Abstract
Coordinated activity across networks of neurons is a hallmark of both resting and active behavioural states in many species1,2,3,4,5. These global patterns alter energy metabolism over seconds to hours, which underpins the widespread use of oxygen consumption and glucose uptake as proxies of neural activity6,7. However, whether changes in neural activity are causally related to metabolic flux in intact circuits on the timescales associated with behaviour is unclear. Here we combine two-photon microscopy of the fly brain with sensors that enable the simultaneous measurement of neural activity and metabolic flux, across both resting and active behavioural states. We demonstrate that neural activity drives changes in metabolic flux, creating a tight coupling between these signals that can be measured across brain networks. Using local optogenetic perturbation, we demonstrate that even transient increases in neural activity result in rapid and persistent increases in cytosolic ATP, which suggests that neuronal metabolism predictively allocates resources to anticipate the energy demands of future activity. Finally, our studies reveal that the initiation of even minimal behavioural movements causes large-scale changes in the pattern of neural activity and energy metabolism, which reveals a widespread engagement of the brain. As the relationship between neural activity and energy metabolism is probably evolutionarily ancient and highly conserved, our studies provide a critical foundation for using metabolic proxies to capture changes in neural activity.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Raw imaging data are available upon request to the corresponding authors.
Code availability
Analysis scripts are available upon request from the corresponding authors.
References
Liska, A., Galbusera, A., Schwarz, A. J. & Gozzi, A. Functional connectivity hubs of the mouse brain. Neuroimage 115, 281–291 (2015).
Ahrens, M. B., Orger, M. B., Robson, D. N., Li, J. M. & Keller, P. J. Whole-brain functional imaging at cellular resolution using light-sheet microscopy. Nat. Methods 10, 413–420 (2013).
Mann, K., Gallen, C. L. & Clandinin, T. R. Whole-brain calcium imaging reveals an intrinsic functional network in Drosophila. Curr. Biol. 27, 2389–2396.e4 (2017).
Prevedel, R. et al. Simultaneous whole-animal 3D imaging of neuronal activity using light-field microscopy. Nat. Methods 11, 727–730 (2014).
Power, J. D., Schlaggar, B. L. & Petersen, S. E. Studying brain organization via spontaneous fMRI signal. Neuron 84, 681–696 (2014).
Logothetis, N. K. What we can do and what we cannot do with fMRI. Nature 453, 869–878 (2008).
Magistretti, P. J. & Allaman, I. A cellular perspective on brain energy metabolism and functional imaging. Neuron 86, 883–901 (2015).
Horwitz, B., Soncrant, T. T. & Haxby, J. V. in Advances in Metabolic Mapping Techniques for Brain Imaging of Behavioral and Learning Functions (eds Gonzalez-Lima, F. et al.) 189–217 (1992).
Sokoloff, L., Kennedy, C. & Smith, C. B. in Carbohydrates and Energy Metabolism (eds Boulton, A. A. et al.) 155–193 (1989).
Passow, S. et al. Default-mode network functional connectivity is closely related to metabolic activity. Hum. Brain Mapp. 36, 2027–2038 (2015).
Schölvinck, M. L., Maier, A., Ye, F. Q., Duyn, J. H. & Leopold, D. A. Neural basis of global resting-state fMRI activity. Proc. Natl Acad. Sci. USA 107, 10238–10243 (2010).
Migault, G. et al. Whole-brain calcium imaging during physiological vestibular stimulation in larval zebrafish. Curr. Biol. 28, 3723–3735.e6 (2018).
Harris, D. T., Kallman, B. R., Mullaney, B. C. & Scott, K. Representations of taste modality in the Drosophila brain. Neuron 86, 1449–1460 (2015).
Lemon, W. C. et al. Whole-central nervous system functional imaging in larval Drosophila. Nat. Commun. 6, 7924 (2015).
Aimon, S. et al. Fast near-whole-brain imaging in adult Drosophila during responses to stimuli and behavior. PLoS Biol. 17, e2006732 (2019).
Palva, J. M. & Palva, S. Infra-slow fluctuations in electrophysiological recordings, blood-oxygenation-level-dependent signals, and psychophysical time series. Neuroimage 62, 2201–2211 (2012).
Mitra, A. et al. Spontaneous infra-slow brain activity has unique spatiotemporal dynamics and laminar structure. Neuron 98, 297–305.e6 (2018).
Biswal, B., Yetkin, F. Z., Haughton, V. M. & Hyde, J. S. Functional connectivity in the motor cortex of resting human brain using echo-planar MRI. Magn. Reson. Med. 34, 537–541 (1995).
San Martín, A. et al. Imaging mitochondrial flux in single cells with a FRET sensor for pyruvate. PLoS ONE 9, e85780 (2014).
Plaçais, P.-Y. et al. Upregulated energy metabolism in the Drosophila mushroom body is the trigger for long-term memory. Nat. Commun. 8, 15510 (2017).
Gonzalez-Gutierrez, A., Ibacache, A., Esparza, A., Felipe Barros, L. & Sierralta, J. Monocarboxylate transport in Drosophila larval brain during low and high neuronal activity. Preprint at https://doi.org/10.1101/610196 (2019).
