Abstract
Cancer is driven by genetic change, and the advent of massively parallel sequencing has enabled systematic documentation of this variation at the whole-genome scale1,2,3. Here we report the integrative analysis of 2,658 whole-cancer genomes and their matching normal tissues across 38 tumour types from the Pan-Cancer Analysis of Whole Genomes (PCAWG) Consortium of the International Cancer Genome Consortium (ICGC) and The Cancer Genome Atlas (TCGA). We describe the generation of the PCAWG resource, facilitated by international data sharing using compute clouds. On average, cancer genomes contained 4–5 driver mutations when combining coding and non-coding genomic elements; however, in around 5% of cases no drivers were identified, suggesting that cancer driver discovery is not yet complete. Chromothripsis, in which many clustered structural variants arise in a single catastrophic event, is frequently an early event in tumour evolution; in acral melanoma, for example, these events precede most somatic point mutations and affect several cancer-associated genes simultaneously. Cancers with abnormal telomere maintenance often originate from tissues with low replicative activity and show several mechanisms of preventing telomere attrition to critical levels. Common and rare germline variants affect patterns of somatic mutation, including point mutations, structural variants and somatic retrotransposition. A collection of papers from the PCAWG Consortium describes non-coding mutations that drive cancer beyond those in the TERT promoter4; identifies new signatures of mutational processes that cause base substitutions, small insertions and deletions and structural variation5,6; analyses timings and patterns of tumour evolution7; describes the diverse transcriptional consequences of somatic mutation on splicing, expression levels, fusion genes and promoter activity8,9; and evaluates a range of more-specialized features of cancer genomes8,10,11,12,13,14,15,16,17,18.
Similar content being viewed by others
Main
Cancer is the second most-frequent cause of death worldwide, killing more than 8 million people every year; the incidence of cancer is expected to increase by more than 50% over the coming decades19,20. ‘Cancer’ is a catch-all term used to denote a set of diseases characterized by autonomous expansion and spread of a somatic clone. To achieve this behaviour, the cancer clone must co-opt multiple cellular pathways that enable it to disregard the normal constraints on cell growth, modify the local microenvironment to favour its own proliferation, invade through tissue barriers, spread to other organs and evade immune surveillance21. No single cellular program directs these behaviours. Rather, there is a large pool of potential pathogenic abnormalities from which individual cancers draw their own combinations: the commonalities of macroscopic features across tumours belie a vastly heterogeneous landscape of cellular abnormalities.
This heterogeneity arises from the stochastic nature of Darwinian evolution. There are three preconditions for Darwinian evolution: characteristics must vary within a population; this variation must be heritable from parent to offspring; and there must be competition for survival within the population. In the context of somatic cells, heritable variation arises from mutations acquired stochastically throughout life, notwithstanding additional contributions from germline and epigenetic variation. A subset of these mutations alter the cellular phenotype, and a small subset of those variants confer an advantage on clones during the competition to escape the tight physiological controls wired into somatic cells. Mutations that provide a selective advantage to the clone are termed driver mutations, as opposed to selectively neutral passenger mutations.
Initial studies using massively parallel sequencing demonstrated the feasibility of identifying every somatic point mutation, copy-number change and structural variant (SV) in a given cancer1,2,3. In 2008, recognizing the opportunity that this advance in technology provided, the global cancer genomics community established the ICGC with the goal of systematically documenting the somatic mutations that drive common tumour types22.
The pan-cancer analysis of whole genomes
The expansion of whole-genome sequencing studies from individual ICGC and TCGA working groups presented the opportunity to undertake a meta-analysis of genomic features across tumour types. To achieve this, the PCAWG Consortium was established. A Technical Working Group implemented the informatics analyses by aggregating the raw sequencing data from different working groups that studied individual tumour types, aligning the sequences to the human genome and delivering a set of high-quality somatic mutation calls for downstream analysis (Extended Data Fig. 1). Given the recent meta-analysis of exome data from the TCGA Pan-Cancer Atlas23,24,25, scientific working groups concentrated their efforts on analyses best-informed by whole-genome sequencing data.
We collected genome data from 2,834 donors (Extended Data Table 1), of which 176 were excluded after quality assurance. A further 75 had minor issues that could affect some of the analyses (grey-listed donors) and 2,583 had data of optimal quality (white-listed donors) (Supplementary Table 1). Across the 2,658 white- and grey-listed donors, whole-genome sequencing data were available from 2,605 primary tumours and 173 metastases or local recurrences. Mean read coverage was 39× for normal samples, whereas tumours had a bimodal coverage distribution with modes at 38× and 60× (Supplementary Fig. 1). RNA-sequencing data were available for 1,222 donors. The final cohort comprised 1,469 men (55%) and 1,189 women (45%), with a mean age of 56 years (range, 1–90 years) across 38 tumour types (Extended Data Table 1 and Supplementary Table 1).
To identify somatic mutations, we analysed all 6,835 samples using a uniform set of algorithms for alignment, variant calling and quality control (Extended Data Fig. 1, Supplementary Fig. 2 and Supplementary Methods 2). We used three established pipelines to call somatic single-nucleotide variations (SNVs), small insertions and deletions (indels), copy-number alterations (CNAs) and SVs. Somatic retrotransposition events, mitochondrial DNA mutations and telomere lengths were also called by bespoke algorithms. RNA-sequencing data were uniformly processed to call transcriptomic alterations. Germline variants identified by the three separate pipelines included single-nucleotide polymorphisms, indels, SVs and mobile-element insertions (Supplementary Table 2).
The requirement to uniformly realign and call variants on approximately 5,800 whole genomes presented considerable computational challenges, and raised ethical issues owing to the use of data from different jurisdictions (Extended Data Table 2). We used cloud computing26,27 to distribute alignment and variant calling across 13 data centres on 3 continents (Supplementary Table 3). Core pipelines were packaged into Docker containers28 as reproducible, stand-alone packages, which we have made available for download. Data repositories for raw and derived datasets, together with portals for data visualization and exploration, have also been created (Box 1 and Supplementary Table 4).
Benchmarking of genetic variant calls
To benchmark mutation calling, we ran the 3 core pipelines, together with 10 additional pipelines, on 63 representative tumour–normal genome pairs (Supplementary Note 1). For 50 of these cases, we performed validation by hybridization of tumour and matched normal DNA to a custom bait set with deep sequencing29. The 3 core somatic variant-calling pipelines had individual estimates of sensitivity of 80–90% to detect a true somatic SNV called by any of the 13 pipelines; more than 95% of SNV calls made by each of the core pipelines were genuine somatic variants (Fig. 1a). For indels—a more-challenging class of variants to identify with short-read sequencing—the 3 core algorithms had individual sensitivity estimates in the range of 40–50%, with precision of 70–95% (Fig. 1b). For individual SV algorithms, we estimated precision to be in the range 80–95% for samples in the 63-sample pilot dataset.
a, Scatter plot of estimated sensitivity and precision for somatic SNVs across individual algorithms assessed in the validation exercise across n = 63 PCAWG samples. Core algorithms included in the final PCAWG call set are shown in blue. b, Sensitivity and precision estimates across individual algorithms for somatic indels. c, Accuracy (precision, sensitivity and F1 score, defined as 2 × sensitivity × precision/(sensitivity + precision)) of somatic SNV calls across variant allele fractions (VAFs) for the core algorithms. The accuracy of two methods of combining variant calls (two-plus, which was used in the final dataset, and logistic regression) is also shown. d, Accuracy of indel calls across variant allele fractions.
Next, we defined a strategy to merge results from the three pipelines into one final call-set to be used for downstream scientific analyses (Methods and Supplementary Note 2). Sensitivity and precision of consensus somatic variant calls were 95% (90% confidence interval, 88–98%) and 95% (90% confidence interval, 71–99%), respectively, for SNVs (Extended Data Fig. 2). For somatic indels, sensitivity and precision were 60% (34–72%) and 91% (73–96%), respectively (Extended Data Fig. 2). Regarding somatic SVs, we estimate the sensitivity of merged calls to be 90% for true calls generated by any one pipeline; precision was estimated as 97.5%. The improvement in calling accuracy from combining different pipelines was most noticeable in variants with low variant allele fractions, which probably originate from tumour subclones (Fig. 1c, d). Germline variant calls, phased using a haplotype-reference panel, displayed a precision of more than 99% and a sensitivity of 92–98% (Supplementary Note 2).
Analysis of PCAWG data
The uniformly generated, high-quality set of variant calls across more than 2,500 donors provided the springboard for a series of scientific working groups to explore the biology of cancer. A comprehensive suite of companion papers that describe the analyses and discoveries across these thematic areas is copublished with this paper4,5,6,7,8,9,10,11,12,13,14,15,16,17,18 (Extended Data Table 3).
Pan-cancer burden of somatic mutations
Across the 2,583 white-listed PCAWG donors, we called 43,778,859 somatic SNVs, 410,123 somatic multinucleotide variants, 2,418,247 somatic indels, 288,416 somatic SVs, 19,166 somatic retrotransposition events and 8,185 de novo mitochondrial DNA mutations (Supplementary Table 1). There was considerable heterogeneity in the burden of somatic mutations across patients and tumour types, with a broad correlation in mutation burden among different classes of somatic variation (Extended Data Fig. 3). Analysed at a per-patient level, this correlation held, even when considering tumours with similar purity and ploidy (Supplementary Fig. 3). Why such correlation should apply on a pan-cancer basis is unclear. It is likely that age has some role, as we observe a correlation between most classes of somatic mutation and age at diagnosis (around 190 SNVs per year, P = 0.02; about 22 indels per year, P = 5 × 10−5; 1.5 SVs per year, P < 2 × 10−16; linear regression with likelihood ratio tests; Supplementary Fig. 4). Other factors are also likely to contribute to the correlations among classes of somatic mutation, as there is evidence that some DNA-repair defects can cause multiple types of somatic mutation30, and a single carcinogen can cause a range of DNA lesions31.
Panorama of driver mutations in cancer
We extracted the subset of somatic mutations in PCAWG tumours that have high confidence to be driver events on the basis of current knowledge. One challenge to pinpointing the specific driver mutations in an individual tumour is that not all point mutations in recurrently mutated cancer-associated genes are drivers32. For genomic elements significantly mutated in PCAWG data, we developed a ‘rank-and-cut’ approach to identify the probable drivers (Supplementary Methods 8.1). This approach works by ranking the observed mutations in a given genomic element based on recurrence, estimated functional consequence and expected pattern of drivers in that element. We then estimate the excess burden of somatic mutations in that genomic element above that expected for the background mutation rate, and cut the ranked mutations at this level. Mutations in each element with the highest driver ranking were then assigned as probable drivers; those below the threshold will probably have arisen through chance and were assigned as probable passengers. Improvements to features that are used to rank the mutations and the methods used to measure them will contribute to further development of the rank-and-cut approach.
We also needed to account for the fact that some bona fide cancer genomic elements were not rediscovered in PCAWG data because of low statistical power. We therefore added previously known cancer-associated genes to the discovery set, creating a ‘compendium of mutational driver elements’ (Supplementary Methods 8.2). Then, using stringent rules to nominate driver point mutations that affect these genomic elements on the basis of prior knowledge33, we separated probable driver from passenger point mutations. To cover all classes of variant, we also created a compendium of known driver SVs, using analogous rules to identify which somatic CNAs and SVs are most likely to act as drivers in each tumour. For probable pathogenic germline variants, we identified all truncating germline point mutations and SVs that affect high-penetrance germline cancer-associated genes.
This analysis defined a set of mutations that we could confidently assert, based on current knowledge, drove tumorigenesis in the more than 2,500 tumours of PCAWG. We found that 91% of tumours had at least one identified driver mutation, with an average of 4.6 drivers per tumour identified, showing extensive variation across cancer types (Fig. 2a). For coding point mutations, the average was 2.6 drivers per tumour, similar to numbers estimated in known cancer-associated genes in tumours in the TCGA using analogous approaches32.
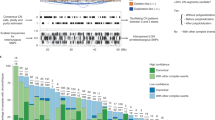
a, Top, putative driver mutations in PCAWG, represented as a circos plot. Each sector represents a tumour in the cohort. From the periphery to the centre of the plot the concentric rings represent: (1) the total number of driver alterations; (2) the presence of whole-genome (WG) duplication; (3) the tumour type; (4) the number of driver CNAs; (5) the number of driver genomic rearrangements; (6) driver coding point mutations; (7) driver non-coding point mutations; and (8) pathogenic germline variants. Bottom, snapshots of the panorama of driver mutations. The horizontal bar plot (left) represents the proportion of patients with different types of drivers. The dot plot (right) represents the mean number of each type of driver mutation across tumours with at least one event (the square dot) and the standard deviation (grey whiskers), based on n = 2,583 patients. b, Genomic elements targeted by different types of mutations in the cohort altered in more than 65 tumours. Both germline and somatic variants are included. Left, the heat map shows the recurrence of alterations across cancer types. The colour indicates the proportion of mutated tumours and the number indicates the absolute count of mutated tumours. Right, the proportion of each type of alteration that affects each genomic element. c, Tumour-suppressor genes with biallelic inactivation in 10 or more patients. The values included under the gene labels represent the proportions of patients who have biallelic mutations in the gene out of all patients with a somatic mutation in that gene. GR, genomic rearrangement; SCNA, somatic copy-number alteration; SGR, somatic genome rearrangement; TSG, tumour suppressor gene; UTR, untranslated region.
