells and follicle-associated epithelium is regulated by TRAF6-mediated NF-κB signaling. J. Exp. Med. 215, 501–519 (2018).
Lefranc, M. P. et al. IMGT, the international ImMunoGeneTics information system. Nucleic Acids Res. 37, D1006–D1012 (2009).
Sato, T. et al. Single Lgr5 stem cells build crypt-villus structures in vitro without a mesenchymal niche. Nature 459, 262–265 (2009).
Ootani, A. et al. Sustained in vitro intestinal epithelial culture within a Wnt-dependent stem cell niche. Nat. Med. 15, 701–706 (2009).
Moon, C., VanDussen, K. L., Miyoshi, H. & Stappenbeck, T. S. Development of a primary mouse intestinal epithelial cell monolayer culture system to evaluate factors that modulate IgA transcytosis. Mucosal Immunol. 7, 818–828 (2014).
Ettayebi, K. et al. Replication of human noroviruses in stem cell-derived human enteroids. Science 353, 1387–1393 (2016).
Acknowledgements
We thank N. Atarashi, M. Kawasumi, S. Onawa, N. Tachibana and A. Ito for their technical support; B. Malissen and T. Honjo for providing Cd3e−/− and Aicda−/− mice, respectively; C. Kuo for providing the R-spondin-1-producing cell line; and S. Koyasu and P. D. Burrows for critical reading and English editing of the manuscript. This study was supported in part by research grants from RIKEN Interdisciplinary Research Program ‘Integrated Symbiology’, RIKEN Pioneering Project ‘Biology of Symbiosis’, Grants-in-Aid for Young Scientists (B) (26850090 to E.M.), AMED-CREST (JP19gm0710009 to H.O.) and Daicel Corporation. T. Takeuchi was supported by RIKEN Junior Research Associate Program.
Author information
Authors and Affiliations
Contributions
T. Takeuchi and H.O. conceived the study. T. Takeuchi designed and performed the experiments and analyses and co-wrote the manuscript. E.M. contributed to the data analyses and discussions and co-wrote the manuscript. T. Kanaya contributed to fluorescence immunohistochemistry and in vitro experiments. T. Kato analysed the human faecal data and performed 16S rRNA gene sequencing. Y.N. quantified the SCFA concentration in the intestines using gas chromatography–tandem mass spectrometry. T.W., T.S. and O.O. contributed to the analyses of the IgA repertoire. T. Taida and H.N. contributed to the animal experiments and FACS analyses. T. Kitami contributed to the immune cell metabolism measurements. S.S. and A.M. developed SCFA-conjugated cellulose. I.K. and I.R.W. provided essential materials and helped to interpret the data. H.O. directed the research, provided essential materials and co-wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
S.S. and A.M. are employees of Daicel Corporation; T. Takeuchi, E.M., S.S., A.M. and H.O. have applied for a patent regarding SCFA-conjugated cellulose; and H.O. received research funds from Daicel Corporation. Otherwise the authors have no competing interests.
Additional information
Peer review information Nature thanks Andrea Cerutti and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Faecal SCFAs are associated with IgA levels in mice and humans.
a, Correlation of total major SCFAs, acetate, propionate and butyrate with SIgA concentration in faeces of antibiotic-treated mice. Mice were administered the following antibiotics via ad libitum drinking water: vancomycin (orange), neomycin (red), ampicillin (aqua), metronidazole (purple) and distilled water (grey) for 2 weeks (n = 4 per group). Faecal samples were sequentially collected at weeks 1 and 2. Total major SCFAs is the sum of acetate, propionate and butyrate. b, Association of human faecal metabolites with SIgA concentration (n = 17 per group). All faecal samples were dichotomized according to the IgA concentration and the relative intensity of specified metabolites was compared between the IgA-low and IgA-high groups. c, Relative intensity of faecal acetate, propionate and butyrate was compared between the IgA-low and high-groups. d–f, Ileal, caecal and colonic concentrations of acetate (d), propionate (e) and butyrate (f) in mice fed the fibre-deprived cWSCA (control)-containing or WSCA-containing diet (n = 4 per group (d), n = 4 per group (e) and n = 4 versus 3 (f)). g, h, Faecal concentration of SCFAs in mice fed the cWSCP (control) or WSCP diet (g), and cWSCB (control) or WSCB diet (h) (n = 4 per group). i, Faecal concentration of SCFAs in mice fed a standard chow diet (CE2, CLEA Japan) (n = 8). c, Box plots indicate the median, upper and lower quartiles, and upper and lower extremes except for outliers. d–i, Data are mean and s.d. *P < 0.05, **P < 0.01, ***P < 0.001; Spearman’s rank-order correlation (a), two-sided Wilcoxon rank-sum test (c) and Kruskal–Wallis test with Dunn’s test (d–i). Exact P values are provided in the Source Data.
