Abstract
Cyclic GMP–AMP synthase (cGAS) is an innate immune sensor for cytosolic microbial DNA1. After binding DNA, cGAS synthesizes the messenger 2′3′-cyclic GMP–AMP (cGAMP)2,3,4, which triggers cell-autonomous defence and the production of type I interferons and pro-inflammatory cytokines via the activation of STING5. In addition to responding to cytosolic microbial DNA, cGAS also recognizes mislocalized cytosolic self-DNA and has been implicated in autoimmunity and sterile inflammation6,7. Specificity towards pathogen- or damage-associated DNA was thought to be caused by cytosolic confinement. However, recent findings place cGAS robustly in the nucleus8,9,10, where tight tethering of chromatin is important to prevent autoreactivity to self-DNA8. Here we show how cGAS is sequestered and inhibited by chromatin. We provide a cryo-electron microscopy structure of the cGAS catalytic domain bound to a nucleosome, which shows that cGAS does not interact with the nucleosomal DNA, but instead interacts with histone 2A–histone 2B, and is tightly anchored to the ‘acidic patch’. The interaction buries the cGAS DNA-binding site B, and blocks the formation of active cGAS dimers. The acidic patch robustly outcompetes agonistic DNA for binding to cGAS, which suggests that nucleosome sequestration can efficiently inhibit cGAS, even when accessible DNA is nearby, such as in actively transcribed genomic regions. Our results show how nuclear cGAS is sequestered by chromatin and provides a mechanism for preventing autoreactivity to nuclear self-DNA.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The electron density reconstruction and final model were deposited at the Electron Microscopy Data Bank (EMDB) with accession code EMD-11601, and the Protein Data Bank (PDB) with accession code 7A08. Source data are provided with this paper.
References
Sun, L., Wu, J., Du, F., Chen, X. & Chen, Z. J. Cyclic GMP-AMP synthase is a cytosolic DNA sensor that activates the type I interferon pathway. Science 339, 786–791 (2013).
Ablasser, A. et al. cGAS produces a 2′-5′-linked cyclic dinucleotide second messenger that activates STING. Nature 498, 380–384 (2013).
Gao, P. et al. Cyclic [G(2′,5′)pA(3′,5′)p] is the metazoan second messenger produced by DNA-activated cyclic GMP-AMP synthase. Cell 153, 1094–1107 (2013).
Zhang, X. et al. Cyclic GMP-AMP containing mixed phosphodiester linkages is an endogenous high-affinity ligand for STING. Mol. Cell 51, 226–235 (2013).
Hopfner, K. P. & Hornung, V. Molecular mechanisms and cellular functions of cGAS-STING signalling. Nat. Rev. Mol. Cell Biol. 21, 501–521 (2020).
Ablasser, A. & Chen, Z. J. cGAS in action: Expanding roles in immunity and inflammation. Science 363, eaat8657 (2019).
Motwani, M., Pesiridis, S. & Fitzgerald, K. A. DNA sensing by the cGAS-STING pathway in health and disease. Nat. Rev. Genet. 20, 657–674 (2019).
Volkman, H. E., Cambier, S., Gray, E. E. & Stetson, D. B. Tight nuclear tethering of cGAS is essential for preventing autoreactivity. eLife 8, e47491 (2019).
Jiang, H. et al. Chromatin-bound cGAS is an inhibitor of DNA repair and hence accelerates genome destabilization and cell death. EMBO J. 38, e102718 (2019).
Gentili, M. et al. The N-terminal domain of cGAS determines preferential association with centromeric dna and innate immune activation in the nucleus. Cell Rep. 26, 2377–2393 (2019).
Kranzusch, P. J. cGAS and CD-NTase enzymes: structure, mechanism, and evolution. Curr. Opin. Struct. Biol. 59, 178–187 (2019).
Civril, F. et al. Structural mechanism of cytosolic DNA sensing by cGAS. Nature 498, 332–337 (2013).
Li, X. et al. Cyclic GMP-AMP synthase is activated by double-stranded DNA-induced oligomerization. Immunity 39, 1019–1031 (2013).
Zhang, X. et al. The cytosolic DNA sensor cGAS forms an oligomeric complex with DNA and undergoes switch-like conformational changes in the activation loop. Cell Rep. 6, 421–430 (2014).
Andreeva, L. et al. cGAS senses long and HMGB/TFAM-bound U-turn DNA by forming protein-DNA ladders. Nature 549, 394–398 (2017).
Du, M. & Chen, Z. J. DNA-induced liquid phase condensation of cGAS activates innate immune signaling. Science 361, 704–709 (2018).