To address the frequency of non-coding driver point mutations, we combined promoters and enhancers that are known targets of non-coding drivers34,35,36,37 with those newly discovered in PCAWG data; this is reported in a companion paper4. Using this approach, only 13% (785 out of 5,913) of driver point mutations were non-coding in PCAWG. Nonetheless, 25% of PCAWG tumours bear at least one putative non-coding driver point mutation, and one third (237 out of 785) affected the TERT promoter (9% of PCAWG tumours). Overall, non-coding driver point mutations are less frequent than coding driver mutations. With the exception of the TERT promoter, individual enhancers and promoters are only infrequent targets of driver mutations4.
Across tumour types, SVs and point mutations have different relative contributions to tumorigenesis. Driver SVs are more prevalent in breast adenocarcinomas (6.4 ± 3.7 SVs (mean ± s.d.) compared with 2.2 ± 1.3 point mutations; P < 1 × 10−16, Mann–Whitney U-test) and ovary adenocarcinomas (5.8 ± 2.6 SVs compared with 1.9 ± 1.0 point mutations; P < 1 × 10−16), whereas driver point mutations have a larger contribution in colorectal adenocarcinomas (2.4 ± 1.4 SVs compared with 7.4 ± 7.0 point mutations; P = 4 × 10−10) and mature B cell lymphomas (2.2 ± 1.3 SVs compared with 6 ± 3.8 point mutations; P < 1 × 10−16), as previously shown38. Across tumour types, there are differences in which classes of mutation affect a given genomic element (Fig. 2b).
We confirmed that many driver mutations that affect tumour-suppressor genes are two-hit inactivation events (Fig. 2c). For example, of the 954 tumours in the cohort with driver mutations in TP53, 736 (77%) had both alleles mutated, 96% of which (707 out of 736) combined a somatic point mutation that affected one allele with somatic deletion of the other allele. Overall, 17% of patients had rare germline protein-truncating variants (PTVs) in cancer-predisposition genes39, DNA-damage response genes40 and somatic driver genes. Biallelic inactivation due to somatic alteration on top of a germline PTV was observed in 4.5% of patients overall, with 81% of these affecting known cancer-predisposition genes (such as BRCA1, BRCA2 and ATM).
PCAWG tumours with no apparent drivers
Although more than 90% of PCAWG cases had identified drivers, we found none in 181 tumours (Extended Data Fig. 4a). Reasons for missing drivers have not yet been systematically evaluated in a pan-cancer cohort, and could arise from either technical or biological causes.
Technical explanations could include poor-quality samples, inadequate sequencing or failures in the bioinformatic algorithms used. We assessed the quality of the samples and found that 4 of the 181 cases with no known drivers had more than 5% tumour DNA contamination in their matched normal sample (Fig. 3a). Using an algorithm designed to correct for this contamination41, we identified previously missed mutations in genes relevant to the respective cancer types. Similarly, if the fraction of tumour cells in the cancer sample is low through stromal contamination, the detection of driver mutations can be impaired. Most tumours with no known drivers had an average power to detect mutations close to 100%; however, a few had power in the 70–90% range (Fig. 3b and Extended Data Fig. 4b). Even in adequately sequenced genomes, lack of read depth at specific driver loci can impair mutation detection. For example, only around 50% of PCAWG tumours had sufficient coverage to call a mutation (≥90% power) at the two TERT promoter hotspots, probably because the high GC content of this region causes biased coverage (Fig. 3c). In fact, 6 hepatocellular carcinomas and 2 biliary cholangiocarcinomas among the 181 cases with no known drivers actually did contain TERT mutations, which were discovered after deep targeted sequencing42.
a, Individual estimates of the percentage of tumour-in-normal contamination across patients with no driver mutations in PCAWG (n = 181). No data were available for myelodysplastic syndromes and acute myeloid leukaemia. Points represent estimates for individual patients, and the coloured areas are estimated density distributions (violin plots). Abbreviations of the tumour types are defined in Extended Data Table 1. b, Average detection sensitivity by tumour type for tumours without known drivers (n = 181). Each dot represents a given sample and is the average sensitivity of detecting clonal substitutions across the genome, taking into account purity and ploidy. Coloured areas are estimated density distributions, shown for cohorts with at least five cases. c, Detection sensitivity for TERT promoter hotspots in tumour types in which TERT is frequently mutated. Coloured areas are estimated density distributions. d, Significant copy-number losses identified by two-sided hypothesis testing using GISTIC2.0, corrected for multiple-hypothesis testing. Numbers in parentheses indicate the number of genes in significant regions when analysing medulloblastomas without known drivers (n = 42). Significant regions with known cancer-associated genes are labelled with the representative cancer-associated gene. e, Aneuploidy in chromophobe renal cell carcinomas and pancreatic neuroendocrine tumours without known drivers. Patients are ordered on the y axis by tumour type and then by presence of whole-genome duplication (bottom) or not (top).
Finally, technical reasons for missing driver mutations include failures in the bioinformatic algorithms. This affected 35 myeloproliferative neoplasms in PCAWG, in which the JAK2V617F driver mutation should have been called. Our somatic variant-calling algorithms rely on ‘panels of normals’, typically from blood samples, to remove recurrent sequencing artefacts. As 2–5% of healthy individuals carry occult haematopoietic clones43, recurrent driver mutations in these clones can enter panels of normals.
With regard to biological causes, tumours may be driven by mutations in cancer-associated genes that are not yet described for that tumour type. Using driver discovery algorithms on tumours with no known drivers, no individual genes reached significance for point mutations. However, we identified a recurrent CNA that spanned SETD2 in medulloblastomas that lacked known drivers (Fig. 3d), indicating that restricting hypothesis testing to missing-driver cases can improve power if undiscovered genes are enriched in such tumours. Inactivation of SETD2 in medulloblastoma significantly decreased gene expression (P = 0.002) (Extended Data Fig. 4c). Notably, SETD2 mutations occurred exclusively in medulloblastoma group-4 tumours (P < 1 × 10−4). Group-4 medulloblastomas are known for frequent mutations in other chromatin-modifying genes44, and our results suggest that SETD2 loss of function is an additional driver that affects chromatin regulators in this subgroup.
Two tumour types had a surprisingly high fraction of patients without identified driver mutations: chromophobe renal cell carcinoma (44%; 19 out of 43) and pancreatic neuroendocrine cancers (22%; 18 out of 81) (Extended Data Fig. 4a). A notable feature of the missing-driver cases in both tumour types was a remarkably consistent profile of chromosomal aneuploidy—patterns that have previously been reported45,46 (Fig. 3e). The absence of other identified driver mutations in these patients raises the possibility that certain combinations of whole-chromosome gains and losses may be sufficient to initiate a cancer in the absence of more-targeted driver events such as point mutations or fusion genes of focal CNAs.
Even after accounting for technical issues and novel drivers, 5.3% of PCAWG tumours still had no identifiable driver events. In a research setting, in which we are interested in drawing conclusions about populations of patients, the consequences of technical issues that affect occasional samples will be mitigated by sample size. In a clinical setting, in which we are interested in the driver mutations in a specific patient, these issues become substantially more important. Careful and critical appraisal of the whole pipeline—including sample acquisition, genome sequencing, mapping, variant calling and driver annotation, as done here—should be required for laboratories that offer clinical sequencing of cancer genomes.
Patterns of clustered mutations and SVs
Some somatic mutational processes generate multiple mutations in a single catastrophic event, typically clustered in genomic space, leading to substantial reconfiguration of the genome. Three such processes have previously been described: (1) chromoplexy, in which repair of co-occurring double-stranded DNA breaks—typically on different chromosomes—results in shuffled chains of rearrangements47,48 (Extended Data Fig. 5a); (2) kataegis, a focal hypermutation process that leads to locally clustered nucleotide substitutions, biased towards a single DNA strand49,50,51 (Extended Data Fig. 5b); and (3) chromothripsis, in which tens to hundreds of DNA breaks occur simultaneously, clustered on one or a few chromosomes, with near-random stitching together of the resulting fragments52,53,54,55 (Extended Data Fig. 5c). We characterized the PCAWG genomes for these three processes (Fig. 4).
a, Kataegis. Top, prevalence of different types of kataegis and their association with SVs (≤1 kb from the focus). Bottom, the distribution of the number of foci of kataegis per sample. Chromoplexy. Prevalence of chromoplexy across cancer types, subdivided into balanced translocations and more complex events. Chromothripsis. Top, frequency of chromothripsis across cancer types. Bottom, for each cancer type a column is shown, in which each row is a chromothripsis region represented by five coloured rectangles relating to its categorization. b, Circos rainfall plot showing the distances between consecutive kataegis events across PCAWG compared with their genomic position. Lymphoid tumours (khaki, B cell non-Hodgkin’s lymphoma; orange, chronic lymphocytic leukaemia) have hypermutation hot spots (≥3 foci with distance ≤1 kb; pale red zone), many of which are near known cancer-associated genes (red annotations) and have associated SVs (≤10 kb from the focus; shown as arcs in the centre). c, Circos rainfall plot as in b that shows the distance versus the position of consecutive chromoplexy and reciprocal translocation footprints across PCAWG. Lymphoid, prostate and thyroid cancers exhibit recurrent events (≥2 footprints with distance ≤10 kb; pale red zone) that are likely to be driver SVs and are annotated with nearby genes and associated SVs, which are shown as bold and thin arcs for chromoplexy and reciprocal translocations, respectively (colours as in a). d, Effect of chromothripsis along the genome and involvement of PCAWG driver genes. Top, number of chromothripsis-induced gains or losses (grey) and amplifications (blue) or deletions (red). Within the identified chromothripsis regions, selected recurrently rearranged (light grey), amplified (blue) and homozygously deleted (magenta) driver genes are indicated. Bottom, interbreakpoint distance between all subsequent breakpoints within chromothripsis regions across cancer types, coloured by cancer type. Regions with an average interbreakpoint distance <10 kb are highlighted. C[T>N]T, kataegis with a pattern of thymine mutations in a Cp TpT context.
Chromoplexy events and reciprocal translocations were identified in 467 (17.8%) samples (Fig. 4a, c). Chromoplexy was prominent in prostate adenocarcinoma and lymphoid malignancies, as previously described47,48, and—unexpectedly—thyroid adenocarcinoma. Different genomic loci were recurrently rearranged by chromoplexy across the three tumour types, mediated by positive selection for particular fusion genes or enhancer-hijacking events. Of 13 fusion genes or enhancer hijacking events in 48 thyroid adenocarcinomas, at least 4 (31%) were caused by chromoplexy, with a further 4 (31%) part of complexes that contained chromoplexy footprints (Extended Data Fig. 5a). These events generated fusion genes that involved RET (two cases) and NTRK3 (one case)56, and the juxtaposition of the oncogene IGF2BP3 with regulatory elements from highly expressed genes (five cases).
Kataegis events were found in 60.5% of all cancers, with particularly high abundance in lung squamous cell carcinoma, bladder cancer, acral melanoma and sarcomas (Fig. 4a, b). Typically, kataegis comprises C > N mutations in a TpC context, which are probably caused by APOBEC activity49,50,51, although a T > N conversion in a TpT or CpT process (the affected T is highlighted in bold) attributed to error-prone polymerases has recently been described57. The APOBEC signature accounted for 81.7% of kataegis events and correlated positively with APOBEC3B expression levels, somatic SV burden and age at diagnosis (Supplementary Fig. 5). Furthermore, 5.7% of kataegis events involved the T > N error-prone polymerase signature and 2.3% of events, most notably in sarcomas, showed cytidine deamination in an alternative GpC or CpC context.
Kataegis events were frequently associated with somatic SV breakpoints (Fig. 4a and Supplementary Fig. 6a), as previously described50,51. Deletions and complex rearrangements were most-strongly associated with kataegis, whereas tandem duplications and other simple SV classes were only infrequently associated (Supplementary Fig. 6b). Kataegis inducing predominantly T > N mutations in CpTpT context was enriched near deletions, specifically those in the 10–25-kilobase (kb) range (Supplementary Fig. 6c).
Samples with extreme kataegis burden (more than 30 foci) comprise four types of focal hypermutation (Extended Data Fig. 6): (1) off-target somatic hypermutation and foci of T > N at CpTpT, found in B cell non-Hodgkin lymphoma and oesophageal adenocarcinomas, respectively; (2) APOBEC kataegis associated with complex rearrangements, notably found in sarcoma and melanoma; (3) rearrangement-independent APOBEC kataegis on the lagging strand and in early-replicating regions, mainly found in bladder and head and neck cancer; and (4) a mix of the last two types. Kataegis only occasionally led to driver mutations (Supplementary Table 5).
We identified chromothripsis in 587 samples (22.3%), most frequently among sarcoma, glioblastoma, lung squamous cell carcinoma, melanoma and breast adenocarcinoma18. Chromothripsis increased with whole-genome duplications in most cancer types (Extended Data Fig. 7a), as previously shown in medulloblastoma58. The most recurrently associated driver was TP5352 (pan-cancer odds ratio = 3.22; pan-cancer P = 8.3 × 10−35; q < 0.05 in breast lobular (odds ratio = 13), colorectal (odds ratio = 25), prostate (odds ratio = 2.6) and hepatocellular (odds ratio = 3.9) cancers; Fisher–Boschloo tests). In two cancer types (osteosarcoma and B cell lymphoma), women had a higher incidence of chromothripsis than men (Extended Data Fig. 7b). In prostate cancer, we observed a higher incidence of chromothripsis in patients with late-onset than early-onset disease59 (Extended Data Fig. 7c).