Extended Data Fig. 2 Acetate, but not other SCFAs, increases colonic IgA production and SIgA binding of commensal bacteria.
a, Representative histograms showing the patterns of faecal SIgA binding to faecal DAPI+ bacteria. b, The IgA concentration in different parts of the intestine was analysed using ELISA (n = 4 per group). Small intestines were divided into duodenum, jejunum and ileum, and large intestines were divided into caecum, proximal and distal colon. c, d, Representative flow cytometry plots of colonic IgA-producing plasma cells (defined as IgA+B220−CD3ε− cells, gated on CD3ε− lymphocytes) (c) and the frequency among colonic lymphocytes and the absolute number (d) (n = 10 per group). e, The frequency of IgA-producing plasma cells among small intestinal lymphocytes and the absolute number (n = 5 per group). f–h, Faecal SIgA concentration at week 4 after the start of the modified diet (f), colonic IgA-producing plasma cells (g) and faecal SIgA-coated (SIgA+) bacteria (h) in mice fed cWSCP (control) or WSCP (n = 4 per group). i–k, Faecal SIgA concentration at week 4 (i), colonic IgA-producing plasma cells (j) and faecal SIgA+ bacteria (k) in mice fed cWSCB (control) or WSCB (n = 4 per group (i, j); n = 3 versus 4 (k)). l, Weighted UniFrac distances between faecal SIgA− and SIgA+ bacteria in the cWSCA (control) or WSCA groups. The distances between all samples within groups are indicated as a reference (n = 4 versus 5). m, The relative abundance of faecal SIgA− and SIgA+ bacteria at the phylum level (n = 4 per group). n, The relative abundance of faecal SIgA− and SIgA+ Bifidobacteriales, Bacteroidales, Erysipelotrichales and Enterobacterales (n = 4 per group). Data are mean and s.d. *P < 0.05, **P < 0.01, ***P < 0.001; Kruskal–Wallis test with Dunn’s test (b, l, n) and two-sided Wilcoxon rank-sum test (d–k). Pooled data from two independent experiments (d). Exact P values are provided in the Source Data.
Extended Data Fig. 3 Acetate increases IgA production in E. coli-monocolonized mice and germ-free mice orally administered heat-killed E. coli.
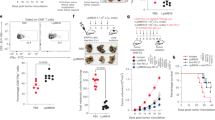
a, Representative flow cytometry plots (gated on DAPI+ population) and summary data of caecal SIgA+ bacteria in control-diet-fed and WSCA-diet-fed mice monocolonized with B. thetaiotaomicron (n = 9 per group) or E. coli (n = 9 versus 10, respectively). b, Representative flow cytometry plots (gated on lymphocytes) and summary data of colonic IgA+B220− plasma cells (PC) and IgA+B220+ B cells (BC) in mice monocolonized with B. thetaiotaomicron (n = 9 per group) or E. coli (n = 9 versus 10). c, Abundance of E. coli in E. coli-monocolonized mice measured as CFU per gram (CFU/g) of colonic contents (n = 5 per group). d, e, Representative flow cytometry plots (gated on CD3ε− lymphocytes) depicting colonic IgA+ plasma cells and B cells in germ-free mice (d) and the absolute number (e) (n = 3 versus 4). f, Faecal SIgA concentration at week 4 in germ-free mice (n = 8 per group). g, Microorganism-reactive SIgA in the caecal contents from control-diet-fed and WSCA-diet-fed germ-free mice orally administered heat-killed B. thetaiotaomicron (n = 4 versus 5, respectively) or E. coli (n = 9 per group). The OD450 in the WSCA diet groups was normalized to that of the cWSCA (control) diet groups. h, The frequency among colonic lymphocytes and the absolute number of colonic IgA-producing plasma cells in control-diet-fed and WSCA-diet-fed germ-free mice orally administered heat-killed B. thetaiotaomicron (n = 4 versus 5, respectively) or E. coli (n = 4 per group). i–k, Skewness (i), kurtosis (j) and inverse Simpson’s index (k) of HCDR3 amino acid length distribution in colonic IgA-producing B cells of control-diet-fed and WSCA-diet-fed mice monocolonized with B. thetaiotaomicron (n = 5 per group) or E. coli (n = 4 versus 5, respectively). l, HCDR3 amino acid sequence distribution in control-diet-fed and WSCA-diet-fed mice monocolonized with B. thetaiotaomicron (n = 5 per group) or E. coli (n = 4 versus 5, respectively). Each column corresponds to the sequencing results from a single mouse and the colour corresponds to the order of frequency. m, The frequency of IgA+ B cell clones expressing the most-dominant HCDR3 amino acid sequence in control-diet-fed and WSCA-diet-fed mice monocolonized with B. thetaiotaomicron (n = 5 per group) or E. coli (n = 4 versus 5, respectively). n, Mode (that is, the value that appears the most frequently in a sample) of HCDR3 amino acid length in control-diet-fed and WSCA-diet-fed mice monocolonized with B. thetaiotaomicron (n = 5 per group) or E. coli (n = 4 versus 5, respectively). b, c, e, f, Data are mean and s.d. a, g–k, m, n, Box plots indicate median, upper and lower quartiles, and upper and lower extremes except for outliers. *P < 0.05, **P < 0.01; two-sided Wilcoxon rank-sum test (a–c, e–g, i–k, m, n) and Kruskal–Wallis test with Dunn’s test (h). a, b, f, g, Pooled data from two independent experiments. Exact P values are provided in the Source Data.
