Abstract
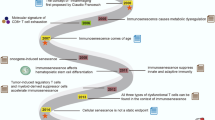
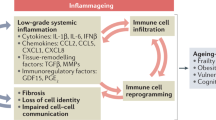
Ageing of the immune system, or immunosenescence, contributes to the morbidity and mortality of the elderly1,2. To define the contribution of immune system ageing to organism ageing, here we selectively deleted Ercc1, which encodes a crucial DNA repair protein3,4, in mouse haematopoietic cells to increase the burden of endogenous DNA damage and thereby senescence5,6,7 in the immune system only. We show that Vav-iCre+/−;Ercc1−/fl mice were healthy into adulthood, then displayed premature onset of immunosenescence characterized by attrition and senescence of specific immune cell populations and impaired immune function, similar to changes that occur during ageing in wild-type mice8,9,10. Notably, non-lymphoid organs also showed increased senescence and damage, which suggests that senescent, aged immune cells can promote systemic ageing. The transplantation of splenocytes from Vav-iCre+/−;Ercc1−/fl or aged wild-type mice into young mice induced senescence in trans, whereas the transplantation of young immune cells attenuated senescence. The treatment of Vav-iCre+/−;Ercc1−/fl mice with rapamycin reduced markers of senescence in immune cells and improved immune function11,12. These data demonstrate that an aged, senescent immune system has a causal role in driving systemic ageing and therefore represents a key therapeutic target to extend healthy ageing.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Reasonable requests for all data presented in this Article will be honoured by the corresponding authors.
References
Kennedy, B. K. et al. Geroscience: linking aging to chronic disease. Cell 159, 709–713 (2014).
St Sauver, J. L. et al. Risk of developing multimorbidity across all ages in an historical cohort study: differences by sex and ethnicity. BMJ Open 5, e006413 (2015).
Robinson, A. R. et al. Spontaneous DNA damage to the nuclear genome promotes senescence, redox imbalance and aging. Redox Biol. 17, 259–273 (2018).
Wang, J., Clauson, C. L., Robbins, P. D., Niedernhofer, L. J. & Wang, Y. The oxidative DNA lesions 8,5′-cyclopurines accumulate with aging in a tissue-specific manner. Aging Cell 11, 714–716 (2012).
Baker, D. J. et al. Naturally occurring p16Ink4a-positive cells shorten healthy lifespan. Nature 530, 184–189 (2016).
Zhu, Y. et al. The Achilles’ heel of senescent cells: from transcriptome to senolytic drugs. Aging Cell 14, 644–658 (2015).
Xu, M. et al. Senolytics improve physical function and increase lifespan in old age. Nat. Med. 24, 1246–1256 (2018).
Pettan-Brewer, C. & Treuting, P. M. Practical pathology of aging mice. Pathobiol. Aging Age Relat. Dis. 1, (2011).
Bumgardner, S. A. et al. Genetic influence on splenic natural killer cell frequencies and maturation among aged mice. Exp. Gerontol. 104, 9–16 (2018).
Liu, Y. et al. Expression of p16INK4a in peripheral blood T-cells is a biomarker of human aging. Aging Cell 8, 439–448 (2009).
Mannick, J. B. et al. mTOR inhibition improves immune function in the elderly. Sci. Transl. Med. 6, 268ra179 (2014).
Mannick, J. B. et al. TORC1 inhibition enhances immune function and reduces infections in the elderly. Sci. Transl. Med. 10, eaaq1564 (2018).
Olshansky, S. J. Articulating the case for the longevity dividend. Cold Spring Harb. Perspect. Med. 6, a025940 (2016).
van Deursen, J. M. The role of senescent cells in ageing. Nature 509, 439–446 (2014).
Kirkland, J. L. & Tchkonia, T. Cellular senescence: a translational perspective. EBioMedicine 21, 21–28 (2017).