Chromothripsis regions coincided with 3.6% of all identified drivers in PCAWG and around 7% of copy-number drivers (Fig. 4d). These proportions are considerably enriched compared to expectation if selection were not acting on these events (Extended Data Fig. 7d). The majority of coinciding driver events were amplifications (58%), followed by homozygous deletions (34%) and SVs within genes or promoter regions (8%). We frequently observed a ≥2-fold increase or decrease in expression of amplified or deleted drivers, respectively, when these loci were part of a chromothripsis event, compared with samples without chromothripsis (Extended Data Fig. 7e).
Chromothripsis manifested in diverse patterns and frequencies across tumour types, which we categorized on the basis of five characteristics (Fig. 4a). In liposarcoma, for example, chromothripsis events often involved multiple chromosomes, with universal MDM2 amplification60 and co-amplification of TERT in 4 of 19 cases (Fig. 4d). By contrast, in glioblastoma the events tended to affect a smaller region on a single chromosome that was distant from the telomere, resulting in focal amplification of EGFR and MDM2 and loss of CDKN2A. Acral melanomas frequently exhibited CCND1 amplification, and lung squamous cell carcinomas SOX2 amplifications. In both cases, these drivers were more-frequently altered by chromothripsis compared with other drivers in the same cancer type and to other cancer types for the same driver (Fig. 4d and Extended Data Fig. 7f). Finally, in chromophobe renal cell carcinoma, chromothripsis nearly always affected chromosome 5 (Supplementary Fig. 7): these samples had breakpoints immediately adjacent to TERT, increasing TERT expression by 80-fold on average compared with samples without rearrangements (P = 0.0004; Mann–Whitney U-test).
Timing clustered mutations in evolution
An unanswered question for clustered mutational processes is whether they occur early or late in cancer evolution. To address this, we used molecular clocks to define broad epochs in the life history of each tumour49,61. One transition point is between clonal and subclonal mutations: clonal mutations occurred before, and subclonal mutations after, the emergence of the most-recent common ancestor. In regions with copy-number gains, molecular time can be further divided according to whether mutations preceded the copy-number gain (and were themselves duplicated) or occurred after the gain (and therefore present on only one chromosomal copy)7.
Chromothripsis tended to have greater relative odds of being clonal than subclonal, suggesting that it occurs early in cancer evolution, especially in liposarcomas, prostate adenocarcinoma and squamous cell lung cancer (Fig. 5a). As previously reported, chromothripsis was especially common in melanomas62. We identified 89 separate chromothripsis events that affected 66 melanomas (61%); 47 out of 89 events affected genes known to be recurrently altered in melanoma63 (Supplementary Table 6). Involvement of a region on chromosome 11 that includes the cell-cycle regulator CCND1 occurred in 21 cases (10 out of 86 cutaneous, and 11 out of 21 acral or mucosal melanomas), typically combining chromothripsis with amplification (19 out of 21 cases) (Extended Data Fig. 8). Co-involvement of other cancer-associated genes in the same chromothripsis event was also frequent, including TERT (five cases), CDKN2A (three cases), TP53 (two cases) and MYC (two cases) (Fig. 5b). In these co-amplifications, a chromothripsis event involving multiple chromosomes initiated the process, creating a derivative chromosome in which hundreds of fragments were stitched together in a near-random order (Fig. 5b). This derivative then rearranged further, leading to massive co-amplification of the multiple target oncogenes together with regions located nearby on the derivative chromosome.
a, Extent and timing of chromothripsis, kataegis and chromoplexy across PCAWG. Top, stacked bar charts illustrate co-occurrence of chromothripsis, kataegis and chromoplexy in the samples. Middle, relative odds of clustered events being clonal or subclonal are shown with bootstrapped 95% confidence intervals. Point estimates are highlighted when they do not overlap odds of 1:1. Bottom, relative odds of the events being early or late clonal are shown as above. Sample sizes (number of patients) are shown across the top. b, Three representative patients with acral melanoma and chromothripsis-induced amplification that simultaneously affects TERT and CCND1. The black points (top) represent sequence coverage from individual genomic bins, with SVs shown as coloured arcs (translocation in black, deletion in purple, duplication in brown, tail-to-tail inversion in cyan and head-to-head inversion in green). Bottom, the variant allele fractions of somatic point mutations.
In these cases of amplified chromothripsis, we can use the inferred number of copies bearing each SNV to time the amplification process. SNVs present on the chromosome before amplification will themselves be amplified and are therefore reported in a high fraction of sequence reads (Fig. 5b and Extended Data Fig. 8). By contrast, late SNVs that occur after the amplification has concluded will be present on only one chromosome copy out of many, and thus have a low variant allele fraction. Regions of CCND1 amplification had few—sometimes zero—mutations at high variant allele fraction in acral melanomas, in contrast to later CCND1 amplifications in cutaneous melanomas, in which hundreds to thousands of mutations typically predated amplification (Fig. 5b and Extended Data Fig. 9a, b). Thus, both chromothripsis and the subsequent amplification generally occurred very early during the evolution of acral melanoma. By comparison, in lung squamous cell carcinomas, similar patterns of chromothripsis followed by SOX2 amplification are characterized by many amplified SNVs, suggesting a later event in the evolution of these cancers (Extended Data Fig. 9c).
Notably, in cancer types in which the mutational load was sufficiently high, we could detect a larger-than-expected number of SNVs on an intermediate number of DNA copies, suggesting that they appeared during the amplification process (Supplementary Fig. 8).
Germline effects on somatic mutations
We integrated the set of 88 million germline genetic variant calls with somatic mutations in PCAWG, to study germline determinants of somatic mutation rates and patterns. First, we performed a genome-wide association study of somatic mutational processes with common germline variants (minor allele frequency (MAF) > 5%) in individuals with inferred European ancestry. An independent genome-wide association study was performed in East Asian individuals from Asian cancer genome projects. We focused on two prevalent endogenous mutational processes: spontaneous deamination of 5-methylcytosine at CpG dinucleotides5 (signature 1) and activity of the APOBEC3 family of cytidine deaminases64 (signatures 2 and 13). No locus reached genome-wide significance (P < 5 × 10−8) for signature 1 (Extended Data Fig. 10a, b). However, a locus at 22q13.1 predicted an APOBEC3B-like mutagenesis at the pan-cancer level65 (Fig. 6a). The strongest signal at 22q13.1 was driven by rs12628403, and the minor (non-reference) allele was protective against APOBEC3B-like mutagenesis (β = −0.43, P = 5.6 × 10−9, MAF = 8.2%, n = 1,201 donors) (Extended Data Fig. 10c). This variant tags a common, approximately 30-kb germline SV that deletes the APOBEC3B coding sequence and fuses the APOBEC3B 3′ untranslated region with the coding sequence of APOBEC3A. The deletion is known to increase breast cancer risk and APOBEC mutagenesis in breast cancer genomes66,67. Here, we found that rs12628403 reduces APOBEC3B-like mutagenesis specifically in cancer types with low levels of APOBEC mutagenesis (βlow = −0.50, Plow = 1 × 10−8; βhigh = 0.17, Phigh = 0.2), and increases APOBEC3A-like mutagenesis in cancer types with high levels of APOBEC mutagenesis (βhigh = 0.44, Phigh = 8 × 10−4; βlow = −0.21, Plow = 0.02). Moreover, we identified a second, novel locus at 22q13.1 that was associated with APOBEC3B-like mutagenesis across cancer types (rs2142833, β = 0.23, P = 1.3 × 10−8). We independently validated the association between both loci and APOBEC3B-like mutagenesis using East Asian individuals from Asian cancer genome projects (βrs12628403 = 0.57, Prs12628403 = 4.2 × 10−12; βrs2142833 = 0.58, Prs2142833 = 8 × 10−15) (Extended Data Fig. 10d). Notably, in a conditional analysis that accounted for rs12628403, we found that rs2142833 and rs12628403 are inherited independently in Europeans (r2<0.1), and rs2142833 remained significantly associated with APOBEC3B-like mutagenesis in Europeans (βEUR = 0.17, PEUR = 3 × 10−5) and East Asians (βASN = 0.25, PASN = 2 × 10−3) (Extended Data Fig. 10e, f). Analysis of donor-matched expression data further suggests that rs2142833 is a cis-expression quantitative trait locus (eQTL) for APOBEC3B at the pan-cancer level (β = 0.19, P = 2 × 10−6) (Extended Data Fig. 10g, h), consistent with cis-eQTL studies in normal cells68,69.
a, Association between common (MAF > 5%) germline variants and somatic APOBEC3B-like mutagenesis in individuals of European ancestry (n = 1,201). Two-sided hypothesis testing was performed with PLINK v.1.9. To mitigate multiple-hypothesis testing, the significance threshold was set to genome-wide significance (P < 5 × 10−8). b, Templated insertion SVs in a BRCA1-associated prostate cancer. Left, chromosome bands (1); SVs ≤ 10 megabases (Mb) (2); 1-kb read depth corrected to copy number 0–6 (3); inter- and intrachromosomal SVs > 10 Mb (4). Right, a complex somatic SV composed of a 2.2-kb tandem duplication on chromosome 2 together with a 232-base-pair (bp) inverted templated insertion SV that is derived from chromosome 5 and inserted inbetween the tandem duplication (bottom). Consensus sequence alignment of locally assembled Oxford Nanopore Technologies long sequencing reads to chromosomes 2 and 5 of the human reference genome (top). Breakpoints are circled and marked as 1 (beginning of tandem duplication), 2 (end of tandem duplication) or 3 (inverted templated insertion). For each breakpoint, the middle panel shows Illumina short reads at SV breakpoints. c, Association between rare germline PTVs (MAF < 0.5%) and somatic CpG mutagenesis (approximately with signature 1) in individuals of European ancestry (n = 1,201). Genes highlighted in blue or red were associated with lower or higher somatic mutation rates. Two-sided hypothesis testing was performed using linear-regression models with sex, age at diagnosis and cancer project as variables. To mitigate multiple-hypothesis testing, the significance threshold was set to exome-wide significance (P < 2.5 × 10−6). The black line represents the identity line that would be followed if the observed P values followed the null expectation; the shaded area shows the 95% confidence intervals. d, Catalogue of polymorphic germline L1 source elements that are active in cancer. The chromosomal map shows germline source L1 elements as volcano symbols. Each volcano is colour-coded according to the type of source L1 activity. The contribution of each source locus (expressed as a percentage) to the total number of transductions identified in PCAWG tumours is represented as a gradient of volcano size, with top contributing elements exhibiting larger sizes.
Second, we performed a rare-variant association study (MAF <0.5%) to investigate the relationship between germline PTVs and somatic DNA rearrangements in individuals with European ancestry (Extended Data Fig. 11a–c). Germline BRCA2 and BRCA1 PTVs were associated with an increased burden of small (less than 10 kb) somatic SV deletions (P = 1 × 10−8) and tandem duplications (P = 6 × 10−13), respectively, corroborating recent studies in breast and ovarian cancer30,70. In PCAWG data, this pattern also extends to other tumour types, including adenocarcinomas of the prostate and pancreas6, typically in the setting of biallelic inactivation. In addition, tumours with high levels of small SV tandem duplications frequently exhibited a novel and distinct class of SVs termed ‘cycles of templated insertions’6. These complex SV events consist of DNA templates that are copied from across the genome, joined into one contiguous sequence and inserted into a single derivative chromosome. We found a significant association between germline BRCA1 PTVs and templated insertions at the pan-cancer level (P = 4 × 10−15) (Extended Data Fig. 11d, e). Whole-genome long-read sequencing data generated for a BRCA1-deficient PCAWG prostate tumour verified the small tandem-duplication and templated-insertion SV phenotypes (Fig. 6b). Almost all (20 out of 21) of BRCA1-associated tumours with a templated-insertion SV phenotype displayed combined germline and somatic hits in the gene. Together, these data suggest that biallelic inactivation of BRCA1 is a driver of the templated-insertion SV phenotype.
Third, rare-variant association analysis revealed that patients with germline MBD4 PTVs had increased rates of somatic C > T mutation rates at CpG dinucleotides (P < 2.5 × 10−6) (Fig. 6c and Extended Data Fig. 11f, g). Analysis of previously published whole-exome sequencing samples from the TCGA (n = 8,134) replicated the association between germline MBD4 PTVs and increased somatic CpG mutagenesis at the pan-cancer level (P = 7.1 × 10−4) (Extended Data Fig. 11h). Moreover, gene-expression profiling revealed a significant but modest correlation between MBD4 expression and somatic CpG mutation rates between and within PCAWG tumour types (Extended Data Fig. 11i–k). MBD4 encodes a DNA-repair gene that removes thymidines from T:G mismatches within methylated CpG sites71, a functionality that would be consistent with a CpG mutational signature in cancer.
