Abstract
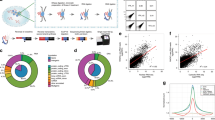
Highly structured RNA molecules usually interact with each other, and associate with various RNA-binding proteins, to regulate critical biological processes. However, RNA structures and interactions in intact cells remain largely unknown. Here, by coupling proximity ligation mediated by RNA-binding proteins with deep sequencing, we report an RNA in situ conformation sequencing (RIC-seq) technology for the global profiling of intra- and intermolecular RNA–RNA interactions. This technique not only recapitulates known RNA secondary structures and tertiary interactions, but also facilitates the generation of three-dimensional (3D) interaction maps of RNA in human cells. Using these maps, we identify noncoding RNA targets globally, and discern RNA topological domains and trans-interacting hubs. We reveal that the functional connectivity of enhancers and promoters can be assigned using their pairwise-interacting RNAs. Furthermore, we show that CCAT1-5L—a super-enhancer hub RNA—interacts with the RNA-binding protein hnRNPK, as well as RNA derived from the MYC promoter and enhancer, to boost MYC transcription by modulating chromatin looping. Our study demonstrates the power and applicability of RIC-seq in discovering the 3D structures, interactions and regulatory roles of RNA.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
RIC-seq and hnRNPK CLIP-seq data that support the findings of this study have been deposited in the GEO under accession number GSE127188.
Code availability
Custom codes used for data analysis in this paper can be found at https://github.com/caochch/RIC-seq.
References
Butcher, S. E. & Pyle, A. M. The molecular interactions that stabilize RNA tertiary structure: RNA motifs, patterns, and networks. Acc. Chem. Res. 44, 1302–1311 (2011).
Gerstberger, S., Hafner, M. & Tuschl, T. A census of human RNA-binding proteins. Nat. Rev. Genet. 15, 829–845 (2014).
Aw, J. G. et al. In vivo mapping of eukaryotic RNA interactomes reveals principles of higher-order organization and regulation. Mol. Cell 62, 603–617 (2016).
Kudla, G., Granneman, S., Hahn, D., Beggs, J. D. & Tollervey, D. Cross-linking, ligation, and sequencing of hybrids reveals RNA–RNA interactions in yeast. Proc. Natl Acad. Sci. USA 108, 10010–10015 (2011).
Lu, Z. et al. RNA duplex map in living cells reveals higher-order transcriptome structure. Cell 165, 1267–1279 (2016).
Nguyen, T. C. et al. Mapping RNA–RNA interactome and RNA structure in vivo by MARIO. Nat. Commun. 7, 12023 (2016).
Ramani, V., Qiu, R. & Shendure, J. High-throughput determination of RNA structure by proximity ligation. Nat. Biotechnol. 33, 980–984 (2015).
Sharma, E., Sterne-Weiler, T., O’Hanlon, D. & Blencowe, B. J. Global mapping of human RNA–RNA interactions. Mol. Cell 62, 618–626 (2016).
Sugimoto, Y. et al. hiCLIP reveals the in vivo atlas of mRNA secondary structures recognized by Staufen 1. Nature 519, 491–494 (2015).
Morf, J. et al. RNA proximity sequencing reveals the spatial organization of the transcriptome in the nucleus. Nat. Biotechnol. 37, 793–802 (2019).
Fazal, F. M. et al. Atlas of subcellular RNA localization revealed by APEX-seq. Cell 178, 473–490.e426 (2019).
Anger, A. M. et al. Structures of the human and Drosophila 80S ribosome. Nature 497, 80–85 (2013).
Durand, N. C. et al. Juicebox provides a visualization system for Hi-C contact maps with unlimited zoom. Cell Syst. 3, 99–101 (2016).
West, J. A. et al. The long noncoding RNAs NEAT1 and MALAT1 bind active chromatin sites. Mol. Cell 55, 791–802 (2014).
Ray, D. et al. A compendium of RNA-binding motifs for decoding gene regulation. Nature 499, 172–177 (2013).