Goronzy, J. J. & Weyand, C. M. Understanding immunosenescence to improve responses to vaccines. Nat. Immunol. 14, 428–436 (2013).
Krishnamurthy, J. et al. Ink4a/Arf expression is a biomarker of aging. J. Clin. Invest. 114, 1299–1307 (2004).
Vicente, R., Mausset-Bonnefont, A. L., Jorgensen, C., Louis-Plence, P. & Brondello, J. M. Cellular senescence impact on immune cell fate and function. Aging Cell 15, 400–406 (2016).
Pinti, M. et al. Aging of the immune system: Focus on inflammation and vaccination. Eur. J. Immunol. 46, 2286–2301 (2016).
Cho, J. S., Kook, S. H., Robinson, A. R., Niedernhofer, L. J. & Lee, B. C. Cell autonomous and nonautonomous mechanisms drive hematopoietic stem/progenitor cell loss in the absence of DNA repair. Stem Cells 31, 511–525 (2013).
Gregg, S. Q. et al. A mouse model of accelerated liver aging caused by a defect in DNA repair. Hepatology 55, 609–621 (2012).
Lavasani, M. et al. Muscle-derived stem/progenitor cell dysfunction limits healthspan and lifespan in a murine progeria model. Nat. Commun. 3, 608 (2012).
Nasto, L. A. et al. Genotoxic stress accelerates age-associated degenerative changes in intervertebral discs. Mech. Ageing Dev. 134, 35–42 (2013).
Niedernhofer, L. J. et al. A new progeroid syndrome reveals that genotoxic stress suppresses the somatotroph axis. Nature 444, 1038–1043 (2006).
Siegemund, S., Shepherd, J., Xiao, C. & Sauer, K. hCD2-iCre and Vav-iCre mediated gene recombination patterns in murine hematopoietic cells. PLoS ONE 10, e0124661 (2015).
Su, Y., Orelli, B., Madireddy, A., Niedernhofer, L. J. & Schärer, O. D. Multiple DNA binding domains mediate the function of the ERCC1-XPF protein in nucleotide excision repair. J. Biol. Chem. 287, 21846–21855 (2012).
Sharpe, A. H. & Pauken, K. E. The diverse functions of the PD1 inhibitory pathway. Nat. Rev. Immunol. 18, 153–167 (2017).
Sharpless, N. E. & Sherr, C. J. Forging a signature of in vivo senescence. Nat. Rev. Cancer 15, 397–408 (2015).
Coppé, J. P., Desprez, P. Y., Krtolica, A. & Campisi, J. The senescence-associated secretory phenotype: the dark side of tumor suppression. Annu. Rev. Pathol. 5, 99–118 (2010).
Zhang, H., Davies, K. J. A. & Forman, H. J. Oxidative stress response and Nrf2 signaling in aging. Free Radic. Biol. Med. 88 (Pt B), 314–336 (2015).
Shimatani, K., Nakashima, Y., Hattori, M., Hamazaki, Y. & Minato, N. PD-1+ memory phenotype CD4+ T cells expressing C/EBPα underlie T cell immunodepression in senescence and leukemia. Proc. Natl Acad. Sci. USA 106, 15807–15812 (2009).
Zhao, J. et al. Quantitative analysis of cellular senescence in culture and in vivo. Curr. Protoc. Cytom. 79, 9.51.1–9.51.25 (2017).
Munshi, R. et al. MCP-1 gene activation marks acute kidney injury. J. Am. Soc. Nephrol. 22, 165–175 (2011).
Argyropoulos, C. P. et al. Rediscovering β-2 microglobulin as a biomarker across the spectrum of kidney diseases. Front. Med. (Lausanne) 4, 73 (2017).
Burd, C. E. et al. Monitoring tumorigenesis and senescence in vivo with a p16INK4a-luciferase model. Cell 152, 340–351 (2013).
