Abstract
Checkpoint blockade therapies have improved cancer treatment, but such immunotherapy regimens fail in a large subset of patients. Conventional type 1 dendritic cells (DC1s) control the response to checkpoint blockade in preclinical models and are associated with better overall survival in patients with cancer, reflecting the specialized ability of these cells to prime the responses of CD8+ T cells1,2,3. Paradoxically, however, DC1s can be found in tumours that resist checkpoint blockade, suggesting that the functions of these cells may be altered in some lesions. Here, using single-cell RNA sequencing in human and mouse non-small-cell lung cancers, we identify a cluster of dendritic cells (DCs) that we name ‘mature DCs enriched in immunoregulatory molecules’ (mregDCs), owing to their coexpression of immunoregulatory genes (Cd274, Pdcd1lg2 and Cd200) and maturation genes (Cd40, Ccr7 and Il12b). We find that the mregDC program is expressed by canonical DC1s and DC2s upon uptake of tumour antigens. We further find that upregulation of the programmed death ligand 1 protein—a key checkpoint molecule—in mregDCs is induced by the receptor tyrosine kinase AXL, while upregulation of interleukin (IL)-12 depends strictly on interferon-γ and is controlled negatively by IL-4 signalling. Blocking IL-4 enhances IL-12 production by tumour-antigen-bearing mregDC1s, expands the pool of tumour-infiltrating effector T cells and reduces tumour burden. We have therefore uncovered a regulatory module associated with tumour-antigen uptake that reduces DC1 functionality in human and mouse cancers.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All mice sequencing data are publicly available (GEO accession code GSE131957). All human sequencing data is available on NCBI with BioProject ID PRJNA609924.
Code availability
Scripts to reproduce clustering and differential expression analyses, as well as for direct reproduction of figures related to computational results, are available at https://github.com/effiken/Maier_et_al_nature_2020.
Change history
05 June 2020
A Correction to this paper has been published: https://doi.org/10.1038/s41586-020-2326-5
References
Sánchez-Paulete, A. R. et al. Intratumoral immunotherapy with XCL1 and sFlt3L encoded in recombinant Semliki Forest Virus-derived fosters dendritic cell-mediated T-cell cross-priming. Cancer Res. 78, 6643–6654 (2018).
Salmon, H. et al. Expansion and activation of CD103+ dendritic cell progenitors at the tumor site enhances tumor responses to therapeutic PD-L1 and BRAF inhibition. Immunity 44, 924–938 (2016).
Broz, M. L. et al. Dissecting the tumor myeloid compartment reveals rare activating antigen-presenting cells critical for T cell immunity. Cancer Cell 26, 638–652 (2014).
Lavin, Y. et al. Innate immune landscape in early lung adenocarcinoma by paired single-cell analyses. Cell 169, 750–765 (2017).
Ginhoux, F. et al. The origin and development of nonlymphoid tissue CD103+ DCs. J. Exp. Med. 206, 3115–3130 (2009).
Sathaliyawala, T. et al. Mammalian target of rapamycin controls dendritic cell development downstream of Flt3 ligand signaling. Immunity 33, 597–606 (2010).
Miller, J. C. et al. Deciphering the transcriptional network of the dendritic cell lineage. Nat. Immunol. 13, 888–899 (2012).
Idoyaga, J. et al. Specialized role of migratory dendritic cells in peripheral tolerance induction. J. Clin. Invest. 123, 844–854 (2013).
Steinman, R. M., Hawiger, D. & Nussenzweig, M. C. Tolerogenic dendritic cells. Annu. Rev. Immunol. 21, 685–711 (2003).
Ohl, L. et al. CCR7 governs skin dendritic cell migration under inflammatory and steady-state conditions. Immunity 21, 279–288 (2004).
Helft, J. et al. Cross-presenting CD103+ dendritic cells are protected from influenza virus infection. J. Clin. Invest. 122, 4037–4047 (2012).
Delamarre, L., Pack, M., Chang, H., Mellman, I. & Trombetta, E. S. Differential lysosomal proteolysis in antigen-presenting cells determines antigen fate. Science 307, 1630–1634 (2005).
Garris, C. S. et al. Successful anti-PD-1 cancer immunotherapy requires T cell-dendritic cell crosstalk involving the cytokines IFN-γ and IL-12. Immunity 49, 1148–1161 (2018).
Stitt, T. N. et al. The anticoagulation factor protein S and its relative, Gas6, are ligands for the Tyro 3/Axl family of receptor tyrosine kinases. Cell 80, 661–670 (1995).