Xie, W. et al. Human cGAS catalytic domain has an additional DNA-binding interface that enhances enzymatic activity and liquid-phase condensation. Proc. Natl Acad. Sci. USA 116, 11946–11955 (2019).
Hooy, R. M. & Sohn, J. The allosteric activation of cGAS underpins its dynamic signaling landscape. eLife 7, e39984 (2018).
Herzner, A. M. et al. Sequence-specific activation of the DNA sensor cGAS by Y-form DNA structures as found in primary HIV-1 cDNA. Nat. Immunol. 16, 1025–1033 (2015).
Mackenzie, K. J. et al. cGAS surveillance of micronuclei links genome instability to innate immunity. Nature 548, 461–465 (2017).
Liu, H. et al. Nuclear cGAS suppresses DNA repair and promotes tumorigenesis. Nature 563, 131–136 (2018).
Zierhut, C. et al. The cytoplasmic DNA sensor cGAS promotes mitotic cell death. Cell 178, 302–315 (2019).
McGinty, R. K. & Tan, S. Recognition of the nucleosome by chromatin factors and enzymes. Curr. Opin. Struct. Biol. 37, 54–61 (2016).
Arya, G. & Schlick, T. A tale of tails: how histone tails mediate chromatin compaction in different salt and linker histone environments. J. Phys. Chem. A 113, 4045–4059 (2009).
Kalia, R. et al. Structural basis of mitochondrial receptor binding and constriction by DRP1. Nature 558, 401–405 (2018).
Harding, S. M. et al. Mitotic progression following DNA damage enables pattern recognition within micronuclei. Nature 548, 466–470 (2017).
Dou, Z. et al. Cytoplasmic chromatin triggers inflammation in senescence and cancer. Nature 550, 402–406 (2017).
Seo, G. J. et al. TRIM56-mediated monoubiquitination of cGAS for cytosolic DNA sensing. Nat. Commun. 9, 613 (2018).
Klinker, H., Haas, C., Harrer, N., Becker, P. B. & Mueller-Planitz, F. Rapid purification of recombinant histones. PLoS One 9, e104029 (2014).
Huynh, V. A., Robinson, P. J. & Rhodes, D. A method for the in vitro reconstitution of a defined “30 nm” chromatin fibre containing stoichiometric amounts of the linker histone. J. Mol. Biol. 345, 957–968 (2005).
Rogge, R. A. et al. Assembly of nucleosomal arrays from recombinant core histones and nucleosome positioning DNA. J. Vis. Exp. 79, e50354 (2013).
Rueden, C. T. et al. ImageJ2: ImageJ for the next generation of scientific image data. BMC Bioinformatics 18, 529 (2017).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Rohou, A. & Grigorieff, N. CTFFIND4: fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Scheres, S. H. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Tan, Y. Z. et al. Addressing preferred specimen orientation in single-particle cryo-EM through tilting. Nat. Methods 14, 793–796 (2017).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004).
Goddard, T. D. et al. UCSF ChimeraX: meeting modern challenges in visualization and analysis. Protein Sci. 27, 14–25 (2018).
Schmid-Burgk, J. L. et al. OutKnocker: a web tool for rapid and simple genotyping of designer nuclease edited cell lines. Genome Res. 24, 1719–1723 (2014).
Acknowledgements
We thank K. Schall and V. Niebauer for guidance with nucleosome preparation and A. Drescher for discussions of SPR data. C.C.O.M. is supported as a Cancer Research Institute/Eugene V. Weissman Fellow. This work was supported by the German Research Council (grants CRC/TRR237 and the Gottfried Wilhelm Leibniz-Prize to K.-P.H. and V.H.; RTG1721 to K.-P.H. and G.W.; HO2489/8 to K.-P.H.; WI3717/3-1 to G.W.; and CRC1054 to K.L.).
Author information
Authors and Affiliations
Contributions
C.C.O.M. and S.M. prepared cryo-EM samples and performed biochemical analysis. C.C.O.M., S.M. and J.B. optimized and screened cryo-EM grids. K.L. collected cryo-EM data. C.C.O.M. performed structure determination and model building with assistance from J.B. and K.L. SPR experiments were performed by G.W. C.S. performed cell-based assays and V.H. supervised cell-based experiments and interpreted data. C.C.O.M. and K.-P.H. designed the overall study, analysed the results and wrote the paper with contributions from all other authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks Yuan He and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 cGAS binds nucleosomes even in the presence of free DNA.