Whyte, W. A. et al. Master transcription factors and mediator establish super-enhancers at key cell identity genes. Cell 153, 307–319 (2013).
Kim, T. K. & Shiekhattar, R. Architectural and functional commonalities between enhancers and promoters. Cell 162, 948–959 (2015).
Dixon, J. R. et al. Topological domains in mammalian genomes identified by analysis of chromatin interactions. Nature 485, 376–380 (2012).
Nora, E. P. et al. Spatial partitioning of the regulatory landscape of the X-inactivation centre. Nature 485, 381–385 (2012).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
Bolzer, A. et al. Three-dimensional maps of all chromosomes in human male fibroblast nuclei and prometaphase rosettes. PLoS Biol. 3, e157 (2005).
Xiang, J. F. et al. Human colorectal cancer-specific CCAT1-L lncRNA regulates long-range chromatin interactions at the MYC locus. Cell Res. 24, 513–531 (2014).
Tseng, Y. Y. et al. PVT1 dependence in cancer with MYC copy-number increase. Nature 512, 82–86 (2014).
Michelotti, E. F., Michelotti, G. A., Aronsohn, A. I. & Levens, D. Heterogeneous nuclear ribonucleoprotein K is a transcription factor. Mol. Cell. Biol. 16, 2350–2360 (1996).
Backe, P. H., Messias, A. C., Ravelli, R. B., Sattler, M. & Cusack, S. X-ray crystallographic and NMR studies of the third KH domain of hnRNP K in complex with single-stranded nucleic acids. Structure 13, 1055–1067 (2005).
Xue, Y. et al. Direct conversion of fibroblasts to neurons by reprogramming PTB-regulated microRNA circuits. Cell 152, 82–96 (2013).
Chen, J. et al. The RNA-binding protein ROD1/PTBP3 cotranscriptionally defines AID-loading sites to mediate antibody class switch in mammalian genomes. Cell Res. 28, 981–995 (2018).
Engreitz, J. M. et al. RNA–RNA interactions enable specific targeting of noncoding RNAs to nascent Pre-mRNAs and chromatin sites. Cell 159, 188–199 (2014).
Qin, G. et al. Long noncoding RNA p53-stabilizing and activating RNA promotes p53 signaling by inhibiting heterogeneous nuclear ribonucleoprotein K deSUMOylation and suppresses hepatocellular carcinoma. Hepatology 71, 112–129 (2020).
Orjalo, A. V., Jr & Johansson, H. E. Stellaris® RNA fluorescence in situ hybridization for the simultaneous detection of immature and mature long noncoding RNAs in adherent cells. Methods Mol. Biol. 1402, 119–134 (2016).
Wang, X. W. et al. A microRNA-inducible CRISPR–Cas9 platform serves as a microRNA sensor and cell-type-specific genome regulation tool. Nat. Cell Biol. 21, 522–530 (2019).
Gilbert, L. A. et al. CRISPR-mediated modular RNA-guided regulation of transcription in eukaryotes. Cell 154, 442–451 (2013).
Zarnegar, B. J. et al. irCLIP platform for efficient characterization of protein–RNA interactions. Nat. Methods 13, 489–492 (2016).
Hagège, H. et al. Quantitative analysis of chromosome conformation capture assays (3C–qPCR). Nat. Protocols 2, 1722–1733 (2007).
Chu, C. et al. Systematic discovery of Xist RNA binding proteins. Cell 161, 404–416 (2015).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnetJ. 17, 10–12 (2011).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Robinson, J. T. et al. Integrative Genomics Viewer. Nat. Biotechnol. 29, 24–26 (2011).
Xue, Y. et al. Genome-wide analysis of PTB–RNA interactions reveals a strategy used by the general splicing repressor to modulate exon inclusion or skipping. Mol. Cell 36, 996–1006 (2009).
Darty, K., Denise, A. & Ponty, Y. VARNA: interactive drawing and editing of the RNA secondary structure. Bioinformatics 25, 1974–1975 (2009).