Extended Data Fig. 4 Acetate alters the composition of mucosa-associated bacteria in an IgA-dependent manner.
a, Ratio of mucosa-associated to luminal bacteria at the order level in wild-type mice and Aicda−/− mice depicted on a log10 scale. Each row represents the sequencing results from a single mouse. Results of unsupervised clustering based on Ward’s method are also shown. The taxa in bold font were significantly altered in terms of their localization upon WSCA diet administration. b, The relative abundance of gut bacteria at the phylum level in the colonic luminal contents and mucus layer of wild-type and Aicda−/− mice. c, Schematic of the experimental protocol. Mice fed cWSCA (control) or WSCA were orally administered 1 × 109 CFU of E. coli to enhance IgA production. On the day of assessment, 1 × 109 CFU of GFP-expressing E. coli were orally administered and, 3 h later, the colonic tissues and contents were sampled. d–g, The frequency of total GFP-expressing E. coli (d), SIgA− (e) and SIgA+ (f) GFP-expressing E. coli, and SIgA+ GFP-negative bacteria (g) in the colonic contents (n = 5 versus 6). d–g, Data are mean and s.d. *P < 0.05; two-sided Wilcoxon rank-sum test (a, d–g). P values were corrected by FDR (a). N.S., not significant. n = 4 per group for wild-type mice and n = 5 per group for Aicda−/− mice (a, b). Exact P values are provided in the Source Data.
Extended Data Fig. 5 Acetate increases germinal-centre B cells in MLNs and the colon.
a, Representative flow cytometry plots (gated on B220+ lymphocytes) depicting GL7+Fas+ germinal-centre B cells in MLNs. b, Frequency among B220+ B cells and the absolute number of GL7+Fas+ germinal-centre B cells (n = 5 per group). c, Representative flow cytometry plots (gated on B220+ lymphocytes) depicting GL7+Fas+ germinal-centre B cells in the colon. d, Frequency among B220+ B cells and the absolute number of GL7+Fas+ germinal-centre B cells (n = 15 per group). b, d, Data are mean and s.d. *P < 0.05, **P < 0.01; two-sided Wilcoxon rank-sum test (b, d). Pooled data from three independent experiments (d). Exact P values are provided in the Source Data.
Extended Data Fig. 6 Acetate increases colonic IgA production and alters IgA reactivity in a CCR6–CCL20-axis-dependent manner.