Lobas, M. A. et al. A genetically encoded single-wavelength sensor for imaging cytosolic and cell surface ATP. Nat. Commun. 10, 711 (2019).
Chen, T.-W. et al. Ultrasensitive fluorescent proteins for imaging neuronal activity. Nature 499, 295–300 (2013).
Jenett, A. et al. A GAL4-driver line resource for Drosophila neurobiology. Cell Rep. 2, 991–1001 (2012).
Ito, K. et al. A systematic nomenclature for the insect brain. Neuron 81, 755–765 (2014).
Dana, H. et al. Sensitive red protein calcium indicators for imaging neural activity. eLife 5, e12727 (2016).
Saitoe, M., Schwarz, T. L., Umbach, J. A., Gundersen, C. B. & Kidokoro, Y. Absence of junctional glutamate receptor clusters in Drosophila mutants lacking spontaneous transmitter release. Science 293, 514–517 (2001).
Klapoetke, N. C. et al. Independent optical excitation of distinct neural populations. Nat. Methods 11, 338–346 (2014).
Stocker, R. F., Heimbeck, G., Gendre, N. & de Belle, J. S. Neuroblast ablation in Drosophila P[GAL4] lines reveals origins of olfactory interneurons. J. Neurobiol. 32, 443–456 (1997).
Wagenmakers, E.-J., Farrell, S. & Ratcliff, R. Estimation and interpretation of 1/fα noise in human cognition. Psychon. Bull. Rev. 11, 579–615 (2004).
Weissman, M. B. 1/f noise and other slow, nonexponential kinetics in condensed matter. Rev. Mod. Phys. 60, 537–571 (1988).
Namiki, S., Dickinson, M. H., Wong, A. M., Korff, W. & Card, G. M. The functional organization of descending sensory-motor pathways in Drosophila. eLife 7, e34272 (2018).
Bidaye, S. S., Machacek, C., Wu, Y. & Dickson, B. J. Neuronal control of Drosophila walking direction. Science 344, 97–101 (2014).
Hugdahl, K., Raichle, M. E., Mitra, A. & Specht, K. On the existence of a generalized non-specific task-dependent network. Front. Hum. Neurosci. 9, 430 (2015).
Cong, L. et al. Rapid whole brain imaging of neural activity in freely behaving larval zebrafish (Danio rerio). eLife 6, e28158 (2017).
Nguyen, J. P. et al. Whole-brain calcium imaging with cellular resolution in freely behaving Caenorhabditis elegans. Proc. Natl Acad. Sci. USA 113, E1074–E1081 (2016).
Steinmetz, N. A., Zatka-Haas, P., Carandini, M. & Harris, K. D. Distributed coding of choice, action and engagement across the mouse brain. Nature 576, 266–273 (2019).
Selemon, L. D. & Goldman-Rakic, P. S. Common cortical and subcortical targets of the dorsolateral prefrontal and posterior parietal cortices in the rhesus monkey: evidence for a distributed neural network subserving spatially guided behavior. J. Neurosci. 8, 4049–4068 (1988).
Oliphant, T. E. Python for scientific computing. Comput. Sci. Eng. 9, 10–20 (2007).
Millman, K. J., Jarrod Millman, K. & Aivazis, M. Python for scientists and engineers. Comput. Sci. Eng. 13, 9–12 (2011).
Friard, O. & Gamba, M. BORIS: a free, versatile open‐source event‐logging software for video/audio coding and live observations. Methods Ecol. Evol. 7, 1325–1330 (2016).
Acknowledgements
We thank C. Gallen for providing discussion and support. This work was supported by the Simons foundation (T.R.C. and S.G.), an NSF Career Award 1845166 (S.G.) and the Stanford Wu Tsai neuroscience institute (K.M.)
Author information
Authors and Affiliations
Contributions
K.M., S.D., S.G. and T.R.C. conceived the project. K.M. collected the data. K.M. and S.D. analysed the data and generated figures. K.M., S.D., S.G. and T.R.C. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks Marcus Raichle and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Normalized iATPSnFR responses in whole brains to ATP.
Normalized ∆F/F values for different concentrations of ATP measured in whole brains expressing iATPSnFR pan-neuronally. n = 10 flies, mean ± s.e.m.
Extended Data Fig. 2 Example traces and correlations of Pyronic, jRGECO1a and iATPSnFR.
a, Pyronic traces over an imaging session in different regions. b, A pair of traces that exhibit high correlation over time. c, Scatter plot of these two regions demonstrating correlation. d, A pair of traces that exhibit lower correlation over time. e, Scatter plot of these two regions demonstrating correlation. f–j, As in a–e, but with jRGECO1a. k–o, As in a–e, but with iATPSnFR.
Extended Data Fig. 3 Correlation matrices of GCaMP6s, Pyronic and iATPSnFR.
a–c, Correlation matrices for GCaMP6s, Pyronic and iATPSnFR, reproduced and enlarged from Fig. 1, and labelling each individual region.
Extended Data Fig. 4 Correspondence of functional networks derived from simultaneous jRGECO1a, Pyronic and iATPSnFR measurements.