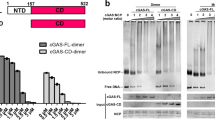
a, Gel-mobility shift binding analysis of purified cGAS and nucleosomes. Nucleosomes without linker DNA (0N0) were tested for binding to purified human (h) and mouse (m) full-length cGAS and cGAScat. Data are representative of two biological replicates. b, Human cGAS activity assay preincubated with dsDNA followed by titration of 0N0 nucleosomes. Data are representative of two biological replicates.
Extended Data Fig. 2 Cryo-EM data processing for cGAS–nucleosome structure.
a, Representative micrograph of the dataset used to determine the cGAS–nucleosome complex structure. b, Left, final reconstruction of the cGAS–nucleosome complex coloured by local resolution. Right, representation of angular distribution of particles contributing to the final map. c, Histogram and directional Fourier shell correlation (FSC) plot for the final 3D reconstitution of the cGAS–nucleosome complex (3.11 Å). A sphericity of 0.9 was determined indicating very isotropic angular distribution (a value of 1 stands for completely isotropic angular distribution). The global resolution was determined to 3.11 Å (0.143 criterion). Directional FSC determination was performed with the 3DFSC software. d, Flow chart for image processing using RELION and cryoSPARC.
Extended Data Fig. 3 Sample density maps for cGAS–nucleosome structure.
a, Representative examples of cryo-EM map areas of cGAS, nucleosomal DNA and histones used for model building. b, Electron density for cGAS (blue) and H2A–H2B (yellow, red) interacting residues in interface I and interface II. cGAS tethering loops 1 and 2 with key interacting residues R222 and R241 are depicted as well as DNA-binding site B residues. Dashed lines represent hydrogen bonds.
Extended Data Fig. 4 Mutational analysis of binding interface between cGAS and the nucleosome.
a, Protein–protein residue interactions across the interface of cGAS with histone H2A and cGAS with histone H2B. Interacting amino acids are joined by coloured lines, each representing a different type of interaction, as per the key below. Interaction maps for the cGAS–nucleosome complex were generated using PDBsum. b, Thermal shift assay derivative melt curve plots of human cGAScat mutants. Respective inflection temperatures are: cGAScat 61.8 °C; cGAScat(K407E/K411E) 61.3 °C; cGAScat(C396A/C397A (55.5 °C; cGAScat(R236E) 61.9 °C, cGAScat(R300E/K301E) 62.5 °C. Data are representative of two biological replicates. c, Coomassie stained SDS–PAGE gels of purified recombinant human and mouse cGAS (7 μg each) constructs used in this study. Gels are representative of one replicate. d, Representative EMSAs for mouse cGAS–nucleosome interface I and interface II mutants binding to fluorescently labelled nucleosomes. Data are representative of three biological replicates. e, EMSAs for mouse cGAS mutants in tethering loops 1 and 2 (R222E, R241E) and DNA-binding site B (R337E, R341E) binding to fluorescently labelled nucleosomal DNA. Data are representative of two biological replicates. f, EMSAs for human cGAS full-length and catalytic domain, DNA-binding site A (K407E/K411E), Zn-thumb (C396A/C397A), site B (R236E) and site C (R300E/K301E) mutants binding to fluorescently labelled nucleosomes. Data are representative of two biological replicates. g, Representative EMSAs for mouse cGAS full-length binding to fluorescently labelled acidic patch mutant nucleosomes apI (H2A(E61A/E64A/D90A)) and apII (H2A(R71A), H2B(H49A/D51A)) and apI + apII (H2A(E61A/E64A/R71A/D90A/E92A), H2B(H49A/D51A)). Data are representative of three biological replicates.
Extended Data Fig. 5 cGAS DNA-binding site B is required for cGAS tethering by the nucleosome.