Almada, A. E., Wu, X., Kriz, A. J., Burge, C. B. & Sharp, P. A. Promoter directionality is controlled by U1 snRNP and polyadenylation signals. Nature 499, 360–363 (2013).
Lestrade, L. & Weber, M. J. snoRNA-LBME-db, a comprehensive database of human H/ACA and C/D box snoRNAs. Nucleic Acids Res. 34, D158–D162 (2006).
Nojima, T. et al. Deregulated expression of mammalian lncRNA through loss of SPT6 induces R-loop formation, replication stress, and cellular senescence. Mol. Cell 72, 970–984 (2018).
Kim, D., Langmead, B. & Salzberg, S. L. HISAT: a fast spliced aligner with low memory requirements. Nat. Methods 12, 357–360 (2015).
Trapnell, C. et al. Transcript assembly and quantification by RNA-seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat. Biotechnol. 28, 511–515 (2010).
Gong, J. et al. RISE: a database of RNA interactome from sequencing experiments. Nucleic Acids Res. 46, D194–D201 (2018).
Krüger, J. & Rehmsmeier, M. RNAhybrid: microRNA target prediction easy, fast and flexible. Nucleic Acids Res. 34, W451–W454 (2006).
Bailey, T. L. et al. MEME SUITE: tools for motif discovery and searching. Nucleic Acids Res. 37, W202–W208 (2009).
Rao, S. S. et al. A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping. Cell 159, 1665–1680 (2014).
Lu, X. J., Bussemaker, H. J. & Olson, W. K. DSSR: an integrated software tool for dissecting the spatial structure of RNA. Nucleic Acids Res. 43, e142 (2015).
Reuter, J. S. & Mathews, D. H. RNAstructure: software for RNA secondary structure prediction and analysis. BMC Bioinformatics 11, 129 (2010).
Zhang, Y. et al. Model-based analysis of ChIP–seq (MACS). Genome Biol. 9, R137 (2008).
Li, X. et al. GRID-seq reveals the global RNA–chromatin interactome. Nat. Biotechnol. 35, 940–950 (2017).
Ørom, U. A. et al. Long noncoding RNAs with enhancer-like function in human cells. Cell 143, 46–58 (2010).
Grant, C. E., Bailey, T. L. & Noble, W. S. FIMO: scanning for occurrences of a given motif. Bioinformatics 27, 1017–1018 (2011).
Acknowledgements
We thank A. M. Pyle, B. Zhou and J. Hu for critical reading of this manuscript; Y. Wang for sharing the PiggyBac-Cas9 system; G. Shan for assistance with the DNA FISH; L.-L. Chen for HT29 cells; G. Zhou for sharing the hnRNPK expression plasmid; and Y. Wang, Y. Feng and S. Li for SIM imaging and analysis. This work was supported by the National Natural Science Foundation of China (91740201, 91940306, 31522015 and 81921003), the Ministry of Science and Technology of China (2017YFA0504400 and 2016YFC0900400) and the Strategic Priority Program of CAS (XDB19000000) to Y.X., by the Beijing Municipal Natural Science Foundation (5182024) to C.C., and by the National Natural Science Foundation of China (31771438, 31970610) to L.J.
Author information
Authors and Affiliations
Contributions
Y.X. initiated and planned the study; Z.C. cultured cells, constructed clones, performed smFISH and created the RIC-seq library; L.J. performed Cas9 deletion, CRISPRi, DNA FISH, ChIRP-MS and functional validation of CCAT1-5L with the help of C.X. and S.W.; C.C. performed the bioinformatics analysis, led the project and prepared the figures with the help of Z.D. and N.H.; R.Y. tested RIC-seq conditions, validated enhancer–promoter interactions and constructed hnRNPK CLIP-seq libraries; D.W. performed 3C-qPCR analysis; J.C. and X. Yu performed co-IP and ChIP–qPCR experiments; S.H., L.W. and X. Yang advised on bioinformatics analysis, Y.X., Z.C. and C.C. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
Z.C., C.C. and Y.X. have filled a joint application for the patent of RIC-seq technology.