a, The frequency among colonic lymphocytes of IgA-producing plasma cells and faecal SIgA concentration in Ccr6GFP/+ (hetero) and Ccr6GFP/GFP (KO) mice. b, Faecal SIgA reactive to B. thetaiotaomicron and to E. coli in Ccr6GFP/+ and Ccr6GFP/GFP mice. c, A principal coordinate analysis plot of weighted UniFrac distances for faecal SIgA− and SIgA+ bacteria in Ccr6GFP/GFP mice (n = 4 per group). The dashed and solid lines indicate 90% confidence levels for the SIgA− and SIgA+ bacteria groups, respectively. d, The relative abundance of faecal SIgA− and SIgA+ bacteria at the phylum level in Ccr6GFP/GFP mice (n = 4 per group). e, The relative abundance of faecal SIgA− and SIgA+ Bacteroidales, Erysipelotrichales and Proteobacteria in Ccr6GFP/GFP mice (n = 4 per group). f, The frequency of faecal SIgA+ bacteria in Ccr6GFP/GFP mice monocolonized with E. coli (n = 8 per group). g, Ccl20 mRNA expression in colonic epithelial cells of wild-type mice (n = 5 versus 6). h, Ccl20 mRNA expression in a monolayer form of mouse colonic organoids stimulated with or without sodium acetate (n = 5 per group). a, b, e–h, Data are mean and s.d. *P < 0.05, **P < 0.01; Kruskal–Wallis test with Dunn’s test (a, b, e) or two-sided Wilcoxon rank-sum test (f–h). n = 6 per group for Ccr6GFP/+ mice and n = 5 versus 4 for Ccr6GFP/GFP mice (a, b). Pooled data from two independent experiments (f). Exact P values are provided in the Source Data.
Extended Data Fig. 7 Acetate increases CCR6-expressing TFH-like cells in the colon.
a, Representative flow cytometry plots (gated on CD3ε+CD4+ lymphocytes) depicting CCR6+ T cells and CCR6+CXCR5+ TFH-like cells in the colon of Ccr6GFP/+ mice. b, c, The frequency among CD3ε+CD4+ T cells and the absolute number of CCR6+ T cells (b) and CCR6+CXCR5+ TFH-like cells (c) in the colon of Ccr6GFP/+ mice (n = 6 per group). d, Representative flow cytometry plots (gated on CD19−CD3ε+CD4+ lymphocytes) depicting CXCR5+PD-1+ TFH-like cells in the colon of wild-type mice. e, f, The frequency among CD3ε+CD4+ T cells and the absolute number of CXCR5+PD-1+ TFH-like cells (e) and RORγt+ TH17 cells (f) in the colon of wild-type mice (n = 5 per group). g, The frequency among CD3ε−CD19− lineage marker (lin) negative cells and the absolute number of RORγt+ innate lymphoid cells (ILCs) in the colon of wild-type mice (n = 5 per group). Data are mean and s.d. (b, c, e–g). *P < 0.05, **P < 0.01; two-sided Wilcoxon rank-sum test (b, c, e–g). Exact P values are provided in the Source Data.
Extended Data Fig. 8 Acetate and E. coli synergistically regulate T cell functions to facilitate IgA production ex vivo and in vitro.
a, Bcl6 mRNA expression in CD4 T cells sorted from MLN cells co-stimulated ex vivo with heat-killed bacteria and 5 mM sodium acetate for 4 h (n = 3 per group). b, IgA concentrations in culture supernatants of B cells with CD11c+ cells and with (n = 5 per group) or without (n = 3 versus 4) CD4 T cells, and co-stimulated with heat-killed bacteria and sodium acetate for 7 days. c, IgA concentrations in culture supernatants of CD19+IgD+ B cells and CD11c+ cells with CD4 T cells from wild-type mice, and co-stimulated with 0.1 μg ml−1 of LPS and sodium acetate for 7 days (n = 10 per group). d, IgA concentrations in culture supernatants of CD19+IgD+ B cells and CD11c+ cells with CD4 T cells from Myd88−/−Ticam1−/− mice, and co-stimulated with 0.1 μg ml−1 of LPS and sodium acetate for 7 days (n = 5 per group). e, Aldh1a2 mRNA expression in CD11c+ cells sorted from MLN cells co-stimulated ex vivo with heat-killed bacteria and 5 mM sodium acetate for 4 h (n = 3 per group). a–e, Data are mean and s.d. *P < 0.05, **P < 0.01, ***P < 0.001; two-way ANOVA with Tukey’s test (a, e) and Kruskal–Wallis test with Dunn’s test (b–d). Pooled data from two independent experiments (c). Exact P values are provided in the Source Data.