Fourth, we assessed long interspersed nuclear elements (LINE-1; L1 hereafter) that mediate somatic retrotransposition events72,73,74. We identified 114 germline source L1 elements capable of active somatic retrotransposition, including 70 that represent insertions with respect to the human reference genome (Fig. 6d and Supplementary Table 7), and 53 that were tagged by single-nucleotide polymorphisms in strong linkage disequilibrium (Supplementary Table 7). Only 16 germline L1 elements accounted for 67% (2,440 out of 3,669) of all L1-mediated transductions10 detected in the PCAWG dataset (Extended Data Fig. 12a). These 16 hot-L1 elements followed two broad patterns of somatic activity (8 of each), which we term Strombolian and Plinian in analogy to patterns of volcanic activity. Strombolian L1s are frequently active in cancer, but mediate only small-to-modest eruptions of somatic L1 activity in cancer samples (Extended Data Fig. 12b). By contrast, Plinian L1s are more rarely seen, but display aggressive somatic activity. Whereas Strombolian elements are typically relatively common (MAF > 2%) and sometimes even fixed in the human population, all Plinian elements were infrequent (MAF ≤ 2%) in PCAWG donors (Extended Data Fig. 12c; P = 0.001, Mann–Whitney U-test). This dichotomous pattern of activity and allele frequency may reflect differences in age and selective pressures, with Plinian elements potentially inserted into the human germline more recently. PCAWG donors bear on average between 50 and 60 L1 source elements and between 5 and 7 elements with hot activity (Extended Data Fig. 12d), but only 38% (1,075 out of 2,814) of PCAWG donors carried ≥1 Plinian element. Some L1 germline source loci caused somatic loss of tumour-suppressor genes (Extended Data Fig. 12e). Many are restricted to individual continental population ancestries (Extended Data Fig. 12f–j).
Replicative immortality
One of the hallmarks of cancer is the ability of cancer to evade cellular senescence21. Normal somatic cells typically have finite cell division potential; telomere attrition is one mechanism to limit numbers of mitoses75. Cancers enlist multiple strategies to achieve replicative immortality. Overexpression of the telomerase gene, TERT, which maintains telomere lengths, is especially prevalent. This can be achieved through point mutations in the promoter that lead to de novo transcription factor binding34,37; hitching TERT to highly active regulatory elements elsewhere in the genome46,76; insertions of viral enhancers upstream of the gene77,78; and increased dosage through chromosomal amplification, as we have seen in melanoma (Fig. 5b). In addition, there is an ‘alternative lengthening of telomeres’ (ALT) pathway, in which telomeres are lengthened through homologous recombination, mediated by loss-of-function mutations in the ATRX and DAXX genes79.
As reported in a companion paper13, 16% of tumours in the PCAWG dataset exhibited somatic mutations in at least one of ATRX, DAXX and TERT. TERT alterations were detected in 270 samples, whereas 128 tumours had alterations in ATRX or DAXX, of which 71 were protein-truncating. In the companion paper, which focused on describing patterns of ALT and TERT-mediated telomere maintenance13, 12 features of telomeric sequence were measured in the PCAWG cohort. These included counts of nine variants of the core hexameric sequence, the number of ectopic telomere-like insertions within the genome, the number of genomic breakpoints and telomere length as a ratio between tumour and normal. Here we used the 12 features as an overview of telomere integrity across all tumours in the PCAWG dataset.
On the basis of these 12 features, tumour samples formed 4 distinct subclusters (Fig. 7a and Extended Data Fig. 13a), suggesting that telomere-maintenance mechanisms are more diverse than the well-established TERT and ALT dichotomy. Clusters C1 (47 tumours) and C2 (42 tumours) were enriched for traits of the ALT pathway—having longer telomeres, more genomic breakpoints, more ectopic telomere insertions and variant telomere sequence motifs (Supplementary Fig. 9). C1 and C2 were distinguished from one another by the latter having a considerable increase in the number of TTCGGG and TGAGGG variant motifs among the telomeric hexamers. Thyroid adenocarcinomas were markedly enriched among C3 samples (26 out of 33 C3 samples; P < 10−16); the C1 cluster (ALT subtype 1) was common among sarcomas; and both pancreatic endocrine neoplasms and low-grade gliomas had a high proportion of samples in the C2 cluster (ALT subtype 2) (Fig. 7b). Notably, some of the thyroid adenocarcinomas and pancreatic neuroendocrine tumours that cluster together (cluster C3) had matched normal samples that also cluster together (normal cluster N3) (Extended Data Fig. 13a) and which share common properties. For example, the GTAGGG repeat was overrepresented among samples in this group (Supplementary Fig. 10).
a, Scatter plot of the clusters of telomere patterns identified across PCAWG using t-distributed stochastic neighbour embedding (t-SNE), based on n = 2,518 tumour samples and their matched normal samples. Axes have arbitrary dimensions such that samples with similar telomere profiles are clustered together and samples with dissimilar telomere profiles are far apart with high probability. b, Distribution of the four tumour-specific clusters of telomere patterns in selected tumour types from PCAWG. c, Distribution of relevant driver mutations associated with alternative lengthening of telomere and normal telomere maintenance across the four clusters. d, Distribution of telomere maintenance abnormalities across tumour types with more than 40 patients in PCAWG. Samples were classified as tumour clusters 1–3 if they fell into a relevant cluster without mutations in TERT, ATRX or DAXX and had no ALT phenotype. TMM, telomere maintenance mechanisms.
Somatic driver mutations were also unevenly distributed across the four clusters (Fig. 7c). C1 tumours were enriched for RB1 mutations or SVs (P = 3 × 10−5), as well as frequent SVs that affected ATRX (P = 6 × 10−14), but not DAXX. RB1 and ATRX mutations were largely mutually exclusive (Extended Data Fig. 13b). By contrast, C2 tumours were enriched for somatic point mutations in ATRX and DAXX (P = 6 × 10−5), but not RB1. The enrichment of RB1 mutations in C1 remained significant when only leiomyosarcomas and osteosarcomas were considered, confirming that this enrichment is not merely a consequence of the different distribution of tumour types across clusters. C3 samples had frequent TERT promoter mutations (30%; P = 2 × 10−6).
There was a marked predominance of RB1 mutations in C1. Nearly a third of the samples in C1 contained an RB1 alteration, which were evenly distributed across truncating SNVs, SVs and shallow deletions (Extended Data Fig. 13c). Previous research has shown that RB1 mutations are associated with long telomeres in the absence of TERT mutations and ATRX inactivation80, and studies using mouse models have shown that knockout of Rb-family proteins causes elongated telomeres81. The association with the C1 cluster here suggests that RB1 mutations can represent another route to activating the ALT pathway, which has subtly different properties of telomeric sequence compared with the inactivation of DAXX—these fall almost exclusively in cluster C2.
Tumour types with the highest rates of abnormal telomere maintenance mechanisms often originate in tissues that have low endogenous replicative activity (Fig. 7d). In support of this, we found an inverse correlation between previously estimated rates of stem cell division across tissues82 and the frequency of telomere maintenance abnormalities (P = 0.01, Poisson regression) (Extended Data Fig. 13d). This suggests that restriction of telomere maintenance is an important tumour-suppression mechanism, particularly in tissues with low steady-state cellular proliferation, in which a clone must overcome this constraint to achieve replicative immortality.
Conclusions and future perspectives
The resource reported in this paper and its companion papers has yielded insights into the nature and timing of the many mutational processes that shape large- and small-scale somatic variation in the cancer genome; the patterns of selection that act on these variations; the widespread effect of somatic variants on transcription; the complementary roles of the coding and non-coding genome for both germline and somatic mutations; the ubiquity of intratumoral heterogeneity; and the distinctive evolutionary trajectory of each cancer type. Many of these insights can be obtained only from an integrated analysis of all classes of somatic mutation on a whole-genome scale, and would not be accessible with, for example, targeted exome sequencing.
The promise of precision medicine is to match patients to targeted therapies using genomics. A major barrier to its evidence-based implementation is the daunting heterogeneity of cancer chronicled in these papers, from tumour type to tumour type, from patient to patient, from clone to clone and from cell to cell. Building meaningful clinical predictors from genomic data can be achieved, but will require knowledge banks comprising tens of thousands of patients with comprehensive clinical characterization83. As these sample sizes will be too large for any single funding agency, pharmaceutical company or health system, international collaboration and data sharing will be required. The next phase of ICGC, ICGC-ARGO (https:// www.icgc-argo.org/), will bring the cancer genomics community together with healthcare providers, pharmaceutical companies, data science and clinical trials groups to build comprehensive knowledge banks of clinical outcome and treatment data from patients with a wide variety of cancers, matched with detailed molecular profiling.
Extending the story begun by TCGA, ICGC and other cancer genomics projects, the PCAWG has brought us closer to a comprehensive narrative of the causal biological changes that drive cancer phenotypes. We must now translate this knowledge into sustainable, meaningful clinical treatments.
Methods
Samples
We compiled an inventory of matched tumour–normal whole-cancer genomes in the ICGC Data Coordinating Centre. Most samples came from treatment-naive, primary cancers, although a small number of donors had multiple samples of primary, metastatic and/or recurrent tumours. Our inclusion criteria were: (1) matched tumour and normal specimen pair; (2) a minimal set of clinical fields; and (3) characterization of tumour and normal whole genomes using Illumina HiSeq paired-end sequencing reads.
We collected genome data from 2,834 donors, representing all ICGC and TCGA donors that met these criteria at the time of the final data freeze in autumn 2014 (Extended Data Table 1). After quality assurance (Supplementary Methods 2.5), data from 176 donors were excluded as unusable, 75 had minor issues that could affect some analyses (grey-listed donors) and 2,583 had data of optimal quality (white-listed donors) (Supplementary Table 1). Across the 2,658 white- and grey-listed donors, whole-genome sequences were available from 2,605 primary tumours and 173 metastases or local recurrences. Matching normal samples were obtained from blood (2,064 donors), tissue adjacent to the primary tumour (87 donors) or from distant sites (507 donors). Whole-genome sequencing data were available for tumour and normal DNA for the entire cohort. The mean read coverage was 39× for normal samples, whereas tumours had a bimodal coverage distribution with modes at 38× and 60× (Supplementary Fig. 1). The majority of specimens (65.3%) were sequenced using 101-bp paired-end reads. An additional 28% were sequenced with 100-bp paired-end reads. Of the remaining specimens, 4.7% were sequenced with read lengths longer than 101 bp, and 1.9% with read lengths shorter than 100 bp. The distribution of read lengths by tumour cohort is shown in Supplementary Fig. 11. Median read length for whole-genome sequencing paired-end reads was 101 bp (mean = 106.2, s.d. = 16.7; minimum–maximum = 50–151). RNA-sequencing data were collected and re-analysed centrally for 1,222 donors, including 1,178 primary tumours, 67 metastases or local recurrences and 153 matched normal tissue samples adjacent to the primary tumour.
Demographically, the cohort included 1,469 men (55%) and 1,189 women (45%), with a mean age of 56 years (range, 1–90 years) (Supplementary Table 1). Using population ancestry-differentiated single nucleotide polymorphisms, the ancestry distribution was heavily weighted towards donors of European descent (77% of total) followed by East Asians (16%), as expected for large contributions from European, North American and Australian projects (Supplementary Table 1).
We consolidated histopathology descriptions of the tumour samples, using the ICD-0-3 tumour site controlled vocabulary89. Overall, the PCAWG dataset comprises 38 distinct tumour types (Extended Data Table 1 and Supplementary Table 1). Although the most common tumour types are included in the dataset, their distribution does not match the relative population incidences, largely owing to differences among contributing ICGC/TCGA groups in the numbers of sequenced samples.
Uniform processing and somatic variant calling
To generate a consistent set of somatic mutation calls that could be used for cross-tumour analyses, we analysed all 6,835 samples using a uniform set of algorithms for alignment, variant calling and quality control (Extended Data Fig. 1, Supplementary Fig. 2, Supplementary Table 3 and Supplementary Methods 2). We used the BWA-MEM algorithm90 to align each tumour and normal sample to human reference build hs37d5 (as used in the 1000 Genomes Project91). Somatic mutations were identified in the aligned data using three established pipelines, which were run independently on each tumour–normal pair. Each of the three pipelines—labelled ‘Sanger’92,93,94,95, ‘EMBL/DKFZ’96,97 and ‘Broad’98,99,100,101 after the computational biology groups that created or assembled them—consisted of multiple software packages for calling somatic SNVs, small indels, CNAs and somatic SVs (with intrachromosomal SVs defined as those >100 bp). Two additional variant algorithms102,103 were included to further improve accuracy across a broad range of clonal and subclonal mutations. We tested different merging strategies using validation data, and choses the optimal method for each variant type to generate a final consensus set of mutation calls (Supplementary Methods S2.4).
Somatic retrotransposition events, including Alu and LINE-1 insertions72, L1-mediated transductions73 and pseudogene formation104, were called using a dedicated pipeline73. We removed these retrotransposition events from the somatic SV call-set. Mitochondrial DNA mutations were called using a published algorithm105. RNA-sequencing data were uniformly processed to quantify normalized gene-level expression, splicing variation and allele-specific expression, and to identify fusion transcripts, alternative promoter usage and sites of RNA editing8.