Additional information
Peer review information Nature thanks Scott Tenenbaum, Jernej Ule and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Characterization of RIC-seq technology.
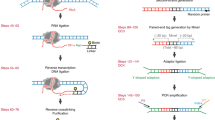
a, The RNA fragment sizes were quantified using a 2100 Bioanalyzer. Top, total RNA after rRNA depletion. Middle, RNA after fragmentation. Bottom, RNA after C1 beads pull-down. b, c, The RIC-seq libraries were purified from the gel (the bracketed region) and quantified using a 2100 Bioanalyzer. −pCp denotes samples without pCp–biotin labelling. d, The mapping pipeline for RIC-seq data. e, Read types and their relationship to reference transcripts. f, The numbers of intra- and intermolecular chimeric reads. g, The intra- and intermolecular chimeric reads show a 4.2- and 5.5-fold increase, respectively, in the intron-to-exon ratio, as compared with RNA-seq data. h, The intra- and intermolecular RIC-seq signals do not decrease after the branch point (red star). i, Transcripts enriched in different subcellular fractions revealed by RNA-seq45 in HeLa cells. Two replicates for each fraction (rep 1 and rep 2). j, The number of RNAs enriched (blue) in each subcellular fraction and the numbers of RNAs that could be captured by RIC-seq (red). k, Cartoon of RBP-mediated RNA proximal interactions and data presentation. The structural models of the RBPs were generated in PyMOL on the basis of PDB accessions 5B16 and 4V6X. Light blue lines between chimeric reads represent gaps. Pairwise junctions are visualized as arc lines or plotted as a heat map. Colour intensity indicates the number of junctions within each 5 × 5-nt pixels. l, Scatter plots showing the reproducibility of RIC-seq-detected pairwise interactions in two biological replicates. The reproducibility for chimeric reads per transcript (R = 0.932) is displayed in a red box. RNA abundance is normalized for both scatter plots. m, Heat map showing the cross-species RNA–RNA interactions observed by the cell mixing strategy. The boxed region in dashed lines represents random ligations. The experiments in b, c were independently repeated three times with similar results.
Extended Data Fig. 2 RIC-seq recapitulates known and newly identified RNA–RNA interactions.
a–c, RIC-seq recapitulates known structures of tRNA, U3 snoRNA and RPPH1 lncRNA. Known structural elements are marked above specific arc lines. For U3 snoRNA, only three clusters of chimeric reads that support stem–loops 1 to 3 are shown. Mismatches based on variation from the reference transcriptome sequence are marked as red dots in b (C>T), and green dots in c (G>A). d, RIC-seq recapitulates known intra- and intermolecular interactions of U4 and U6 snRNAs. Red dots, C>T mismatches. e, RIC-seq recapitulates snoRNA-interacting sites in 28S rRNA. The red arrow indicates known modification sites. The boxed region represents the D′ box. f–h, PARIS signal, RIC-seq signal and U1-motif density in three different groups of U1–MALAT1 contact sites. Common regions, n = 11; RIC-seq-specific regions, n = 8; PARIS-specific regions, n = 2. i, The genomic distribution of snoRNA-interacting sites detected by RIC-seq. j, The SNORD22 interacting sites in SPHK2 and BCL2L2 RNA. The D box is shown in blue. k, qPCR showing reduced mRNA levels of BCL2L2 and SPHK2 upon knockdown of SNORD22 with ASOs. LENG8 and PM20D2 served as negative controls. Data are mean ± s.d., n = 3 biological replicates. Two-tailed unpaired t-test was used to calculate the P values in f–h, k.