Extended Data Fig. 9 Acetate does not increase colonic IgA production or SIgA binding of Enterobacterales in T-cell-deficient mice.
a, b, Representative flow cytometry plots (gated on DAPI+ population) illustrating faecal SIgA+ bacteria in Cd3e−/− mice (a) and summary data (b) (n = 4 per group). c, Faecal SIgA concentration in Cd3e−/− mice (n = 6 per group). d, Representative flow cytometry plots (gated on lymphocytes) depicting colonic IgA+B220− plasma cells in Cd3e−/− mice. e, f, The frequency among colonic lymphocytes (e) and the absolute number (f) of IgA-producing plasma cells (n = 6 per group). g, Representative flow cytometry plots (gated on B220+ lymphocytes) depicting GL7+Fas+ germinal-centre B cells in MLNs of Cd3e−/− mice. h, i, The frequency of germinal-centre B cells among MLN lymphocytes (h) and the absolute number in Cd3e−/− mice (i) (n = 5 per group). j, A principal coordinate analysis (PCoA) plot of weighted UniFrac distances for faecal SIgA− and SIgA+ bacteria in Cd3e−/− mice (n = 6 per group). The dashed and solid lines indicate 90% confidence levels for the SIgA− and SIgA+ bacteria groups, respectively. k, The relative abundance of faecal SIgA− and SIgA+ bacteria at the phylum level in Cd3e−/− mice (n = 6 per group). l–n, The relative abundance of faecal SIgA− and SIgA+ Bacteroidales (l), Erysipelotrichales (m) and Enterobacterales (n) in Cd3e−/− mice (n = 6 per group). o, A principal coordinate analysis plot of weighted UniFrac distances for luminal and mucosa-associated bacteria in Cd3e−/− mice (n = 6 per group). The dashed and solid lines indicate 90% confidence levels for the luminal and mucosa-associated bacteria groups, respectively. p, The relative abundance of gut bacteria at the phylum level in the colonic luminal contents and mucus layer of Cd3e−/− mice (n = 6 per group). q, Ratio of mucosa-associated to luminal bacteria at the order level in Cd3e−/− mice depicted on a log10 scale, with unsupervised clustering based on Ward’s method (n = 6 per group). The taxa highlighted in bold were significantly altered in terms of their localization upon WSCA diet administration. b, c, e, f, h, i, l–n, Data are mean and s.d. *P < 0.05, ***P < 0.001; two-sided Wilcoxon rank-sum test (b, e, f, h, i, q), two-way ANOVA with Tukey’s test (c) and Kruskal–Wallis test with Dunn’s test (l–n). P values were corrected by FDR (q). Exact P values are provided in the Source Data.
Extended Data Fig. 10 Toll-like receptor signalling in CD4 T cells is necessary for acetate to increase colonic IgA production.
a, Microorganism-reactive SIgA in the caecal contents from Cd3e−/− mice transferred with wild-type (n = 9 per group) or Myd88−/−Ticam1−/− (DKO; n = 8 versus 9) CD4 T cells. b, The absolute number of colonic CD4 T cells in Cd3e−/− mice transferred with wild-type (n = 9 per group) or Myd88−/−Ticam1−/− (n = 8 versus 9) CD4 T cells. c, d, The frequency of caecal SIgA+ bacteria (c) and the absolute number of colonic IgA-producing plasma cells (d) in Cd3e−/− mice monocolonized with E. coli. The mice were transferred with CD4 T cells from either wild-type (n = 4 versus 6) or Myd88−/−Ticam1−/− (n = 4 per group) mice two to three weeks before the dietary intervention. e, f, The frequency of faecal SIgA+ bacteria (e) and the absolute number of colonic IgA-producing plasma cells (f) in Gpr43+/− (n = 5 per group) and Gpr43−/− (n = 5 versus 6) mice. Data are mean and s.d. (a–f). *P < 0.05, **P < 0.01, ***P < 0.001; Kruskal–Wallis test with Dunn’s test (a–f). Pooled data from two independent experiments (a, c, d). Exact P values are provided in the Source Data.
Supplementary information
Supplementary Figure 1
This file contains Supplementary Figure 1 and the accompanying legend for Supplementary Figure 1.
Source data
Rights and permissions
About this article
Cite this article
Takeuchi, T., Miyauchi, E., Kanaya, T. et al. Acetate differentially regulates IgA reactivity to commensal bacteria. Nature 595, 560–564 (2021). https://doi.org/10.1038/s41586-021-03727-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-021-03727-5
This article is cited by
-
Gut microbiota as a target in the bone health of livestock and poultry: roles of short-chain fatty acids
Animal Diseases (2023)
-
Colonization and development of the gut microbiome in calves
Journal of Animal Science and Biotechnology (2023)
-
Gut microbial carbohydrate metabolism contributes to insulin resistance
Nature (2023)
-
Polymeric immunoglobulin receptor deficiency exacerbates autoimmune hepatitis by inducing intestinal dysbiosis and barrier dysfunction
Cell Death & Disease (2023)
-
Bacterial induction of B cell senescence promotes age-related changes in the gut microbiota
Nature Cell Biology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.