Integration, phasing and validation of germline variant call-sets
Calls of common (≥1% frequency in PCAWG) and rare (<1%) germline variants including single-nucleotide polymorphisms, indels, SVs and mobile-element insertions (MEIs) were generated using a population-scale genetic polymorphism-detection approach91,106. The uniform germline data-processing workflow comprised variant identification using six different variant-calling algorithms96,107,108 and was orchestrated using the Butler workflow system109.
We performed call-set benchmarking, merging, variant genotyping and statistical haplotype-block phasing91 (Supplementary Methods 3.4). Using this strategy, we identified 80.1 million germline single-nucleotide polymorphisms, 5.9 million germline indels, 1.8 million multi-allelic short (<50 bp) germline variants, as well as germline SVs ≥ 50 bp in size including 29,492 biallelic deletions and 27,254 MEIs (Supplementary Table 2). We statistically phased this germline variant set using haplotypes from the 1000 Genomes Project91 as a reference panel, yielding an N50-phased block length of 265 kb based on haploid chromosomes from donor-matched tumour genomes. Precision estimates for germline SNVs and indels were >99% for the phased merged call-set, and sensitivity estimates ranged from 92% to 98%.
Core alignment and variant calling by cloud computing
The requirement to uniformly realign and call variants on nearly 5,800 whole genomes (tumour plus normal) presented considerable computational challenges, and raised ethical issues owing to the use of data from different jurisdictions (Extended Data Table 2). To process the data, we adopted a cloud-computing architecture26 in which the alignment and variant calling was spread across 13 data centres on 3 continents, representing a mixture of commercial, infrastructure-as-a-service, academic cloud compute and traditional academic high-performance computer clusters (Supplementary Table 3). Together, the effort used 10 million CPU-core hours.
To generate reproducible variant calling across the 13 data centres, we built the core pipelines into Docker containers28, in which the workflow description, required code and all associated dependencies were packaged together in stand-alone packages. These heavily tested, extensively validated workflows are available for download (Box 1).
Validation, benchmarking and merging of somatic variant calls
To evaluate the performance of each of the mutation-calling pipelines and determine an integration strategy, we performed a large-scale deep-sequencing validation experiment (Supplementary Notes 1). We selected a pilot set of 63 representative tumour–normal pairs, on which we ran the 3 core pipelines, together with a set of 10 additional somatic variant-calling pipelines contributed by members of the PCAWG SNV Calling Methods Working Group. Sufficient DNA remained for 50 of the 63 cases for validation, which was performed by hybridization of tumour and matched normal DNA to a custom RNA bait set, followed by deep sequencing, as previously described29. Although performed using the same sequencing chemistry as the original whole-genome sequencing analyses, the considerably greater depth achieved in the validation experiment enabled accurate assessment of sensitivity and precision of variant calls. Variant calls in repeat-masked regions were not tested, owing to the challenge of designing reliable validation probes in these areas.
The 3 core pipelines had individual estimates of sensitivity of 80–90% to detect a true somatic SNV called by any of the 13 pipelines; with >95% of SNV calls made by each of the core pipelines being genuine somatic variants (Fig. 1a). For indels—a more-challenging class of variants to identify in short-read sequencing data—the 3 core algorithms had individual sensitivity estimates in the range of 40–50%, with precision 70–95% (Fig. 1b). Validation of SV calls is inherently more difficult, as methods based on PCR or hybridization to RNA baits often fail to isolate DNA that spans the breakpoint. To assess the accuracy of SV calls, we therefore used the property that an SV must either generate a copy-number change or be balanced, whereas artefactual calls will not respect this property. For individual SV-calling algorithms, we estimated precision to be in the range of 80–95% for samples in the 63-sample pilot dataset.
Next, we examined multiple methods for merging calls made by several algorithms into a single definitive call-set to be used for downstream analysis. The final consensus calls for SNVs were based on a simple approach that required two or more methods to agree on a call. For indels, because methods were less concordant, we used stacked logistic regression110,111 to integrate the calls. The merged SV set includes all calls made by two or more of the four primary SV-calling algorithms96,100,112,113. Consensus CNA calls were obtained by joining the outputs of six individual CNA-calling algorithms with SV consensus breakpoints to obtain base-pair resolution CNAs (Supplementary Methods 2.4.3). Consensus purity and ploidy were derived, and a multitier system was developed for consensus copy-number calls (Supplementary Methods 2.4.3, and described in detail elsewhere7).
Overall, the sensitivity and precision of the consensus somatic variant calls were 95% (90% confidence interval, 88–98%) and 95% (90% confidence interval, 71–99%), respectively, for SNVs (Extended Data Fig. 2). For somatic indels, sensitivity and precision were 60% (90% confidence interval, 34–72%) and 91% (90% confidence interval, 73–96%), respectively. Regarding SVs, we estimate the sensitivity of the merging algorithm to be 90% for true calls generated by any one calling pipeline; precision was estimated to be 97.5%. That is, 97.5% of SVs in the merged SV call-set had an associated copy-number change or balanced partner rearrangement. The improvement in calling accuracy from combining different pipelines was most noticeable in variants that had low variant allele fractions, which are likely to originate from subclonal populations of the tumour (Fig. 1c, d). There remains much work to be done to improve indel calling software; we still lack sensitivity for calling even fully clonal complex indels from short-read sequencing data.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this paper.
Data availability
The PCAWG-generated alignments, somatic variant calls, annotations and derived datasets are available for general research use for browsing and download at http://dcc.icgc.org/pcawg/ (Box 1 and Supplementary Table 4). In accordance with the data access policies of the ICGC and TCGA projects, most molecular, clinical and specimen data are in an open tier which does not require access approval. To access potentially identifying information, such as germline alleles and underlying read data, researchers will need to apply to the TCGA Data Access Committee (DAC) via dbGaP (https://dbgap.ncbi.nlm.nih.gov/aa/wga.cgi?page=login) for access to the TCGA portion of the dataset, and to the ICGC Data Access Compliance Office (DACO; http://icgc.org/daco) for the ICGC portion. In addition, to access somatic single nucleotide variants derived from TCGA donors, researchers will also need to obtain dbGaP authorization.
Beyond the core sequence data and variant call-sets, the analyses in this paper used a number of datasets that were derived from the variant calls (Supplementary Table 4). The individual datasets are available at Synapse (https://www.synapse.org/), and are denoted with synXXXXX accession numbers; all these datasets are also mirrored at https://dcc.icgc.org, with full links, filenames, accession numbers and descriptions detailed in Supplementary Table 4. The datasets encompass: clinical data from each patient including demographics, tumour stage and vital status (syn10389158); harmonized tumour histopathology annotations using a standardised hierarchical ontology (syn1038916); inferred purity and ploidy values for each tumour sample (syn8272483); driver mutations for each patient from their cancer genome spanning all classes of variant, and coding versus non-coding drivers (syn11639581); mutational signatures inferred from PCAWG donors (syn11804065), including APOBEC mutagenesis (syn7437313); and transcriptional data from RNA sequencing, including gene expression levels (syn5553985, syn5553991, syn8105922) and gene fusions (syn10003873, syn7221157).
Code availability
Computational pipelines for calling somatic mutations are available to the public at https://dockstore.org/organizations/PCAWG/collections/PCAWG. A range of data-visualization and -exploration tools are also available for the PCAWG data (Box 1).
Change history
25 January 2023
A Correction to this paper has been published: https://doi.org/10.1038/s41586-022-05598-w
References
Pleasance, E. D. et al. A comprehensive catalogue of somatic mutations from a human cancer genome. Nature 463, 191–196 (2010).
Pleasance, E. D. et al. A small-cell lung cancer genome with complex signatures of tobacco exposure. Nature 463, 184–190 (2010).
Ley, T. J. et al. DNA sequencing of a cytogenetically normal acute myeloid leukaemia genome. Nature 456, 66–72 (2008).
Rheinbay, E. et al. Analyses of non-coding somatic drivers in 2,693 cancer whole genomes. Nature https://doi.org/10.1038/s41586-020-1965-x (2020).
Alexandrov, L. B. et al. The repertoire of mutational signatures in human cancer. Nature https://doi.org/10.1038/s41586-020-1943-3 (2020).
Li, Y. et al. Patterns of somatic structural variation in human cancer genomes. Nature https://doi.org/10.1038/s41586-019-1913-9 (2020).
Gerstung, M. et al. The evolutionary history of 2,658 cancers. Nature https://doi.org/10.1038/s41586-019-1907-7 (2020).
PCAWG Transcriptome Core Group et al. Genomic basis of RNA alterations in cancer. Nature https://doi.org/10.1038/s41586-020-1970-0 (2020).
Zhang, Y. et al. High-coverage whole-genome analysis of 1,220 cancers reveals hundreds of genes deregulated by rearrangement-mediated cis-regulatory alterations. Nat. Commun. https://doi.org/10.1038/s41467-019-13885-w (2020).
Rodriguez-Martin, B. et al. Pan-cancer analysis of whole genomes identifies driver rearrangements promoted by LINE-1 retrotransposition. Nat. Genet. https://doi.org/10.1038/s41588-019-0562-0 (2020).
Zapatka, M. et al. The landscape of viral associations in human cancers. Nat. Genet. https://doi.org/10.1038/s41588-019-0558-9 (2020).
Jiao, W. et al. A deep learning system can accurately classify primary and metastatic cancers based on patterns of passenger mutations. Nat. Commun. https://doi.org/10.1038/s41467-019-13825-8 (2020).
Sieverling, L. et al. Genomic footprints of activated telomere maintenance mechanisms in cancer. Nat. Commun. https://doi.org/10.1038/s41467-019-13824-9 (2020).
Yuan, Y. et al. Comprehensive molecular characterization of mitochondrial genomes in human cancers. Nat. Genet. https://doi.org/10.1038/s41588-019-0557-x (2020).
Akdemir, K. C. et al. Chromatin folding domains disruptions by somatic genomic rearrangements in human cancers. Nat. Genet. https://doi.org/10.1038/s41588-019-0564-y (2020).
Reyna, M. A. et al. Pathway and network analysis of more than 2,500 whole cancer genomes. Nat. Commun. https://doi.org/10.1038/s41467-020-14351-8 (2020).
Bailey, M. H. et al. Retrospective evaluation of whole exome and genome mutation calls in 746 cancer samples. Nat. Commun. (2020).
Cortes-Ciriano, I. et al. Comprehensive analysis of chromothripsis in 2,658 human cancers using whole-genome sequencing. Nat. Genet. https://doi.org/10.1038/s41588-019-0576-7 (2020).
Bray, F., Ren, J.-S., Masuyer, E. & Ferlay, J. Global estimates of cancer prevalence for 27 sites in the adult population in 2008. Int. J. Cancer 132, 1133–1145 (2013).
Tarver, T. Cancer Facts & Figures 2012. American Cancer Society (ACS). J. Consum. Health Internet 16, 366–367 (2012).
Hanahan, D. & Weinberg, R. A. Hallmarks of cancer: the next generation. Cell 144, 646–674 (2011).
International Cancer Genome Consortium. International network of cancer genome projects. Nature 464, 993–998 (2010).
Bailey, M. H. et al. Comprehensive characterization of cancer driver genes and mutations. Cell 173, 371–385 (2018).
Sanchez-Vega, F. et al. Oncogenic signaling pathways in The Cancer Genome Atlas. Cell 173, 321–337 (2018).
Hoadley, K. A. et al. Cell-of-origin patterns dominate the molecular classification of 10,000 tumors from 33 types of cancer. Cell 173, 291–304 (2018).
Stein, L. D., Knoppers, B. M., Campbell, P., Getz, G. & Korbel, J. O. Data analysis: create a cloud commons. Nature 523, 149–151 (2015).
Phillips, M. et al. Genomics: data sharing needs international code of conduct. Nature https://doi.org/10.1038/d41586-020-00082-9 (2020).
Krochmalski, J. Developing with Docker (Packt Publishing, 2016).
Welch, J. S. et al. The origin and evolution of mutations in acute myeloid leukemia. Cell 150, 264–278 (2012).
Nik-Zainal, S. et al. Landscape of somatic mutations in 560 breast cancer whole-genome sequences. Nature 534, 47–54 (2016).
Meier, B. et al. C. elegans whole-genome sequencing reveals mutational signatures related to carcinogens and DNA repair deficiency. Genome Res. 24, 1624–1636 (2014).
Martincorena, I. et al. Universal patterns of selection in cancer and somatic tissues. Cell 171, 1029–1041 (2017).
Tamborero, D. et al. Cancer Genome Interpreter annotates the biological and clinical relevance of tumor alterations. Genome Med. 10, 25 (2018).
Huang, F. W. et al. Highly recurrent TERT promoter mutations in human melanoma. Science 339, 957–959 (2013).
Rheinbay, E. et al. Recurrent and functional regulatory mutations in breast cancer. Nature 547, 55–60 (2017).
Fredriksson, N. J., Ny, L., Nilsson, J. A. & Larsson, E. Systematic analysis of noncoding somatic mutations and gene expression alterations across 14 tumor types. Nat. Genet. 46, 1258–1263 (2014).
Horn, S. et al. TERT promoter mutations in familial and sporadic melanoma. Science 339, 959–961 (2013).
Ciriello, G. et al. Emerging landscape of oncogenic signatures across human cancers. Nat. Genet. 45, 1127–1133 (2013).
Rahman, N. Realizing the promise of cancer predisposition genes. Nature 505, 302–308 (2014).