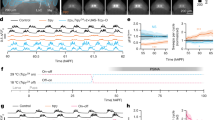
a, Left, traces displaying iATPSnFR (green) and corresponding jRGECO1a signal (blue). Right, Pyronic signals (orange) and corresponding jRGECO1a signals (blue) across six different brain regions b, Correlation matrix derived from jRGECO1a in the simultaneous imaging experiments from a and Fig. 2. c, Correlation matrix derived from Pyronic in the simultaneous imaging experiments from a and Fig. 2. d, Scatter plot of the pairwise correlations between jRGECO1a and Pyronic. e–g, As in b–d, but with jRGECO1a and iATPSnFR. n = 23 flies for Pyronic and n = 9 flies for iATPSnFR. h–m, Comparison of jRGECO1a and Pyronic signals within a single brain region (saddle (SAD)). h, Traces of Pyronic and jRGECO1a signals including all frequency components. i, Pairwise comparison of Pyronic and jRGECO1a signals including all frequency components and the correlation between these signals. j, k, As in h, i, but filtered to include only low-frequency (<0.1 Hz) components. l, m, As in h, i, but filtered to include only high-frequency (>0.1 Hz) components.
Extended Data Fig. 5 Neural activity drives metabolic flux in the brain.
a, jRGECO1a (blue), Pyronic (orange) and iATPSnFR (green) traces in three different brain regions before (left) and after (right) application of TTX. b, Region-by-region correlations between jRGECO1a and Pyronic signals (orange) and between jRGECO1a and iATPSnFR signals (green), across all flies, before TTX application (top row) and after TTX application (bottom row). Mean ± s.e.m. c, GCaMP6s response to 100-ms activation pulse in flies that lack CsChrimson. n = 45 ROIs, mean ± s.e.m. d, As in c, but with iATPSnFR. n = 45 ROIs, mean ± s.e.m.
Extended Data Fig. 6 Example model predictions of behaviour and CsChrimson controls.
a, Schematic of the data processing and analysis pipeline used: (i) traces of Pyronic, iATPSnFR, jRGECO1a and behaviour (movement of the legs); (ii) half of the dataset was used to train a logistic regression model relating neural activity and metabolic flux to behaviour; (iii) predicted behavioural outputs were generated using the withheld data and were compared to the actual behaviour during those time periods; and (iv) model prediction was evaluated by correlating predicted behaviour to observed behaviour. b, Left, four example flies showing the prediction based on the model for jRGECO1a (blue) with the corresponding behaviour trace (black). Correlation between signals shown above each trace. Right, weights for each ROI generated by the model shown on right (oriented as in Fig. 4c). c, As in b, but with Pyronic (orange). d, e, As in b, c, but with a different set of four flies, with jRGECO1a (blue), iATPSnFR (green) and behaviour trace (black). f, Correlation between model weights derived from iATPSnFR and jRGECO1a. g, Correlation between model weights derived from Pyronic and jRGECO1a.
Extended Data Fig. 7 Frequency spectra of jRGECO1a, Pyronic, iATPSnFR and behaviour.
Normalized spectra from data presented in Fig. 4.
Extended Data Fig. 8 Correlation of model weights for GCaMP6s and descending-neuron innervation.
a, Model weights for each brain region generated using GCaMP6s. b, The number of descending-neuron processes in each brain region (abbreviations defined as in ref. 32). c, Graphical representation of model weights, similar to Fig. 4c. d, Correlation between model weights and descending-neuron innervation by each region. e, Correlation between model weights derived from GCaMP6s and jRGECO1a.
Extended Data Fig. 9 Changes in correlations across regions during behaviour for both jRGECO1a and Pyronic.
a, Functional connectivity map of jRGECO1a during bouts of rest. b, Functional connectivity map of jRGECO1a during bouts of activity. c, Correlation of functional connectivity maps during resting and behaving bouts. Correlations increase across the vast majority of regions (P = 0.004, n = 12 flies, one-tailed t-test). d–f, As in a–c, but for Pyronic (P = 0.13, n = 7 flies, one-tailed t-test). g–i, As in a–c, but for iATPSnFR (P = 0.38, n = 13 flies, one-tailed t-test).
Supplementary information
Rights and permissions
About this article
Cite this article
Mann, K., Deny, S., Ganguli, S. et al. Coupling of activity, metabolism and behaviour across the Drosophila brain. Nature 593, 244–248 (2021). https://doi.org/10.1038/s41586-021-03497-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-021-03497-0
This article is cited by
-
Multimodal measures of spontaneous brain activity reveal both common and divergent patterns of cortical functional organization
Nature Communications (2024)
-
High-temporal resolution functional PET/MRI reveals coupling between human metabolic and hemodynamic brain response
European Journal of Nuclear Medicine and Molecular Imaging (2024)
-
Glia instruct axon regeneration via a ternary modulation of neuronal calcium channels in Drosophila
Nature Communications (2023)
-
The spatial and temporal structure of neural activity across the fly brain
Nature Communications (2023)
-
Brain mitochondrial diversity and network organization predict anxiety-like behavior in male mice
Nature Communications (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.