Extended Data Fig. 3 Comparison with existing methods.
a, The sensitivity, accuracy and resolution of existing methods for detecting the structure of pre-miRNA. Sensitivity, the number of detected pre-miRNAs; accuracy, the percentage of chimeric reads that support correct stem–loop structures; resolution, the mean distance of chimeric junctions from the apical loop. Notably, RNA proximity ligation (RPL) did not detect pre-miRNA. N.A., not available. b, U1 binding sites in MALAT1 detected by mapping RNA interactome in vivo (MARIO), ligation of interacting RNA followed by high-throughput sequencing (LIGR-seq), sequencing of psoralen crosslinked, ligated, and selected hybrids (SPLASH) and RPL methods. U1 motifs, purple lines; y axis, chimeric read number. c, Venn diagram showing overlapping duplexes detected by RIC-seq and PARIS methods in different cell lines. RNAs with fragments per kilobase of transcript per million mapped reads (FPKM) values ≥ 1 in both cell types are used for the analysis. d, The transcriptomic span distance of RIC-seq-specific (red, n = 69,351) and overlapped clusters (blue, n = 10,951). The transcriptomic span excludes introns. Two-sided Kolmogorov–Smirnov test was used to calculate the P value. e, Box plot showing the minimum free energy (MFE) of duplexes among RIC-seq-specific and overlapped groups. f, Venn diagram showing the proximal RNAs detected by RIC-seq and proximity RNA-seq in two different cell lines. Two-sided Fisher’s exact test was used to calculate the P value. g, Cartoon depicting the number of colocalized RNAs in four subcellular compartments revealed by APEX-seq. Dashed lines, in silico random contacts. h, The percentage of intracompartment RNA–RNA interactions observed by RIC-seq (n = 71) is higher than that of random contacts. The one-sided binomial test was used to calculate the P value. i, Box plot showing that the RIC-seq signal strength of intracompartment RNA–RNA interactions is stronger than for intercompartment interactions. Two-tailed unpaired t-test was used to calculate the P values in e and i. For the box plots in e and i, the centre line represents the median, the box borders represent the first (Q1) and third (Q3) quartiles, and the whiskers are the most extreme data points within 1.5× the interquartile range (from Q1 to Q3).
Extended Data Fig. 4 RIC-seq precisely recaptures rRNA and lncRNA 3D interactions.
a, The chimeric reads mapped to rRNA. b, The RNA–RNA interactions of 28S rRNA in pCp- samples. c, The RIC-seq and cryo-EM data of 28S rRNA are highly correlated. The RIC-seq signal markedly decreases with increased distance to known pairwise-interacting sites in 28S rRNA (x axis) and with the increased spatial distance (y axis). d, The true-positive and true-negative datasets are generated from the cryo-EM model of 28S rRNA. e, The boundaries of RNA duplexes in 28S rRNA tend to be occupied by RBPs. The spatial distance to the nearest amino acid is calculated from the cryo-EM model. Regions absent from the structure are shown in light grey. f, RIC-seq captures the G4000 duplex protruding from the RBP (in different colours) complex. The arrowhead represents MNase random cut and pCp–biotin labelling position. g, The intermolecular interactions between 5.8S, 18S and 28S rRNA. The histogram on the inner circle represents the number of chimeric reads at the given positions. The red arc lines marked interactions between 5.8S and 28S rRNA are shown on the right. h, The modelled structure of 28S rRNA (PDB ID 4V6X). i, The secondary structure deduced from RIC-seq data for two missing 28S rRNA regions (nt 2951–3246 and nt 3301–3561). The bases are marked by different colours based on in vivo click selective 2-hydroxyl acylation and profiling experiment (icSHAPE) scores, and the base pairs are denoted by different lines on the basis of the strength of the RIC-seq signal. j–l, The structural model, physical interaction map and RIC-seq interaction map of 7SK 5′-hairpin, SRP (7SL RNA) and RPPH1. Grey regions in the physical maps indicate no structural data available. m, RIC-seq showed comparable performance to PARIS in detecting the structure of 7SK, SRP RNA and RPPH1. Dashed line, random classifier.