Pearl, L. H., Schierz, A. C., Ward, S. E., Al-Lazikani, B. & Pearl, F. M. G. Therapeutic opportunities within the DNA damage response. Nat. Rev. Cancer 15, 166–180 (2015).
Taylor-Weiner, A. et al. DeTiN: overcoming tumor-in-normal contamination. Nat. Methods 15, 531–534 (2018).
Fujimoto, A. et al. Whole-genome mutational landscape and characterization of noncoding and structural mutations in liver cancer. Nat. Genet. 48, 500–509 (2016).
Shlush, L. I. Age-related clonal hematopoiesis. Blood 131, 496–504 (2018).
Northcott, P. A. et al. The whole-genome landscape of medulloblastoma subtypes. Nature 547, 311–317 (2017).
Scarpa, A. et al. Whole-genome landscape of pancreatic neuroendocrine tumours. Nature 543, 65–71 (2017).
Davis, C. F. et al. The somatic genomic landscape of chromophobe renal cell carcinoma. Cancer Cell 26, 319–330 (2014).
Berger, M. F. et al. The genomic complexity of primary human prostate cancer. Nature 470, 214–220 (2011).
Baca, S. C. et al. Punctuated evolution of prostate cancer genomes. Cell 153, 666–677 (2013).
Nik-Zainal, S. et al. The life history of 21 breast cancers. Cell 149, 994–1007 (2012).
Nik-Zainal, S. et al. Mutational processes molding the genomes of 21 breast cancers. Cell 149, 979–993 (2012).
Roberts, S. A. et al. Clustered mutations in yeast and in human cancers can arise from damaged long single-strand DNA regions. Mol. Cell 46, 424–435 (2012).
Rausch, T. et al. Genome sequencing of pediatric medulloblastoma links catastrophic DNA rearrangements with TP53 mutations. Cell 148, 59–71 (2012).
Stephens, P. J. et al. Massive genomic rearrangement acquired in a single catastrophic event during cancer development. Cell 144, 27–40 (2011).
Korbel, J. O. & Campbell, P. J. Criteria for inference of chromothripsis in cancer genomes. Cell 152, 1226–1236 (2013).
Zhang, C.-Z. et al. Chromothripsis from DNA damage in micronuclei. Nature 522, 179–184 (2015).
The Cancer Genome Atlas Research Network. Integrated genomic characterization of papillary thyroid carcinoma. Cell 159, 676–690 (2014).
Supek, F. & Lehner, B. Clustered mutation signatures reveal that error-prone DNA repair targets mutations to active genes. Cell 170, 534–547 (2017).
Mardin, B. R. et al. A cell-based model system links chromothripsis with hyperploidy. Mol. Syst. Biol. 11, 828 (2015).
Weischenfeldt, J. et al. Integrative genomic analyses reveal an androgen-driven somatic alteration landscape in early-onset prostate cancer. Cancer Cell 23, 159–170 (2013).
Garsed, D. W. et al. The architecture and evolution of cancer neochromosomes. Cancer Cell 26, 653–667 (2014).
Durinck, S. et al. Temporal dissection of tumorigenesis in primary cancers. Cancer Discov. 1, 137–143 (2011).
Hayward, N. K. et al. Whole-genome landscapes of major melanoma subtypes. Nature 545, 175–180 (2017).
The Cancer Genome Atlas Network. Genomic classification of cutaneous melanoma. Cell 161, 1681–1696 (2015).
Alexandrov, L. B. et al. Signatures of mutational processes in human cancer. Nature 500, 415–421 (2013).
Chan, K. et al. An APOBEC3A hypermutation signature is distinguishable from the signature of background mutagenesis by APOBEC3B in human cancers. Nat. Genet. 47, 1067–1072 (2015).
Nik-Zainal, S. et al. Association of a germline copy number polymorphism of APOBEC3A and APOBEC3B with burden of putative APOBEC-dependent mutations in breast cancer. Nat. Genet. 46, 487–491 (2014).
Middlebrooks, C. D. et al. Association of germline variants in the APOBEC3 region with cancer risk and enrichment with APOBEC-signature mutations in tumors. Nat. Genet. 48, 1330–1338 (2016).
Westra, H.-J. et al. Systematic identification of trans eQTLs as putative drivers of known disease associations. Nat. Genet. 45, 1238–1243 (2013).
Stranger, B. E. et al. Population genomics of human gene expression. Nat. Genet. 39, 1217–1224 (2007).
Menghi, F. et al. The tandem duplicator phenotype as a distinct genomic configuration in cancer. Proc. Natl Acad. Sci. USA 113, E2373–E2382 (2016).
Hendrich, B., Hardeland, U., Ng, H. H., Jiricny, J. & Bird, A. The thymine glycosylase MBD4 can bind to the product of deamination at methylated CpG sites. Nature 401, 301–304 (1999).
Lee, E. et al. Landscape of somatic retrotransposition in human cancers. Science 337, 967–971 (2012).
Tubio, J. M. C. et al. Extensive transduction of nonrepetitive DNA mediated by L1 retrotransposition in cancer genomes. Science 345, 1251343–1251343 (2014).
Helman, E. et al. Somatic retrotransposition in human cancer revealed by whole-genome and exome sequencing. Genome Res. 24, 1053–1063 (2014).
Shay, J. W. & Wright, W. E. Hayflick, his limit, and cellular ageing. Nat. Rev. Mol. Cell Biol. 1, 72–76 (2000).
Peifer, M. et al. Telomerase activation by genomic rearrangements in high-risk neuroblastoma. Nature 526, 700–704 (2015).
Totoki, Y. et al. Trans-ancestry mutational landscape of hepatocellular carcinoma genomes. Nat. Genet. 46, 1267–1273 (2014).
Paterlini-Bréchot, P. et al. Hepatitis B virus-related insertional mutagenesis occurs frequently in human liver cancers and recurrently targets human telomerase gene. Oncogene 22, 3911–3916 (2003).
Heaphy, C. M. et al. Prevalence of the alternative lengthening of telomeres telomere maintenance mechanism in human cancer subtypes. Am. J. Pathol. 179, 1608–1615 (2011).
Barthel, F. P. et al. Systematic analysis of telomere length and somatic alterations in 31 cancer types. Nat. Genet. 49, 349–357 (2017).
García-Cao, M., Gonzalo, S., Dean, D. & Blasco, M. A. A role for the Rb family of proteins in controlling telomere length. Nat. Genet. 32, 415–419 (2002).
Tomasetti, C. & Vogelstein, B. Variation in cancer risk among tissues can be explained by the number of stem cell divisions. Science 347, 78–81 (2015).
Gerstung, M. et al. Precision oncology for acute myeloid leukemia using a knowledge bank approach. Nat. Genet. 49, 332–340 (2017).
O’Connor, B. D. et al. The Dockstore: enabling modular, community-focused sharing of Docker-based genomics tools and workflows. F1000Res. 6, 52 (2017).
Zhang, J. et al. The International Cancer Genome Consortium Data Portal. Nat. Biotechnol. 37, 367–369 (2019).
Miller, C. A., Qiao, Y., DiSera, T., D’Astous, B. & Marth, G. T. bam.iobio: a web-based, real-time, sequence alignment file inspector. Nat. Methods 11, 1189–1189 (2014).
Goldman, M. et al. The UCSC Xena platform for public and private cancer genomics data visualization and interpretation. Preprint at https://www.biorxiv.org/content/10.1101/326470v6 (2019).
Papatheodorou, I. et al. Expression Atlas: gene and protein expression across multiple studies and organisms. Nucleic Acids Res. 46, D246–D251 (2018).
NCI SEER. ICD-O-3 Coding Materials (2018).
Li, H. & Durbin, R. Fast and accurate long-read alignment with Burrows–Wheeler transform. Bioinformatics 26, 589–595 (2010).
1000 Genomes Project Consortium. A global reference for human genetic variation. Nature 526, 68–74 (2015).
Raine, K. M. et al. ascatNgs: identifying somatically acquired copy-number alterations from whole-genome sequencing data. Curr. Protoc. Bioinformatics 56, 15.9.1–15.9.17 (2016).
Jones, D. et al. cgpCaVEManWrapper: simple execution of CaVEMan in order to detect somatic single nucleotide variants in NGS data. Curr. Protoc. Bioinformatics 56, 15.10.1–15.10.18 (2016).
Raine, K. M. et al. cgpPindel: identifying somatically acquired insertion and deletion events from paired end sequencing. Curr. Protoc. Bioinformatics 52, 15.7.1–15.7.12 (2015).
Ye, K., Schulz, M. H., Long, Q., Apweiler, R. & Ning, Z. Pindel: a pattern growth approach to detect break points of large deletions and medium sized insertions from paired-end short reads. Bioinformatics 25, 2865–2871 (2009).
Rausch, T. et al. DELLY: structural variant discovery by integrated paired-end and split-read analysis. Bioinformatics 28, i333–i339 (2012).
Rimmer, A. et al. Integrating mapping-, assembly- and haplotype-based approaches for calling variants in clinical sequencing applications. Nat. Genet. 46, 912–918 (2014).
Cibulskis, K. et al. Sensitive detection of somatic point mutations in impure and heterogeneous cancer samples. Nat. Biotechnol. 31, 213–219 (2013).
Carter, S. L. et al. Absolute quantification of somatic DNA alterations in human cancer. Nat. Biotechnol. 30, 413–421 (2012).
Drier, Y. et al. Somatic rearrangements across cancer reveal classes of samples with distinct patterns of DNA breakage and rearrangement-induced hypermutability. Genome Res. 23, 228–235 (2013).
Ramos, A. H. et al. Oncotator: cancer variant annotation tool. Hum. Mutat. 36, E2423–E2429 (2015).
Moncunill, V. et al. Comprehensive characterization of complex structural variations in cancer by directly comparing genome sequence reads. Nat. Biotechnol. 32, 1106–1112 (2014).
Fan, Y. et al. MuSE: accounting for tumor heterogeneity using a sample-specific error model improves sensitivity and specificity in mutation calling from sequencing data. Genome Biol. 17, 178 (2016).
Cooke, S. L. et al. Processed pseudogenes acquired somatically during cancer development. Nat. Commun. 5, 3644 (2014).
Ju, Y. S. et al. Origins and functional consequences of somatic mitochondrial DNA mutations in human cancer. eLife 3, e02935 (2014).
Sudmant, P. H. et al. An integrated map of structural variation in 2,504 human genomes. Nature 526, 75–81 (2015).
Garrison, E. & Marth, G. Haplotype-based variant detection from short-read sequencing. Preprint at https://arxiv.org/abs/1207.3907 (2012).
DePristo, M. A. et al. A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nat. Genet. 43, 491–498 (2011).
Yakneen, S., Waszak, S. M., Gertz, M. & Korbel, J. O. & PCAWG Consortium. Butler enables rapid cloud-based analysis of thousands of human genomes. Nat. Biotechnol. https://doi.org/10.1038/s41587-019-0360-3 (2020).
Kim, S. Y., Jacob, L. & Speed, T. P. Combining calls from multiple somatic mutation-callers. BMC Bioinformatics 15, 154 (2014).
Breiman, L. Stacked regressions. Mach. Learn. 24, 49–64 (1996).
Campbell, P. J. et al. Identification of somatically acquired rearrangements in cancer using genome-wide massively parallel paired-end sequencing. Nat. Genet. 40, 722–729 (2008).
Wala, J. A. et al. SvABA: genome-wide detection of structural variants and indels by local assembly. Genome Res. 28, 581–591 (2018).
Acknowledgements
We thank research participants who donated samples and data, the physicians and clinical staff who contributed to sample annotation and collection, and the numerous funding agencies that contributed to the collection and analysis of this dataset.