Ahmad, A. et al. ERCC1-XPF endonuclease facilitates DNA double-strand break repair. Mol. Cell. Biol. 28, 5082–5092 (2008).
Yousefzadeh, M. J. et al. Mouse models of accelerated cellular senescence. Methods Mol. Biol. 1896, 203–230 (2019).
Tabibian, J. H., O’Hara, S. P., Splinter, P. L., Trussoni, C. E. & LaRusso, N. F. Cholangiocyte senescence by way of N-ras activation is a characteristic of primary sclerosing cholangitis. Hepatology 59, 2263–2275 (2014).
Yousefzadeh, M. J. et al. Circulating levels of monocyte chemoattractant protein-1 as a potential measure of biological age in mice and frailty in humans. Aging Cell 17, e12706 (2018).
Ladiges, W. Pathology assessment is necessary to validate translational endpoints in preclinical aging studies. Pathobiol. Aging Age Relat. Dis. 6, 31478 (2016).
Farndale, R. W., Buttle, D. J. & Barrett, A. J. Improved quantitation and discrimination of sulphated glycosaminoglycans by use of dimethylmethylene blue. Biochim. Biophys. Acta 883, 173–177 (1986).
Acknowledgements
This work was supported by the National Institutes of Health (NIH) grants P01 AG043376 (P.D.R., L.J.N., E.E.K., J.H.), RO1 AG063543 (L.J.N.), R56 AG059676 (L.J.N.), U19 AG056278 (P.D.R., L.J.N., W.C.L.), P01 AG062413 (P.D.R., L.J.N.), R56 AG058543 (W.C.L.), R01 AG044376 (N.V.V.) and the Glenn Foundation (L.J.N., C.E.B.). M.J.Y. is supported by The Irene Diamond Fund/American Federation on Aging Research Postdoctoral Transition Award. Mass cytometry and panel design were performed by S. Farwana and K. D. Pavelko at the Mayo Clinic Immune Monitoring Core. We thank J. Zhao, C. Bukata, K. Melos and M. Calubag for their assistance in measuring senescence. All mouse illustrations were made with BioRender.
Author information
Authors and Affiliations
Contributions
M.J.Y., R.R.F., P.D.R. and L.J.N. conceived and designed the study. M.J.Y., L.A., S.J.M., T.S. and R.D.O. performed most of the mouse manipulations and in vivo physiological analysis. M.J.Y., R.W.B., J.I.K. and M.P.B. performed ex vivo senescence analysis. C.E.B. provided p16-luciferase reporter mice. E.A.W. assisted with the IVIS analysis of p16-luciferase with L.A. and S.J.M. C.E.T. performed and N.F.L. contributed to p16 in situ hybridization studies. R.R.F. performed immune population analysis. M.J.Y. and K.L. prepared samples for and Y.Z. performed and analysed data from CyTOF with assistance from Z.C.S., A.L.B. and I.M.S. M.J.Y. and T.S. performed immune function tests. M.J.Y. and L.A. performed transplantation experiments. M.J.Y. performed and S.E.L., E.E.K., Y.C. and Y.W. contributed to the analysis of oxidative stress and DNA damage analysis. A.L. and J.H. performed analysis of cardiotoxin-injected muscle tissues. D.W. and Q.D. performed and N.V.V. contributed to intervertebral disc and spine analysis. J.K., S.P.S.P. and W.C.L. performed histopathological analysis and C.A.M. performed the grip strength analysis. M.J.Y. and R.R.F. wrote the manuscript with input from all co-authors. M.J.Y., R.R.F., P.D.R. and L.J.N. oversaw all experimental design, data analysis and manuscript preparation.
Corresponding authors
Ethics declarations
Competing interests
L.J.N. and P.D.R. are co-founders of NRTK Biosciences, a start-up biotechnology company developing senolytic drugs.