Extended Data Fig. 5 Monte Carlo simulation to identify significant RNA–RNA interactions and lncRNA targets.
a, Intermolecular RNA–RNA interactions (n = 2,088,874) were plotted against the average pairwise interaction counts from 100,000 simulations. b, A single random dataset (n = 3,536,556) was taken for further 100,000 simulations and plotted as in a. c, The observed pairwise interactions were compared with simulated random counts to identify high-confidence interactions (P ≤ 0.05, red dots). d, Box plot showing how the P value distribution changed as the Monte Carlo simulation progressed using n = 2,088,874 intermolecular RNA–RNA interaction events. The 100,000 simulations were divided into 100 batches. e, RNA–RNA interaction types and their numbers of RIC-seq chimeric reads. f, The violin plot shows the expression levels of MALAT1- and NEAT1-targeting genes. g, The enriched motifs among MALAT1 or NEAT1 chimeric targets. h, Overlaps of MALAT1 and NEAT1 binding sites identified by RIC-seq and CHART-seq. Two-sided Fisher’s exact test was used to calculate the P value. i, Summary of NEAT1 foci and their overlap with MALAT1 in 15 cells. j, Structured illumination microscopy analysis showing the localization of MALAT1 with NEAT1 5′ region, NEAT1 middle region or NEAT1 3′ non-contact region. The regions marked by white boxes are magnified at the top right. The violin plot illustrates the distance from NEAT1 foci to the nearest MALAT1 puncta in 20 cells. 5′ end, n = 131; middle, n = 169; 3′ non-contact region, n = 166. Two-tailed unpaired t-test was used to calculate the P values in f and j. For the violin or box plots in d, f and j, the white centre point represents the median, the box limits represent the Q1 and Q3, the whiskers are the most extreme data points within 1.5 × the interquartile range (from Q1 to Q3), and the upper–lower limits represent the maximum–minimum values.
Extended Data Fig. 6 The features of RNA topological domain.
a, The number and distribution of topological domains in genes. b, The length distribution of the topological domains. Dashed line, median length. c, The domain boundaries frequently overlap with exons. Grey line, random control. d, The RBP binding pattern around topological domains (black box) in pre-mRNA and pre-lncRNA. CLIP-seq coverage of each RBP is shown at the bottom. e, RBP binding profiles are plotted around the domain boundary. The dashed lines in colours represent random controls.
Extended Data Fig. 7 The characterization of hub RNAs.
a, The classification of hub RNAs. b, Hub lncRNA (n = 29) show stronger trans interaction strength than hub pre-mRNA (n = 610). c–i, Comparing the expression levels, RIC-seq signals, conservation, Pol II signals, H3K4me3 signals, H3K27ac signals and H3K27me3 signals of hub RNA (n = 642) to control RNA (n = 5,466). Two-tailed unpaired t-test was used to calculate the P values in b and c. The two-sided Kolmogorov–Smirnov test was used to calculate the P values in d–i. j, t-SNE visualization of 642 hub RNAs on the basis of the strength of their interaction with target RNAs. k, Hub RNAs from the same chromosome tend to interact with each other. l, Expression pattern of hub RNAs in different subcellular fractions. m, Hub RNAs prefer to interact with each other in the same subcellular fraction (chromatin, n = 493; nucleoplasmic, n = 61; cytoplasmic, n = 80). n, o, Hub RNAs and their targets are classified into three distinct clusters based on 187 RBP motifs. p, Hub RNAs in the same cluster tends to interact with each other. Cluster 1 to 3, n = 292, 79, 271. q, Hub RNAs in the same and different Gene Ontology terms are clustered on the basis of their relative interaction strength. From 1 to 12, n = 186, 379, 253, 51, 213, 65, 79, 206, 61, 108, 54 and 59. r, Over-represented motifs in super-enhancer-related hub RNAs. s, RBPs enriched in super-enhancer-related hub RNAs. t, The eCLIP peaks enriched in super-enhancer-related hub RNAs. For the violin plots in b and c, the white centre point represents the median, the box limits represent Q1 and Q3, the whiskers are the most extreme data points within 1.5× the interquartile range (from Q1 to Q3), and the upper–lower limits represent the maximum–minimum values. The two-sided binomial test was used to calculate the P values shown in k, m, p and q.