Author information
Authors and Affiliations
Consortia
Contributions
Writing committee leads: Peter J. Campbell, Gad Getz, Jan O. Korbel, Joshua M. Stuart, Jennifer L. Jennings, Lincoln D. Stein. Head of project management: Jennifer L. Jennings. Sample collection: major contributions from Marc D. Perry, Hardeep K. Nahal-Bose; led by B. F. Francis Ouellette. Histopathology harmonization: major contribution from Constance H. Li; further contributions from Esther Rheinbay, G. Petur Nielsen, Dennis C. Sgroi, Chin-Lee Wu, William C. Faquin, Vikram Deshpande, Paul C. Boutros, Alexander J. Lazar, Katherine A. Hoadley; led by Lincoln D. Stein, David N. Louis. Uniform processing, somatic, germline variant calling: major contribution from L. Jonathan Dursi; further contributions from Christina K. Yung, Matthew H. Bailey, Gordon Saksena, Keiran M. Raine, Ivo Buchhalter, Kortine Kleinheinz, Matthias Schlesner, Junjun Zhang, Wenyi Wang, David A. Wheeler; led by Li Ding, Jared T. Simpson. Core alignment, variant calling by cloud computing: major contributions from Christina K. Yung, Brian D. O’Connor, Sergei Yakneen, Junjun Zhang; further contributions from Kyle Ellrott, Kortine Kleinheinz, Naoki Miyoshi, Keiran M. Raine, Adam P. Butler, Romina Royo, Gordon Saksena, Matthias Schlesner, Solomon I. Shorser, Miguel Vazquez. Integration, phasing, validation of germline variant callsets: major contributions from Tobias Rausch, Grace Tiao, Sebastian M. Waszak, Bernardo Rodriguez-Martin, Suyash Shringarpure, Dai-Ying Wu; further contributions from Sergei Yakneen, German M. Demidov, Olivier Delaneau, Shuto Hayashi, Seiya Imoto, Nina Habermann, Ayellet V. Segre, Erik Garrison, Andy Cafferkey, Eva G. Alvarez, José María Heredia-Genestar, Francesc Muyas, Oliver Drechsel, Alicia L. Bruzos, Javier Temes, Jorge Zamora, L. Jonathan Dursi, Adrian Baez-Ortega, Hyung-Lae Kim, Matthew H. Bailey, R. Jay Mashl, Kai Ye, Ivo Buchhalter, Anthony DiBiase, Kuan-lin Huang, Ivica Letunic, Michael D. McLellan, Steven J. Newhouse, Matthias Schlesner, Tal Shmaya, Sushant Kumar, David C. Wedge, Mark H. Wright, Venkata D. Yellapantula, Mark Gerstein, Ekta Khurana, Tomas Marques-Bonet, Arcadi Navarro, Carlos D. Bustamante, Jared T. Simpson, Li Ding, Reiner Siebert, Hidewaki Nakagawa, Douglas F. Easton; led by Stephan Ossowski, Jose M. C. Tubio, Gad Getz, Francisco M. De La Vega, Xavier Estivill, Jan O. Korbel. Validation, benchmarking, merging of somatic variant calls: major contribution from L. Jonathan Dursi; further contributions from David A. Wheeler, Christina K. Yung; led by Li Ding, Jared T. Simpson. Data and code availability: major contribution from Junjun Zhang; further contributions from Christina K. Yung, Sergei Yakneen, Denis Yuen, George L. Mihaiescu, Larsson Omberg; led by Vincent Ferretti. Pan-cancer burden of somatic mutations: major contribution from Junjun Zhang; led by Peter J. Campbell. Panorama of driver mutations in human cancer: led by Radhakrishnan Sabarinathan, Oriol Pich, Abel Gonzalez-Perez. PCAWG tumours with no apparent driver mutations: major contribution from Esther Rheinbay; further contributions from Amaro Taylor-Weiner, Radhakrishnan Sabarinathan; led by Peter J. Campbell, Gad Getz. Patterns, oncogenicity of kataegis, chromoplexy: major contributions from Matthew W. Fittall, Jonas Demeulemeester, Maxime Tarabichi; further contributions from Nicola D. Roberts, Peter J. Campbell, Jan O. Korbel; led by Peter Van Loo. Patterns, oncogenicity of chromothripsis: major contributions from Maxime Tarabichi, Jonas Demeulemeester, Matthew W. Fittall; further contributions from Isidro Cortes-Ciriano, Lara Urban, Peter J. Park, Peter J. Campbell, Jan O. Korbel; led by Peter Van Loo. Timing-clustered mutational processes during tumour evolution: major contributions from Jonas Demeulemeester, Maxime Tarabichi, Matthew W. Fittall; further contributions from Jan O. Korbel, Peter J. Campbell; led by Peter Van Loo. Germline effects on somatic mutation: major contributions from Sebastian M. Waszak, Bin Zhu, Bernardo Rodriguez-Martin, Esa Pitkanen, Tobias Rausch; further contributions from Yilong Li, Natalie Saini, Leszek J. Klimczak, Joachim Weischenfeldt, Nikos Sidiropoulos, Ludmil B. Alexandrov, Francesc Muyas, Raquel Rabionet, Georgia Escaramis, Adrian Baez-Ortega, Mattia Bosio, Aliaksei Z. Holik, Hana Susak, Eva G. Alvarez, Alicia L. Bruzos, Javier Temes, Aparna Prasad, Nina Habermann, Serap Erkek, Lara Urban, Claudia Calabrese, Benjamin Raeder, Eoghan Harrington, Simon Mayes, Daniel Turner, Sissel Juul, Steven A. Roberts, Lei Song, Roelof Koster, Lisa Mirabello, Xing Hua, Tomas J. Tanskanen, Marta Tojo, David C. Wedge, Jorge Zamora, Jieming Chen, Lauri A. Aaltonen, Gunnar Ratsch, Roland F. Schwarz, Atul J. Butte, Alvis Brazma, Peter J. Campbell, Stephen J. Chanock, Nilanjan Chatterjee, Oliver Stegle, Olivier Harismendy; led by G. Steven Bova, Dmitry A. Gordenin, Jose M. C. Tubio, Douglas F. Easton, Xavier Estivill, Jan O. Korbel. Replicative immortality: major contribution from David Haan; further contributions from Lina Sieverling, Lars Feuerbach; led by Lincoln D. Stein, Joshua M. Stuart. Ethical considerations of genomic cloud computing: led by Don Chalmers, Yann Joly, Bartha Knoppers, Fruzsina Molnar-Gabor, Jan O. Korbel, Mark Phillips, Adrian Thorogood, David Townend. Online resources for data access, visualization, exploration and analysis: major contributions from Mary Goldman, Junjun Zhang, Nuno A. Fonseca; further contributions from Qian Xiang, Brian Craft, Elena Pineiro-Yanez, Alfonso Munoz, Robert Petryszak, Anja Fullgrabe, Fatima Al-Shahrour, Maria Keays, David Haussler, John Weinstein, Wolfgang Huber, Alfonso Valencia, Irene Papatheodorou, Jingchun Zhu; led by Brian D. O’Connor, Lincoln D. Stein, Alvis Brazma, Vincent Ferretti, Miguel Vazquez. The 63-sample pilot-analysis validation process: major contribution from L. Jonathan Dursi; further contributions from Christina K. Yung, Matthew H. Bailey, Gordon Saksena, Keiran M. Raine, Ivo Buchhalter, Kortine Kleinheinz, Matthias Schlesner, Yu Fan, David Torrents, Matthias Bieg, Paul C. Boutros, Ken Chen, Zechen Chong, Kristian Cibulskis, Oliver Drechsel, Roland Eils, Robert S. Fulton, Josep Gelpi, Mark Gerstein, Santiago Gonzalez, Gad Getz, Ivo G. Gut, Faraz Hach, Michael Heinold, Taobo Hu, Vincent Huang, Barbara Hutter, Hyung-Lae Kim, Natalie Jager, Jongsun Jung, Sushant Kumar, Yogesh Kumar, Christopher Lalansingh, Ignaty Leshchiner, Ivica Letunic, Dimitri Livitz, Eric Z. Ma, Yosef E. Maruvka, R. Jay Mashl, Michael D. McLellan, Ana Milovanovic, Morten Muhlig Nielsen, Brian D. O’Connor, Stephan Ossowski, Nagarajan Paramasivam, Jakob Skou Pedersen, Marc D. Perry, Montserrat Puiggros, Romina Royo, Esther Rheinbay, S. Cenk Sahinalp, Iman Sarrafi, Chip Stewart, Miranda D. Stobbe, Grace Tiao, Jeremiah A. Wala, Jiayin Wang, Wenyi Wang, Sebastian M. Waszak, Joachim Weischenfeldt, Michael Wendl, Johannes Werner, Zhenggang Wu, Hong Xue, Sergei Yakneen, Takafumi N. Yamaguchi, Kai Ye, Venkata Yellapantula, Junjun Zhang, David A. Wheeler; led by Li Ding, Jared T. Simpson. Processing of validation data: major contributions from Christina K. Yung, Brian D. O’Connor, Sergei Yakneen, Junjun Zhang; further contributions from Kyle Ellrott, Kortine Kleinheinz, Naoki Miyoshi, Keiran M. Raine, Romina Royo, Gordon Saksena, Matthias Schlesner, Solomon I. Shorser, Miguel Vazquez, Joachim Weischenfeldt, Denis Yuen, Adam P. Butler, Brandi N. Davis-Dusenbery, Roland Eils, Vincent Ferretti, Robert L. Grossman, Olivier Harismendy, Youngwook Kim, Hidewaki Nakagawa, Steven J. Newhouse, David Torrents; led by Lincoln D. Stein. Whole-genome sequencing somatic variant calling: major contribution from Junjun Zhang; further contributions from Christina K. Yung, Solomon I. Shorser. Whole-genome alignment: Keiran M. Raine, Junjun Zhang, Brian D. O’Connor. DKFZ pipeline: Kortine Kleinheinz, Tobias Rausch, Jan O. Korbel, Ivo Buchhalter, Michael C. Heinold, Barbara Hutter, Natalie Jager, Nagarajan Paramasivam, Matthias Schlesner. EMBL pipeline: Joachim Weischenfeldt. Sanger pipeline: Keiran M. Raine, Jonathan Hinton, David R. Jones, Andrew Menzies, Lucy Stebbings, Adam P. Butler. Broad pipeline: Gordon Saksena, Dimitri Livitz, Esther Rheinbay, Julian M. Hess, Ignaty Leshchiner, Chip Stewart, Grace Tiao, Jeremiah A. Wala, Amaro Taylor-Weiner, Mara Rosenberg, Andrew J. Dunford, Manaswi Gupta, Marcin Imielinski, Matthew Meyerson, Rameen Beroukhim, Gad Getz. MuSE Pipeline: Yu Fan, Wenyi Wang. Consensus somatic SNV/indel annotation: Andrew Menzies, Matthias Schlesner, Juri Reimand, Priyanka Dhingra, Ekta Khurana. Somatic SNV, indel merging: major contribution from L. Jonathan Dursi; further contributions from Christina K. Yung, Matthew H. Bailey, Gordon Saksena, Keiran M. Raine, Ivo Buchhalter, Kortine Kleinheinz, Matthias Schlesner, Yu Fan, David Torrents, Matthias Bieg, Paul C. Boutros, Ken Chen, Zechen Chong, Kristian Cibulskis, Oliver Drechsel, Roland Eils, Robert S. Fulton, Josep L. Gelpi, Mark Gerstein, Santiago Gonzalez, Gad Getz, Ivo G. Gut, Faraz Hach, Michael Heinold, Taobo Hu, Vincent Huang, Barbara Hutter, Hyung-Lae Kim, Natalie Jager, Jongsun Jung, Sushant Kumar, Yogesh Kumar, Christopher Lalansingh, Ignaty Leshchiner, Ivica Letunic, Dimitri Livitz, Eric Z. Ma, Yosef E. Maruvka, R. Jay Mashl, Michael D. McLellan, Ana Milovanovic, Morten Muhlig Nielsen, Brian D. O’Connor, Stephan Ossowski, Nagarajan Paramasivam, Jakob Skou Pedersen, Marc D. Perry, Montserrat Puiggros, Romina Royo, Esther Rheinbay, S. Cenk Sahinalp, Iman Sarrafi, Chip Stewart, Miranda D. Stobbe, Grace Tiao, Jeremiah A. Wala, Jiayin Wang, Wenyi Wang, Sebastian M. Waszak, Joachim Weischenfeldt, Michael Wendl, Johannes Werner, Zhenggang Wu, Hong Xue, Sergei Yakneen, Takafumi N. Yamaguchi, Kai Ye, Venkata Yellapantula, Junjun Zhang, David A. Wheeler; major contributions from Li Ding, Jared T. Simpson. Somatic SV merging: Joachim Weischenfeldt, Francesco Favero, Yilong Li. Somatic CNA merging: Stefan Dentro, Jeff Wintersinger, Ignaty Leshchiner. Oxidative artefact filtration: Dimitri Livitz, Ignaty Leshchiner, Chip Stewart, Esther Rheinbay, Gordon Saksena, Gad Getz. Strand bias filtration: Matthias Bieg, Ivo Buchhalter, Johannes Werner, Matthias Schlesner. miniBAM generation: Jeremiah A. Wala, Gordon Saksena, Rameen Beroukhim, Gad Getz. Germline variant identification from whole-genome sequencing: major contributions from Tobias Rausch, Grace Tiao, Sebastian M. Waszak, Bernardo Rodriguez-Martin, Suyash Shringarpure, Dai-Ying Wu; further contributions from Sergei Yakneen, German M. Demidov, Olivier Delaneau, Shuto Hayashi, Seiya Imoto, Nina Habermann, Ayellet V. Segre, Erik Garrison, Andy Cafferkey, Eva G. Alvarez, Alicia L. Bruzos, Jorge Zamora, José María Heredia-Genestar, Francesc Muyas, Oliver Drechsel, L. Jonathan Dursi, Adrian Baez-Ortega, Hyung-Lae Kim, Matthew H. Bailey, R. Jay Mashl, Kai Ye, Ivo Buchhalter, Vasilisa Rudneva, Ji Wan Park, Eun Pyo Hong, Seong Gu Heo, Anthony DiBiase, Kuan-lin Huang, Ivica Letunic, Michael D. McLellan, Steven J. Newhouse, Matthias Schlesner, Tal Shmaya, Sushant Kumar, David C. Wedge, Mark H. Wright, Venkata D. Yellapantula, Mark Gerstein, Ekta Khurana, Tomas Marques-Bonet, Arcadi Navarro, Carlos D. Bustamante, Jared T. Simpson, Li Ding, Reiner Siebert, Hidewaki Nakagawa, Douglas F. Easton; led by Stephan Ossowski, Jose M. C. Tubio, Gad Getz, Francisco M. De La Vega, Xavier Estivill, Jan O. Korbel. RNA-sequencing analysis: major contributions from Nuno A. Fonseca, Andre Kahles, Kjong-Van Lehmann, Lara Urban, Cameron M. Soulette, Yuichi