Additional information
Peer review information Nature thanks Joan Mannick, Björn Schumacher and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Molecular changes in Vav-iCre+/−;Ercc1−/fl mice.
a, Expression of Ercc1 was measured in tissues from 8–10-month-old Vav-iCre+/−;Ercc1−/fl and Ercc1−/fl control mice (n = 5–7 Vav-iCre+/−;Ercc1−/fl; n = 3–5 Ercc1−/fl, depending on the tissue (see Supplementary Table 3 for sample size details). b, Detection of ERCC1 in splenic lysates from a 9-month-old Vav-iCre+/−;Ercc1−/fl mice and littermate control by immunoblot. c, Superoxide anion levels were measured by electron paramagnetic resonance (EPR) in splenic tissue from 6–8-month-old Vav-iCre+/−;Ercc1−/fl and littermate control mice (n = 5 mice per group). d, e, Expression of the transcription factor NRF (Nfe2l2) and its downstream targets (Cat, Nqo1, Hmox1) measured by qRT–PCR in spleen (d) and bone marrow (e) of Vav-iCre+/−;Ercc1−/fl and littermate control mice at several ages (n = 3 at 3- and 5-months-old; n = 5 8–10-months-old) and in two-year-old wild-type mice (n = 5). f, Catalase activity measured in splenic tissue from 8–10-month-old Vav-iCre+/−;Ercc1−/fl (n = 6) and Vav-iCre+/− (n = 3) mice (Methods). g, Catalase activity in 4-month-old (n = 3) and 24-month-old (n = 6) wild-type mice. h, i, The ratio of reduced to oxidized glutathione (GSH/GSSG) (Methods) (h) and levels of HNE protein adducts (i) measured by ELISA in splenic lysates of Vav-iCre+/−;Ercc1−/fl and littermate control mice at 8–11 months of age (n = 6 mice per group). j, Levels of four cyclopurine adducts in splenic tissue from 8–10-month-old Vav-iCre+/−;Ercc1−/fl mice and littermate controls (n = 4–5 Vav-iCre+/−;Ercc1−/fl; n = 5 Vav-iCre+/−; see Supplementary Table 3 for sample size details) measured by liquid chromatograph–tandem mass spectrometry (LC–MS/MS/MS) (Methods). k, Total splenocyte cell counts from 8–10-month-old Vav-iCre+/−;Ercc1−/fl (n = 14) mice and Vav-iCre+/− (n = 12) mice. l, The absolute number of T (CD4+, CD8+) and B (B220+CD19+) cells in spleens from the same mice (n = 10/4 Vav-iCre+/−;Ercc1−/fl; n = 8/3 Vav-iCre+/− for CD4+ or CD8+/ B220+CD19+ measures, respectively) (Methods). m, Total splenocyte cell counts from young (7-month-old; n = 10) and old (24-month-old; n = 17) wild-type mice. n, The absolute number of CD4+, CD8+ and B220+CD19+ cell in spleens from the same mice (n = 8/3 young WT; n = 17/7 old wild-type mice for CD4+ or CD8+/B220+CD19+ measures, respectively). o, Analysis of CD8+ splenocytes from 8–10-month-old mice for memory (CD44+CD127+), exhaustion (PD-1+) and apoptosis (VAD-FMK+) markers (n = 6 mice per group). p, Thymic weight normalized to total body weight (n = 3 at 3 months old; n = 4–5 at 8–10 months old per group). q, Histology images (20×) of spleen and lymph nodes from 8–10-month-old Vav-iCre mice. Scale bar, 100 μm. GC, germinal centres. Data are mean ± s.d. *P < 0.05, ∞P < 0.01, ∇P < 0.001, #P < 0.0001, unpaired two-tailed Student’s t-test (a, c–e for the 3- and 5-month-old mice, g–p), one-way ANOVA (d, e for the 8–10-month-old mice) or two-way (f) with Tukey’s test.