Extended Data Fig. 8 Features and validation of enhancer–promoter interaction.
a, Pie chart shows the strand preference of eRNAs interacting with promoter RNAs. b, eRNAs tend to interact with other RNAs within TADs. Colour intensity indicates read density in 40-kb windows. The purple line denotes the boundary of TADs. c, Circos plot showing whole-genome enhancer–promoter contacts on the basis of their pairwise-interacting RNAs. Red circle, super-enhancers; blue circle, typical enhancers; yellow circle, promoters. Red and light green arc lines illustrate inferred super-enhancer–promoter and typical-enhancer–promoter interactions, respectively. d, qPCR validation of enhancer–promoter interactions at four typical enhancers upon depletion of eRNAs with LNA ASOs. Specific-enhancer-linked promoter reads are shown as blue arc lines above the genes at positive and negative strands. C5AR1, LRSAM1, AK1, SPTAN1, ZDHHC12, LINC02398, SLCO1C1 and SLCO1B1 served as locus-specific controls. The relative fold change is normalized to the LNA control. e, Inferred enhancer–promoter and promoter–promoter interaction networks on chromosome 8. qPCR showing the expression level of super-enhancer-638-linked genes (box) upon depletion of super-enhancer RNAs with LNA ASOs. PCAT1 and GSDMC served as locus-specific negative controls. Data in d, e are mean ± s.d.; n = 3 biological replicates, two-tailed, unpaired t-test.
Extended Data Fig. 9 Characterization of CCAT1-5L and its interaction partner.
a, Genotyping results showing no integration of human papillomavirus at CCAT1-5L locus. b, Northern blot analysis of CCAT1-5L expression across diverse cell lines. c, Western blot showing the purity of nuclear and cytoplasmic fractions. d, e, CCAT1-5L is localized in the nucleus revealed by qPCR and smFISH. CCAT1-E2, CCAT1 exon 2. f, DNA FISH showing CCAT1-5L, MYC and PVT1 loci are colocalized. g, qPCR showing efficient knockdown of hub RNAs DANT2 and PDE3A with LNA ASOs. Non-targeting LNAs, NC-a and NC-b. h, qPCR showing reduced expression of CCAT1-5L, CCAT1-E2, MYC and PVT1 upon CCAT1-5L knockdown with LNA ASOs. i, Western blot showing reduced MYC levels upon CCAT1-5L knockdown. j, Western blot showing reduced MYC levels upon the deletion of the extra extended region of CCAT1-5L by CRISPR–Cas9. k, qPCR showing reduced levels of CCAT1-5L and MYC in mutant cell lines. l, A cartoon depicts the transcriptional blocking of the extra extended region of CCAT1-5L by CRISPRi. m, qPCR showing the expression levels of CCAT1-5L, MYC and CCAT1 exon 1 and 2 (CCAT1 (E1 + E2)) upon CRISPRi. n, Western blot showing reduced MYC levels upon the CRISPRi of CCAT1-5L. o, Motifs enriched in CCAT-5L, MYC and PVT1. p, Enriched motifs in CCAT1-5L RNA. q, HnRNPK monomer and dimer CLIP-seq are highly correlated in HeLa cells (n = 313,762 windows). r, Consensus motifs identified by hnRNPK CLIP-seq. s, 3C–qPCR analysis of the long-distance interactions at the CCAT1-5L, MYC and PVT1 loci upon hnRNPK knockdown with siRNA. t, Co-IP showing hnRNPK (T389A/Q391A) mutant (Flag_K MT) did not form a dimer in HeLa cells. Data in d, g, h, k, m are mean ± s.d.; Data in s are mean ± s.e.m.; n = 3 biological replicates, two-tailed, unpaired t-test. The experiments in b, c, e, f, i, j, n, t were independently repeated three times with similar results.