a, SPR analysis of single-cycle-kinetics experiment with immobilized nucleosomes via biotinylated DNA and mouse cGAScat and cGAScat(R241E) mutant as analytes. Shown are injections of 1.1, 3.3, 9, 10, 30 and 90 nM mouse cGAScat and cGAScat(R241E). Data are representative of two biological replicates. b, SPR analysis with acidic patch mutant nucleosomes (H2A(E61A/E64A/D90A)) immobilized via biotinylated DNA and cGAScat as analyte. Shown are buffer injections, injections of 1.1, 3.3, 9, 10, 30 and 90 nM mouse cGAScat and the cGAScat background-corrected data. Mouse cGAScat has orders of magnitude lower affinity to acidic patch mutant nucleosomes than wild-type nucleosomes. Data are representative of two biological replicates. c, Human cGAS(R236E) mutant activity assay preincubated with dsDNA followed by titration of 0N0 nucleosome. Data are representative of two biological replicates. d, Human cGAS(R236E) mutant activity assay pre-incubated with 0N0 nucleosome followed by titration of dsDNA. Data are representative of two biological replicates. e, Mouse cGAS mutations tested affecting cGAS–nucleosome interactions were tested for DNA-dependent activation with plasmid DNA in the presence of 0N0 nucleosomes. cGAMP production was assayed by thin-layer chromatography. Data are representative of two biological replicates. f, Mouse cGAS mutations affecting cGAS–nucleosome interactions were tested for DNA-dependent activation with 147-bp nucleosomal DNA in the presence of 0N0 nucleosomes. Data are representative of two biological replicates. g, Mouse cGAS single and double mutations of tethering loop and DNA-binding site B were tested for cGAMP production in the presence of plasmid DNA alone or plasmid DNA and 0N0 nucleosome. Data are representative of two biological replicates. h, Mouse cGAS mutants R222E, K240E and R241E require plasmid DNA for activation. Data are representative of two biological replicates. i, Mouse cGAS mutants R222E, K240E and R241E were tested for DNA-dependent activation with plasmid DNA in the presence of 0N0 nucleosomes. Mutation of DNA-binding site A abolishes activation by plasmid DNA. Data are representative of two biological replicates. j, Human cGAS activity assay in the presence of dsDNA, followed by titration of 0N0 and 0N0 acidic patch mutant I (apI; H2A(E61A/E64A/D90A)). Data are representative of two biological replicates. k, Wild-type and R236E mutant cGAS activity assays with 40N40 nucleosomes (40-bp linker DNA on each side). Data are representative of two biological replicates. l, Fluorescence anisotropy analysis of human cGAScat human cGAScat site A mutant (K407E/K411E) binding to fluorescently labelled 20-bp dsDNA and in the presence of 0N0 nucleosomes. Data are representative of two biological replicates. m, Agarose gel of micrococcal nuclease (MNase)-digested synthetic 601 chromatin indicating a regular nucleosomal structure. Data are representative of two biological replicates.
Extended Data Fig. 6 IP-10 cytokine and ISG production upon self and non-self DNA recognition.
a, PMA-differentiated wild-type or knockout human CGAS−/− THP-1 cells were treated with doxycycline (1 μg ml−1) overnight and either left untreated or treated with HT-DNA (975 ng per 550 μl) for 8 h. Cell lysates were separated on SDS–PAGE gels, western blotted and probed with the indicated antibodies. Data are representative of three biological replicates. b, PMA-differentiated THP-1 cells were left untreated or treated with doxycycline overnight to express the indicated human cGAS mutants, followed by stimulation using HT-DNA (200 ng per well) for 8 h. Supernatant was collected and IP-10 cytokines were measured from supernatant using ELISA. Data are mean and s.e.m. of four biological replicates. ***P < 0.001, **P < 0.01, two-way ANOVA. ns, not significant. Doxycycline R236E P < 0.001; doxycycline R255E P < 0.001; doxycycline + HT-DNA R236E P = 0.002; doxycycline + HT-DNA R255E P < 0.001.
Extended Data Fig. 7 Structurally conserved loop for protein–protein interactions in MAB21 family nucleotidyltransferases.
Structures of cGAS–nucleosome (blue) and MID49–DRP1 (red, PDB code 5WP9). The structurally conserved loop is depicted in green, showing interacting residues. Dashed lines represent hydrogen bonds. Positively charged, basic amino acids on the nucleotidyltransferase loop interact with the acidic patch of the nucleosome or with the protein DRP1, respectively.
Supplementary information
Rights and permissions
About this article
Cite this article
Michalski, S., de Oliveira Mann, C.C., Stafford, C. et al. Structural basis for sequestration and autoinhibition of cGAS by chromatin. Nature 587, 678–682 (2020). https://doi.org/10.1038/s41586-020-2748-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-2748-0
This article is cited by
-
DNA sensing and repair systems unexpectedly team up against cancer
Nature (2024)
-
TXNRD1 drives the innate immune response in senescent cells with implications for age-associated inflammation
Nature Aging (2024)
-
DAMP sensing and sterile inflammation: intracellular, intercellular and inter-organ pathways
Nature Reviews Immunology (2024)
-
The CRL5–SPSB3 ubiquitin ligase targets nuclear cGAS for degradation
Nature (2024)
-
When DNA-damage responses meet innate and adaptive immunity
Cellular and Molecular Life Sciences (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.