Extended Data Fig. 2 Vav-iCre+/−;Ercc1−/fl mice maintain normal weight and body composition.
a, Body weight (BW) of three different age groups of mice by genotype. b–d, Percentage fat (b), lean mass (c) and fluid (d) measured by NMR at three ages of mice (n = 8–25 mice per group) (see Supplementary Table 3 for sample size details). Data are mean ± s.d. P values (not significant) were determined by one-way ANOVA with Tukey’s test.
Extended Data Fig. 3 Measurement of immune function and lymphoid organ senescence in Vav-iCre+/−;Ercc1−/fl mice.
a, Footpad swelling measurements at several time points after antigenic (KLH) challenge, separated by mouse genotype (n = 7 Vav-iCre+/−;Ercc1−/fl; n = 6 Vav-iCre+−; n = 5 old WT mice). b, Footpad swelling by genotype (n = 7 Vav-iCre+/−;Ercc1−/fl; n = 6 Vav-iCre+/−; n = 5 WT mice). c, Quantification of anti-KLH antibodies by ELISA one month after antigenic challenge (n = 3/7 Vav-iCre+/−;Ercc1−/fl; n = 3/6 Vav-iCre+/−; n = 3/3 Ercc1−/fl for naive (uninjected) and KLH-challenged (injected) mice, respectively). d, DTH assay in 2-month-old mice Vav-iCre+/−;Ercc1−/fl (n = 5) and Vav-iCre+/− (n = 8) controls. Data are mean ± s.d. ∞P < 0.01, ∇P < 0.001, #P < 0.0001, two-way ANOVA with Tukey’s test. e, Quantification of Ercc1 expression by qRT–PCR in flow-sorted immune cell populations isolated from spleen (T and natural killer cells) and bone marrow (B cells and macrophages) of 5-month-old Vav-iCre+/−;Ercc1−/fl mice and littermate controls (n = 4 Vav-iCre+/−;Ercc1−/fl; n = 3 Vav-iCre+/−). Data are mean ± s.d. ∇P < 0.001, #P < 0.0001, two-tailed unpaired Student’s t-test. f, Senescence marker expression in CD3+ peripheral T cells from 8–11-month-old Vav-iCre and old wild-type mice (n = 3–9 Vav-iCre+/−; n = 6–9; Vav-iCre+/−;Ercc1−/fl; n = 4–7 two-year-old wild-type, depending on the gene) (see Supplementary Table 3 for sample size details). g, Measurement of senescence marker expression by qRT–PCR in splenic tissue and bone marrow from 2–3-month-old Vav-iCre mice (n = 3). h, Expression of senescence markers in splenic tissue from 8–10-month-old Vav-iCre+/−;Ercc1−/fl and old wild-type mice relative to controls by gender (n = 3–4/3–4 Vav-iCre+/−; n = 3–5/3–4 Vav-iCre+/−;Ercc1−/fl; n = 3–6/3–5 4-month-old WT; n = 5/5 two-year-old WT males/females, respectively) (see Supplementary Table 3 for sample size details). i, Expression of senescence markers in bone marrow of Vav-iCre+/−;Ercc1−/fl mice (n = 3–4/3–4 Vav-iCre+/−; n = 3–4/4–5 Vav-iCre+/−;Ercc1−/fl males/females, respectively). Data are mean ± s.d. *P < 0.05, ∞P < 0.01, ∇P < 0.001, #P < 0.0001, unpaired two-tailed Student’s t-test (e, g), one-way ANOVA (f) or two-way ANOVA (a–d, h, i) with Tukey’s test.
Extended Data Fig. 4 Identification of immune cell types and senescent cells by CyTOF.
a, viSNE analysis of total CD45+ cells to identify immune cell types. Representative viSNE plots are from a Vav-iCre+/− control mouse at 10–12 months of age. See Supplemental Fig. 5 for the gating strategy. b, The proportion of the indicated immune cell subsets that express p16, p21 or CENP-B from 10–12-month-old Vav-iCre+/−;Ercc1−/fl (n = 6), Vav-iCre+/− (n = 6), and >2-year-old wild-type (n = 7) mice was determined by CyTOF. Each dot is an independent mouse. Data are mean ± s.d. *P < 0.05, **P < 0.01, ***P < 0.001, Kruskal–Wallis test with Dunn’s correction for multiple comparisons.