Rothlin, C. V., Carrera-Silva, E. A., Bosurgi, L. & Ghosh, S. TAM receptor signaling in immune homeostasis. Annu. Rev. Immunol. 33, 355–391 (2015).
Skinner, H. D. et al. Integrative analysis identifies a novel AXL-PI3 kinase-PD-L1 signaling axis associated with radiation resistance in head and neck cancer. Clin. Cancer Res. 23, 2713–2722 (2017).
Tsukita, Y. et al. Axl kinase drives immune checkpoint and chemokine signalling pathways in lung adenocarcinomas. Mol. Cancer 18, 24 (2019).
Hugo, W. et al. Genomic and transcriptomic features of response to anti-PD-1 therapy in metastatic melanoma. Cell 165, 35–44 (2016).
Coussens, L. M., Zitvogel, L. & Palucka, A. K. Neutralizing tumor-promoting chronic inflammation: a magic bullet? Science 339, 286–291 (2013).
Mantovani, A. & Allavena, P. The interaction of anticancer therapies with tumor-associated macrophages. J. Exp. Med. 212, 435–445 (2015).
Zahner, S. P. et al. Conditional deletion of TGF-βR1 using Langerin-Cre mice results in Langerhans cell deficiency and reduced contact hypersensitivity. J. Immunol. 187, 5069–5076 (2011).
Xue, W. et al. Response and resistance to NF-κB inhibitors in mouse models of lung adenocarcinoma. Cancer Discov. 1, 236–247 (2011).
Stoeckius, M. et al. Simultaneous epitope and transcriptome measurement in single cells. Nat. Methods 14, 865–868 (2017).
Stoeckius, M. et al. Cell Hashing with barcoded antibodies enables multiplexing and doublet detection for single cell genomics. Genome Biol. 19, 224 (2018).
Jaitin, D. A. et al. Massively parallel single-cell RNA-seq for marker-free decomposition of tissues into cell types. Science 343, 776–779 (2014).
Paul, F. et al. Transcriptional heterogeneity and lineage commitment in myeloid progenitors. Cell 163, 1663–1677 (2015).
Baran, Y. et al. MetaCell: analysis of single cell RNA-seq data using K-nn graph partitions. Genome Biol. 20, 206 (2019).
Gubin, M. M. et al. High-dimensional analysis delineates myeloid and lymphoid compartment remodeling during successful immune-checkpoint cancer therapy. Cell 175, 1443 (2018).
Lambrechts, D. et al. Phenotype molding of stromal cells in the lung tumor microenvironment. Nat. Med. 24, 1277–1289 (2018).
Holland, S. J. et al. R428, a selective small molecule inhibitor of Axl kinase, blocks tumor spread and prolongs survival in models of metastatic breast cancer. Cancer Res. 70, 1544–1554 (2010).
Agudo, J. et al. GFP-specific CD8 T cells enable targeted cell depletion and visualization of T-cell interactions. Nat. Biotechnol. 33, 1287–1292 (2015).
Acknowledgements
This work was supported by National Institutes of Health (NIH) grants R01 CA154947, R01 s (to M.M.), 1R01CA212376 (to S.G. and C.V.R), F30CA243210 (to S.T.C.) and 5T32CA078207 (to A.M.L.). We thank C. Berin for helpful discussions; D. Farber and P. Dogra for critical comments on the manuscript; and the Mount Sinai flow cytometry core, Human Immune Monitoring Center and Mount Sinai Biorepository for support. Research support was provided by Regeneron and Takeda.
Author information
Authors and Affiliations
Contributions
M.M. conceived the project. B.M., B.D.B. and M.M. designed the experiments. B.M., A.M.L., B.D.B. and M.M. wrote the manuscript. A.M.L. and E.K. performed computational analysis. T.M. provided intellectual input and facilitated access to human samples. A.H.R. provided input to single-cell mapping strategies. B.M., S.T.C., N.T., C.C., A.C., S.M., J.L. and L.W. performed experiments. J.P.F. and N.B. provided B16-BFP/OVA cells. B.R. and M.E.K. provided OP4-DL1 cells. C.V.R. and S.G. provided Axl−/− and Axl−/− Mertk−/− bone marrow, and assisted with experiment design. A.W. and R.F. provided human tumour lesions. N.R.D. funded part of the study.
Corresponding author
Ethics declarations
Competing interests
Research support for these studies was provided by Regeneron and Takeda. The authors declare no other competing financial interests.