Extended Data Fig. 10 CCAT1-5L promotes cell proliferation and metastasis via MYC.
a, smFISH showing the ectopically expressed CCAT1-5L lncRNA and MYC promoter RNA are colocalized. CCAT1-5L, red; MYC, green; DAPI, blue. b, Dual RNA–DNA FISH showing the ectopically expressed CCAT1-5L RNA is colocalized with MYC locus. CCAT1-5L lncRNA, red; MYC locus, green; DAPI, blue. c, Western blot showing the ectopically expressed CCAT1-5L boosts MYC levels in HeLa cells. GAPDH served as a loading control. The MYC level is quantified in the middle. d, The depletion or ectopic expression of CCAT1-5L (5L) in HeLa cells influences the proliferation rate. e, Knockdown or ectopic expression of CCAT1-5L affects colony formation. f, CCAT1-5L is critical for cell metastasis in a transwell assay. Scale bar, 50 μm. The experiments in a–c were independently repeated three times with similar results. Data in d–f are mean ± s.d.; n = 3 biological replicates, two-tailed, unpaired t-test.
Supplementary information
Supplementary Figure
Supplementary Figure 1. Original gel source data. This figure contains original gel source data for the western blots and northern blot, with molecular weight markers and an indication of how the gels were cropped.
Supplementary Information
Supplementary Note. This file contains a discussion of the challenges for the experimental study of in vivo RNA structure and interactions.
Supplementary Table
Supplementary Table 1. The mapping results of RIC-seq libraries. This table contains the numbers of raw reads, clean reads, chimeric reads, and chimeric percentage.
Supplementary Table
Supplementary Table 2. RNA enriched in different subcellular fractions. This table contains the transcripts that are preferably enriched in chromatin, nucleoplasmic, or cytoplasmic fractions revealed by RNA-seq.
Supplementary Table
Supplementary Table 3. Comparison between RIC-seq and existing methods. This table contains a comparison of RIC-seq with previous methods, including PARIS, MARIO, LIGR-seq, SPLASH, and RPL.
Supplementary Table
Supplementary Table 4. Summary of RNA-RNA interactions. This table contains the numbers of intra- and inter-molecular RNA-RNA interactions detected by RIC-seq in HeLa cells.
Supplementary Table
Supplementary Table 5. List of MALAT1, NEAT1, and CCAT1 targets in vivo. This table contains genomic positions of MALAT1, NEAT1, and CCAT1 interacting RNA targets.
Supplementary Table
Supplementary Table 6. Summary of RNA topological domains. This table contains genomic positions of identified RNA topological domains in HeLa cells.
Supplementary Table
Supplementary Table 7. Summary of hub RNAs. This table contains genomic positions of identified hub RNAs in HeLa cells.
Supplementary Table
Supplementary Table 8. List of RIC-seq detected enhancer-promoter and promoter-promoter interactions. This table contains genomic positions of RIC-seq deduced enhancer-promoter and promoter-promoter interactions in HeLa cells.
Supplementary Table
Supplementary Table 9. List of primers and probes used in this study. This table contains the sequence of primers, probes, LNA oligos, and siRNAs used in this study.
Rights and permissions
About this article
Cite this article
Cai, Z., Cao, C., Ji, L. et al. RIC-seq for global in situ profiling of RNA–RNA spatial interactions. Nature 582, 432–437 (2020). https://doi.org/10.1038/s41586-020-2249-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-2249-1
This article is cited by
-
KARR-seq reveals cellular higher-order RNA structures and RNA–RNA interactions
Nature Biotechnology (2024)
-
Exploring ethylene-related genes in Cannabis sativa: implications for sexual plasticity
Plant Reproduction (2024)
-
RNA structure: implications in viral infections and neurodegenerative diseases
Advanced Biotechnology (2024)
-
3D genomics and its applications in precision medicine
Cellular & Molecular Biology Letters (2023)
-
A novel super-enhancer-related gene signature predicts prognosis and immune microenvironment for breast cancer
BMC Cancer (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.