Extended Data Fig. 5 Co-localization of p16 mRNA with parenchymal markers in non-lymphoid tissues from Vav-iCre+/−;Ercc1−/fl mice.
a, Measurement of senescence marker expression in the organs of 5-month-old Vav-iCre mice (n = 3 mice per group). b–e, Representative images of p16 in situ hybridization with immunostain or chemical stain for parenchymal markers of liver (b, c), kidney (d) and lung (e) sections from 8–11-month-old Vav-iCre+/−;Ercc1−/fl mice, littermate control Vav-iCre+/− mice and 2-year-old wild-type mice stained for albumin (liver), phalloidin (liver and lung) or kidney-specific (KSP)-cadherin in the red channel, DAPI (blue), and p16 LNA probe (green). The full set of images from Fig. 3c is shown in b. Original magnification, ×40. Scale bar, 10 μm. f, Representative images of SA-β-gal staining on tissues from 8–10-month-old Vav-iCre+/−;Ercc1−/fl and littermate controls. Original magnification, ×20. Scale bar, 50 μm. g, Senescence marker expression in the livers of 8–11-month-old Vav-iCre+/−;Ercc1−/fl (n = 5 male and 4–5 female) (see Supplementary Table 3 for sample size details by gender and gene) and littermate control mice (n = 3 male and 3–4 female) as well as 4-month-old (n = 3 male and 3–4 female) and 2-year-old (n = 6 male and 4–5 female) wild-type mice. h, Levels of circulating SASP factor proteins measured by multiplex ELISA in serum from Vav-iCre+/−;Ercc1−/fl (n = 3 male and 3–4 female) and Vav-iCre+/− (n = 3 male and 3 female) mice. Data are mean ± s.d. *P < 0.05, ∞P < 0.01, ∇P < 0.001, #P < 0.0001, unpaired two-tailed Student’s t-test (a) or two-way ANOVA with Tukey’s test (g, h).
Extended Data Fig. 6 Cell non-autonomous effects of an aged immune system in non-lymphoid tissues of Vav-iCre+/−;Ercc1−/fl mice.
a, Cyclopurine adducts were measured in the liver and kidneys of 8–11-month-old Vav-iCre+/−;Ercc1−/fl (n = 5) and littermate control Vav-iCre+/− (n = 5 for liver and n = 6 for kidney) by LC–MS/MS/MS (Methods). b, Markers of oxidative stress including HNE protein adducts and the ratio of reduced to oxidized glutathione (GSH/GSSG) measured in the kidneys from 8–11-month-old Vav-iCre+/−;Ercc1−/fl and littermate control Vav-iCre+/− mice (n = 6 mice per group). HNE measure by ELISA. GSH/GSSG measured by chromogenic assay (Methods). c, 8-oxo-guanine DNA adducts measured by ELISA in the spleen, liver and kidney of mice at various ages (n = 5–6/5–6/5 Vav-iCre+/−;Ercc1−/fl; n = 5–6/5–6/5 Vav-iCre+/−; n = 5/5/10 old WT mice for spleen/liver/kidney, respectively) (see Supplementary Table 3 for sample size details by genotype and tissue). d, Urinary levels of pro-geronic factor β2-microglobulin and MCP-1 measured by ELISA in 8–11-month-old Vav-iCre+/−;Ercc1−/fl and littermate controls (n = 9 mice per group). e, Renal Mcp1 (also known as Ccl2) expression in 8–11-month-old Vav-iCre+/−;Ercc1−/fl mice (n = 7 per