Additional information
Peer review information Nature thanks Cornels Melief and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 mregDCs are a distinct dendritic-cell cluster present in numerous tumour models.
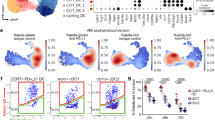
a, Digested lungs of naive or KP-tumour-bearing mice at day 28 post tumour-cell injection were stained with antibodies conjugated either to fluorophores for FACS or to oligonucleotides for CITE-seq analysis. CD45+ Siglec F− Ly6G− MHCII+ CD11c+ cells were sorted and loaded onto a 10x Chromium chip for scRNA-seq and CITE-seq analysis. Dendritic-cell clusters were identified according to marker-gene expression after clustering of transcriptomes. Heat maps show UMI counts of lineage genes across all clusters after downsampling to 2,000 UMIs per cell. b, Left, gene–gene correlation of highly variable genes, with relevant gene modules outlined and annotated; right, scRNA expression divided by cluster. Genes on left and right panels are aligned. c, CD45+ Siglec F− Ly6G− MHCII+ CD11c+ cells from lungs of naive or B16-BFP/OVA-tumour-bearing mice at day 22 were sorted and loaded onto a 10x Chromium platform for scRNA-seq. DCs were mapped to the clusters generated for the experiment shown in Fig. 1 by maximum-likelihood classification. Heat maps show UMI counts of lineage genes across all clusters after downsampling to 2,000 UMIs per cell. d, Mouse-tumour public scRNA data for immune cells from an M38 model13 and a T3 sarcoma model28 were accessed from GEO. Top, broad cell types were sorted in silico using gene lists, resulting in pure DC populations. pDC, plasmacytoid DC. Bottom left, DC1s, DC2s and mregDCs were identified using scores generated from gene lists that defined these populations. Bottom middle, annotations in the heat map were derived from k-means clustering (k = 3) of coordinates in the dendritic-cell-score scatter plot. Bottom right, DCs of each annotation are quantified. Gene lists defining cell types for in silico sorting and stratification of dendritic-cell subtypes are in Supplementary Table 2. e, Lung DC1s and migratory DC1s (migDC1) from DLNs were sorted and analysed by RNA-seq. Genes highlighted in red identify a reference set of genes from migratory DCs7. f, Lung DC1s and migratory DC1s from DLNs in both naive and KP-tumour-bearing mice were sorted and analysed by RNA-seq. The plot compares migDC1 gene expression with lung DC1 expression by log2FC in naive (x-axis) and KP-tumour-bearing (y-axis) mice. Genes upregulated in mregDCs relative to DC1s (log2FC greater than 2; Benjamini–Hochberg-adjusted P-value less than 0.01), as assayed by scRNA-seq, are shown in gold. g, Stratification of dendritic-cell transcriptomes using dendritic-cell subtype scores in naive and KP-tumour-bearing lungs. Scores for each subtype were generated from gene lists that were differentially expressed among clusters. Single cells are coloured by cluster identification (left) or CITE-seq surface marker expression (colour-bar units are log10(1 + ADT counts)). Gene scores are the same as in d (lower left).
Extended Data Fig. 2 The mregDC program is enriched in both canonical dendritic-cell subsets upon tumour-antigen uptake.
a, b, CD45+ Siglec F− Ly6G− MHCII+ CD11c+ cells were sorted from lungs of Ccr7−/− mice and loaded onto the 10x Chromium followed by scRNA-seq. Transcriptomes were mapped to the clusters generated for the wild-type experiment shown in Fig. 1 by maximum-likelihood classification. a, The heat map shows UMI counts of selected genes in dendritic-cell clusters after downsampling to 2,000 UMIs per cell, comparing cells from Ccr7−/− mice to cells from WT mice. b, Comparison of differential expression analyses between mregDCs and resting DCs in WT mice (x-axis) and Ccr7−/− mice (y-axis) (b). c, Frequencies of mregDCs as a percentage of total DCs, as measured by scRNA-seq in naive and KP–GFP-tumour-bearing mice. d, Gating strategy for subsets of conventional lung DCs. e, Flow cytometry of GFP+ versus GFP− DC2s (CD11b+ CD103−) from KP–GFP-tumour-bearing mice. f, Flow cytometry of GFP+ versus GFP− DC1s or DC2s from KP–GFP-tumour-bearing mice. g, Flow cytometry of BFP+ versus BFP− DC1s from B16-BFP/OVA tumour-bearing mice. The experiment shown is representative of two independent experiments; *P < 0.05; **P < 0.01, ***P < 0.001, ****P < 0.0001 (Student’s t-test); data are means ± s.d. (e–g). h, KP–GFP cells were exposed to ultraviolet radiation for 30 min, rested for 24 h, and stained with annexin V and propidium iodide in order to confirm induction of apoptosis before experiments involving coculture of DCs. i, Differential expression between mregDCs identified by transcriptome from KP-tumour-bearing and naive mice. Genes in green are significantly differentially expressed (Benjamini–Hochberg-adjusted P-value of less than 0.15); selected immune genes are shown in orange.