group) measured by qRT–PCR. f, Representative Coomassie-stained gel of urine samples from 8–11-month-old Vav-iCre+/−;Ercc1−/fl and littermate control mice demonstrating increased proteinuria. Recombinant albumin (Alb) was loaded on the gel as a control (lanes 6, 12) and its approximate molecular mass denoted by a box (marker ladder lanes 1, 5, 7, 11, 13–14). Each lane represents a unique mouse. g, Representative images from tissue sections stained for aggrecan (red) and DAPI (blue) in the nucleus pulposus (NP) of intervertebral discs from 8–11-month-old Vav-iCre+/−;Ercc1−/fl and littermate control mice. h, Quantification of aggrecan staining (n = 4 Vav-iCre+/−; n = 7 Vav-iCre+/−;Ercc1−/fl). i, Measurement of senescence marker expression in the intervertebral discs of 8–11-month-old Vav-iCre+/−;Ercc1−/fl and littermate control mice (n = 4 mice per group) by qRT–PCR. j, Representative images of sections of intervertebral discs from 9-month-old mice stained with haematoxylin and eosin (H&E) and safranin O to detect proteoglycans. Arrows point to the anulus fibrosus. k, Representative images of gastrocnemius muscle sections from 8–11-month-old Vav-iCre+/−;Ercc1−/fl and littermate control mice after cardiotoxin injury (Methods) stained with haematoxylin and eosin or immunostained for M1 (CD68, green) and M2 (CD163, red) macrophages. l, Quantification of the ratio of M2/M1 macrophages (n = 4 Vav-iCre+/−; n = 8 Vav-iCre+/−;Ercc1−/fl mice). m, Body weight and grip strength of Vav-iCre+/−;Ercc1−/fl and littermate controls at the indicated ages (n = 8 Vav-iCre+/−;Ercc1−/fl; n = 4 Vav-iCre+/− at both ages). Data are mean ± s.d. *P < 0.05, ∞P < 0.01, ∇P < 0.001, #P < 0.0001, two-tailed unpaired Student’s t-test (a, b, d, e, h, i, l, m) or two-way ANOVA with Tukey’s test (c).
Extended Data Fig. 7 Age-associated increase in serum SASP factors of Vav-iCre+/−;Ercc1−/fl mice and wild-type mice.
a, This is an extension of the data shown in Fig. 3k, including two younger ages of mice. Circulating SASP factors were measured by ELISA (n = 3 for 2–3-month-old, n = 4 for 5-month-old, n = 5–7 for 8–11-month-old, n = 7 for 24-month-old) (see Supplementary Table 3 for sample size details). Data are mean ± s.d. *P < 0.05, ∞P < 0.01, ∇P < 0.001, #P < 0.0001, two-way ANOVA with Tukey’s test. b, Haematoxylin and eosin sections of brain, kidney and liver from 17-month-old Vav-iCre+/−;Ercc1−/fl and littermate controls (n = 5 mice per group) were scored for age-associated histopathological lesions using the Geropathology Grading Platform to generate a composite lesion score for each organ (CLS). Data are mean ± s.d. P values (not significant) were determined by two-tailed unpaired Student’s t-test.
Extended Data Fig. 8 Time course of bioluminescence signal in p16-luciferase mice transplanted with splenocytes and tissue distribution of transplanted cells.
a, Splenocytes from 8–10-month-old Vav-iCre+/−;Ercc1−/fl, Vav-iCre+/− controls, or 2-year-old wild-type mice were injected retro-orbitally into 3–4-month-old p16Ink4+/Luc senescence reporter mice (n = 2 donor mice per genotype) as described in Fig. 4. Tissues were collected from recipient mice 2 weeks after the final imaging and the expression of p21 was measured by qRT–PCR. Data are mean ± s.d. *P < 0.05, ∞P < 0.01, one-way ANOVA with Tukey’s test. b, Splenocytes (5 × 106 cells) from 9–10-month-old Vav-iCre+/−;Ercc1−/fl and Vav-iCre+/− mice were injected retro-orbitally into 3–4-month-old p16Luc/Luc senescence reporter mice (n = 2 donor mice per genotype; n = 4 p16Luc/Luc recipient mice for Vav-iCre+/−;Ercc1−/fl splenocytes; n = 3 receiving Vav-iCre+/− splenocytes). Weekly measurements of luminescence in recipient reporter mice. Data are mean ± s.d. *P < 0.05, ∞P < 0.01, two-tailed unpaired Student’s t-test. c, Splenocytes from 7- or 26-month-old male mice were injected retro-orbitally into female mice to track distribution of the transplanted cells (n = 2 donor mice and n = 3 recipient mice per age group; n = 2 uninjected controls). Tissues were collected 24 h after injection. Expression of the Sry gene on the Y chromosome measured by qRT–PCR in RNA isolated from tissues of recipient mice was used to track homing of immune cells to various recipient mouse organs. There was little difference in immune cell homing if the donor mice were young or old.
Extended Data Fig. 9 Suppression of senescence in progeroid mice by transplantation of young immune cells.
a, Schematic of adoptive transfer: 3-month-old progeroid Ercc1−/∆ mice (n = 4 mice/treatment group) were injected retro-orbitally with 5x106 splenocytes from 2-month-old WT mice or vehicle only (PBS) (n = 6 donors). One month later, the recipient mice and uninjected age-matched wild-type mice were euthanized, and tissues collected. b, Expression of senescence markers in organs of recipient mice (n = 4 Ercc1−/∆ + splenocytes; n = 3–6 Ercc1−/∆ + PBS) (see Supplementary Table 3 for sample size details by organ/endpoint). Gene expression was normalized to that of uninjected, age-matched wild-type controls (n = 4–7) represented as horizontal dashed line. c, SASP factor proteins MCP-1 and TNF were measured in the serum of recipient mice by multiplex ELISA (n = 4 Ercc1−/∆ + splenocytes; n = 6 Ercc1−/∆ + PBS) and compared to untreated, age-matched wild-type mice (n = 4–6). d, Footpad swelling of mice describe above and in Fig. 5f at several time points after antigenic challenge (n = 3/7 Vav-iCre+/−;Ercc1−/fl or n = 3/6 Vav-iCre+/− mice +/− rapamycin, respectively). e, Expression of p21 in PBMCs, measured by qRT–PCR. f, Serum MCP-1 and TNF measured by multiplex ELISA (n = 3/5 +/− rapamycin, respectively). Data are mean ± s.d. *P < 0.05, ∞P < 0.01, ∇P < 0.001, #P < 0.0001 one-way ANOVA (b, c) or two-way ANOVA (d–f) with Tukey’s test. Mouse images in schematic were used with permission from BioRender.
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-6, Supplementary Tables 1-3 and the Supplementary Discussion.
Rights and permissions
About this article
Cite this article
Yousefzadeh, M.J., Flores, R.R., Zhu, Y. et al. An aged immune system drives senescence and ageing of solid organs. Nature 594, 100–105 (2021). https://doi.org/10.1038/s41586-021-03547-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-021-03547-7
This article is cited by
-
Early-life exercise induces immunometabolic epigenetic modification enhancing anti-inflammatory immunity in middle-aged male mice
Nature Communications (2024)
-
Transcription of endogenous retroviruses in senescent cells contributes to the accumulation of double-stranded RNAs that trigger an anti-viral response that reinforces senescence
Cell Death & Disease (2024)
-
Derailed protein turnover in the aging mammalian brain
Molecular Systems Biology (2024)
-
Clonal haematopoiesis, ageing and kidney disease
Nature Reviews Nephrology (2024)
-
Inhibition of S6K lowers age-related inflammation and increases lifespan through the endolysosomal system
Nature Aging (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.