Extended Data Fig. 3 The mregDC program is independent of MyD88/TRIF, inflammasome signalling and lymphocytes.
a–d, Flow-cytometry analysis of DC1s isolated from KP–GFP-tumour-bearing lungs in Asc−/− (a), Il1r−/− (b), Myd88−/−Trif−/− (c) and Rag1−/− (d) mice. e, Heat map showing average TAM receptor RNA expression in mregDC, DC1 and DC2 scRNA-seq clusters. f, Differential expression of TH2 response genes across dendritic-cell clusters identified by scRNA-seq, showing a log2FC between average mregDC expression and resting dendritic-cell expression. Genes in green are differentially expressed (Benjamini–Hochberg-adjusted P-value less than 0.15); TH2 response genes are in orange. g, k, Mice were injected with KP–GFP tumour cells, treated with anti-IL-4 (αIL-4) or control IgG on days 21, 24 and 26, and analysed on day 28. GFP+ DC1s carrying tumour antigens in lung and DLNs (g) and T cells in DLNs (k) were analysed by flow cytometry. h, KP–GFP-tumour-bearing mice were injected with an agonistic CD40 antibody (CD40a) on days 25 and 27; lungs were analysed on day 28. i, KP–GFP-tumour-bearing mice were injected with polyI:C on day 27, and lungs were analysed on day 28. j, GFP+ conventional DC1s were purified from KP–GFP-tumour-bearing lungs from B6D2 mice treated either with anti-IL-4 or control IgG and cocultured with naive CD8+ JEDI T cells isolated from JEDI mouse spleens. JEDI T cells were analysed on day 2. One experiment, representative of two independent experiments, is shown (a–d, g–k). *P < 0.05; **P < 0.01; ***P < 0.001; ****P < 0.0001 (Student’s t-test (a–d, g, k) or one-way ANOVA and Tukey’s test (h, i)). Data are shown as means ± s.d. (a–d, g–i, k).
Extended Data Fig. 4 Protein expression profile of mregDCs in human NSCLC lesions.
a, Average CITE-seq surface protein staining intensity of dendritic-cell clusters in non-involved lung (nLung) and tumour lesions isolated from human NSCLC resections (n = 7). b, scRNA-seq data from a published dataset29 of matched nLung and tumour from resection specimens of eight patients with NSCLC were mapped to the clusters generated for the NSCLC data in Fig. 4 by maximum-likelihood classification. Heat maps show downsampled UMI counts in dendritic-cell clusters after downsampling cells to 2,000 UMIs per cell and evenly sampling cells from dendritic-cell types.
Supplementary information
Supplementary Table 1
Single-cell RNA-seq sample quality metrics.
Supplementary Table 2
Gene sets used in this study, including gene sets used to filter cells prior to clustering, gene sets used to filter cells from public datasets, and gene sets used for scoring dendritic cell subpopulations.
Rights and permissions
About this article
Cite this article
Maier, B., Leader, A.M., Chen, S.T. et al. A conserved dendritic-cell regulatory program limits antitumour immunity. Nature 580, 257–262 (2020). https://doi.org/10.1038/s41586-020-2134-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-2134-y
This article is cited by
-
The dynamic shifts of IL-10-producing Th17 and IL-17-producing Treg in health and disease: a crosstalk between ancient "Yin-Yang" theory and modern immunology
Cell Communication and Signaling (2024)
-
Immunosuppressive tumor microenvironment and immunotherapy of hepatocellular carcinoma: current status and prospectives
Journal of Hematology & Oncology (2024)
-
Acidovorax temperans skews neutrophil maturation and polarizes Th17 cells to promote lung adenocarcinoma development
Oncogenesis (2024)
-
A new step in understanding mouse cDC ontogeny
Nature Immunology (2024)
-
Coordinated chemokine expression defines macrophage subsets across tissues
Nature Immunology (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.