Shiraishi, Fenglin Liu, Yao He, Deniz Demircioglu, Natalie R. Davidson, Claudia Calabrese, Junjun Zhang, Marc D. Perry, Qian Xiang; further contributions from Liliana Greger, Siliang Li, Dongbing Liu, Stefan G. Stark, Fan Zhang, Samirkumar B. Amin, Peter Bailey, Aurelien Chateigner, Isidro Cortes-Ciriano, Brian Craft, Serap Erkek, Milana Frenkel-Morgenstern, Mary Goldman, Katherine A. Hoadley, Yong Hou, Matthew R. Huska, Ekta Khurana, Helena Kilpinen, Jan O. Korbel, Fabien C. Lamaze, Chang Li, Xiaobo Li, Xinyue Li, Xingmin Liu, Maximillian G. Marin, Julia Markowski, Tannistha Nandi, Morten Muhlig Nielsen, Akinyemi I. Ojesina, Qiang Pan-Hammarstrom, Peter J. Park, Chandra Sekhar Pedamallu, Jakob Skou Pedersen, Reiner Siebert, Hong Su, Patrick Tan, Bin Tean Teh, Jian Wang, Sebastian M. Waszak, Heng Xiong, Sergei Yakneen, Chen Ye, Christina Yung, Xiuqing Zhang, Liangtao Zheng, Jingchun Zhu, Shida Zhu, Philip Awadalla, Chad J. Creighton, Matthew Meyerson, B. F. Francis Ouellette, Kui Wu, Huanming Yang; led by Jonathan Goke, Roland F. Schwarz, Oliver Stegle, Zemin Zhang, Alvis Brazma, Gunnar Ratsch, Angela N. Brooks. Clustering of tumour genomes based on telomere maintenance-related features: major contribution from David Haan; led by Lincoln D. Stein, Joshua M. Stuart. Clustered mutational processes in PCAWG: major contributions from Jonas Demeulemeester, Maxime Tarabichi, Matthew W. Fittall; led by Peter J. Campbell, Jan O. Korbel, Peter Van Loo. Tumours without detected driver mutations: Esther Rheinbay, Amaro Taylor-Weiner, Radhakrishnan Sabarinathan, Peter J. Campbell, Gad Getz. Panorama of driver mutations in human cancer: major contributions from Radhakrishnan Sabarinathan, Oriol Pich; further contributions from Inigo Martincorena, Carlota Rubio-Perez, Malene Juul, Jeremiah A. Wala, Steven Schumacher, Ofer Shapira, Nikos Sidiropoulos, Sebastian M. Waszak, David Tamborero, Loris Mularoni, Esther Rheinbay, Henrik Hornshoj, Jordi Deu-Pons, Ferran Muinos, Johanna Bertl, Qianyun Guo, Chad J. Creighton, Joachim Weischenfeldt, Jan O. Korbel, Gad Getz, Peter J. Campbell, Jakob Skou Pedersen, Rameen Beroukhim; led by Abel Gonzalez-Perez. Pilot benchmarking, variant consensus development and validation: major contribution from L. Jonathan Dursi; further contributions from Christina K. Yung, Matthew H. Bailey, Gordon Saksena, Keiran M. Raine, Ivo Buchhalter, Kortine Kleinheinz, Matthias Schlesner, Yu Fan, David Torrents, Matthias Bieg, Paul C. Boutros, Ken Chen, Zechen Chong, Kristian Cibulskis, Oliver Drechsel, Roland Eils, Robert S. Fulton, Josep Gelpi, Mark Gerstein, Santiago Gonzalez, Gad Getz, Ivo G. Gut, Faraz Hach, Michael Heinold, Taobo Hu, Vincent Huang, Barbara Hutter, Hyung-Lae Kim, Natalie Jager, Jongsun Jung, Sushant Kumar, Yogesh Kumar, Christopher Lalansingh, Ignaty Leshchiner, Ivica Letunic, Dimitri Livitz, Eric Z. Ma, Yosef E. Maruvka, R. Jay Mashl, Michael D. McLellan, Ana Milovanovic, Morten Muhlig Nielsen, Brian D. O’Connor, Stephan Ossowski, Nagarajan Paramasivam, Jakob Skou Pedersen, Marc D. Perry, Montserrat Puiggros, Romina Royo, Esther Rheinbay, S. Cenk Sahinalp, Iman Sarrafi, Chip Stewart, Miranda D. Stobbe, Grace Tiao, Jeremiah A. Wala, Jiayin Wang, Wenyi Wang, Sebastian M. Waszak, Joachim Weischenfeldt, Michael Wendl, Johannes Werner, Zhenggang Wu, Hong Xue, Sergei Yakneen, Takafumi N. Yamaguchi, Kai Ye, Venkata Yellapantula, Junjun Zhang, David A. Wheeler; led by Li Ding, Jared T. Simpson. Production somatic variant calling on the PCAWG compute cloud: major contributions from Christina K. Yung, Brian D. O’Connor, Sergei Yakneen, Junjun Zhang; further contributions from Kyle Ellrott, Kortine Kleinheinz, Naoki Miyoshi, Keiran M. Raine, Romina Royo, Gordon Saksena, Matthias Schlesner, Solomon I. Shorser, Miguel Vazquez, Joachim Weischenfeldt, Denis Yuen, Adam P. Butler, Brandi N. Davis-Dusenbery, Roland Eils, Vincent Ferretti, Robert L. Grossman, Olivier Harismendy, Youngwook Kim, Hidewaki Nakagawa, Steven J Newhouse, David Torrents; led by Lincoln D. Stein. PCAWG data portals: major contributions from Mary Goldman, Junjun Zhang, Nuno A. Fonseca, Isidro Cortes-Ciriano; further contributions from Qian Xiang, Brian Craft, Elena Pineiro-Yanez, Brian D O’Connor, Wojciech Bazant, Elisabet Barrera, Alfonso Munoz, Robert Petryszak, Anja Fullgrabe, Fatima Al-Shahrour, Maria Keays, David Haussler, John Weinstein, Wolfgang Huber, Alfonso Valencia, Irene Papatheodorou, Jingchun Zhu; led by Vincent Ferretti, Miguel Vazquez.
Corresponding authors
Ethics declarations
Competing interests
Gad Getz receives research funds from IBM and Pharmacyclics and is an inventor on patent applications related to MuTect, ABSOLUTE, MutSig, MSMuTect, MSMutSig and POLYSOLVER. Hikmat Al-Ahmadie is consultant for AstraZeneca and Bristol-Myers Squibb. Samuel Aparicio is a founder and shareholder of Contextual Genomics. Pratiti Bandopadhayay receives grant funding from Novartis for an unrelated project. Rameen Beroukhim owns equity in Ampressa Therapeutics. Andrew Biankin receives grant funding from Celgene, AstraZeneca and is a consultant for or on advisory boards of AstraZeneca, Celgene, Elstar Therapeutics, Clovis Oncology and Roche. Ewan Birney is a consultant for Oxford Nanopore, Dovetail and GSK. Marcus Bosenberg is a consultant for Eli Lilly. Atul Butte is a cofounder of and consultant for Personalis, NuMedii, a consultant for Samsung, Geisinger Health, Mango Tree Corporation, Regenstrief Institute and in the recent past a consultant for 10x Genomics and Helix, a shareholder in Personalis, a minor shareholder in Apple, Twitter, Facebook, Google, Microsoft, Sarepta, 10x Genomics, Amazon, Biogen, CVS, Illumina, Snap and Sutro and has received honoraria and travel reimbursement for invited talks from Genentech, Roche, Pfizer, Optum, AbbVie and many academic institutions and health systems. Carlos Caldas has served on the Scientific Advisory Board of Illumina. Lorraine Chantrill acted on an advisory board for AMGEN Australia in the past 2 years. Andrew D. Cherniack receives research funding from Bayer. Helen Davies is an inventor on a number of patent applications that encompass the use of mutational signatures. Francisco De La Vega was employed at Annai Systems during part of the project. Ronny Drapkin serves on the scientific advisory board of Repare Therapeutics and Siamab Therapeutics. Rosalind Eeles has received an honorarium for the GU-ASCO meeting in San Francisco in January 2016 as a speaker, a honorarium and support from Janssen for the RMH FR meeting in November 2017 as a speaker (title: genetics and prostate cancer), a honorarium for an University of Chicago invited talk in May 2018 as speaker and an educational honorarium paid by Bayer & Ipsen to attend GU Connect ‘Treatment sequencing for mCRPC patients within the changing landscape of mHSPC’ at a venue at ESMO, Barcelona, on 28 September 2019. Paul Flicek is a member of the scientific advisory boards of Fabric Genomics and Eagle Genomics. Ronald Ghossein is a consultant for Veracyte. Dominik Glodzik is an inventor on a number of patent applications that encompass the use of mutational signatures. Eoghan Harrington is a full-time employee of Oxford Nanopore Technologies and is a stock holder. Yann Joly is responsible for the Data Access Compliance Office (DACO) of ICGC 2009-2018. Sissel Juul is a full-time employee of Oxford Nanopore Technologies and is a stock holder. Vincent Khoo has received personal fees and non-financial support from Accuray, Astellas, Bayer, Boston Scientific and Janssen. Stian Knappskog is a coprincipal investigator on a clinical trial that receives research funding from AstraZeneca and Pfizer. Ignaty Leshchiner is a consultant for PACT Pharma. Carlos López-Otín has ownership interest (including stock and patents) in DREAMgenics. Matthew Meyerson is a scientific advisory board chair of, and consultant for, OrigiMed, has obtained research funding from Bayer and Ono Pharma and receives patent royalties from LabCorp. Serena Nik-Zainal is an inventor on a number of patent applications that encompass the use of mutational signatures. Nathan Pennell has done consulting work with Merck, Astrazeneca, Eli Lilly and Bristol-Myers Squibb. Xose S. Puente has ownership interest (including stock and patents in DREAMgenics. Benjamin J. Raphael is a consultant for and has ownership interest (including stock and patents) in Medley Genomics. Jorge Reis-Filho is a consultant for Goldman Sachs and REPARE Therapeutics, member of the scientific advisory board of Volition RX and Paige.AI and an ad hoc member of the scientific advisory board of Ventana Medical Systems, Roche Tissue Diagnostics, InVicro, Roche, Genentech and Novartis. Lewis R. Roberts has received grant support from ARIAD Pharmaceuticals, Bayer, BTG International, Exact Sciences, Gilead Sciences, Glycotest, RedHill Biopharma, Target PharmaSolutions and Wako Diagnostics and has provided advisory services to Bayer, Exact Sciences, Gilead Sciences, GRAIL, QED Therapeutics and TAVEC Pharmaceuticals. Richard A. Scolyer has received fees for professional services from Merck Sharp & Dohme, GlaxoSmithKline Australia, Bristol-Myers Squibb, Dermpedia, Novartis Pharmaceuticals Australia, Myriad, NeraCare GmbH and Amgen. Tal Shmaya is employed at Annai Systems. Reiner Siebert has received speaker honoraria from Roche and AstraZeneca. Sabina Signoretti is a consultant for Bristol-Myers Squibb, AstraZeneca, Merck, AACR and NCI and has received funding from Bristol-Myers Squibb, AstraZeneca, Exelixis and royalties from Biogenex. Jared Simpson has received research funding and travel support from Oxford Nanopore Technologies. Anil K. Sood is a consultant for Merck and Kiyatec, has received research funding from M-Trap and is a shareholder in BioPath. Simon Tavaré is on the scientific advisory board of Ipsen and a consultant for Kallyope. John F. Thompson has received honoraria and travel support for attending advisory board meetings of GlaxoSmithKline and Provectus and has received honoraria for participation in advisory boards for MSD Australia and BMS Australia. Daniel Turner is a full-time employee of Oxford Nanopore Technologies and is a stock holder. Naveen Vasudev has received speaker honoraria and/or consultancy fees from Bristol-Myers Squibb, Pfizer, EUSA pharma, MSD and Novartis. Jeremiah A. Wala is a consultant for Nference. Daniel J. Weisenberger is a consultant for Zymo Research. Dai-Ying Wu is employed at Annai Systems. Cheng-Zhong Zhang is a cofounder and equity holder of Pillar Biosciences, a for-profit company that specializes in the development of targeted sequencing assays. The other authors declare no competing interests.
Additional information
Peer review information Nature thanks Arul Chinnaiyan, Ben Lehner, Nicolas Robine and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Flow-chart showing key steps in the analysis of PCAWG genomes.
After alignment to the genome, somatic mutations were identified by three pipelines, with subsequent merging into a consensus variant set used for downstream scientific analyses. Subs, substitutions; DKFZ/EMBL, the German Cancer Research Centre (DKFZ) and Europen Molecular Biology Laboratory (EMBL).
Extended Data Fig. 2 Distribution of accuracy estimates across algorithms and samples from validation data.
a, F1 accuracy, precision and sensitivity estimates for somatic SNVs across the core algorithms and different approaches to merging the call-sets. The box plots demarcate the interquartile range and median of estimates across the n = 50 samples in the validation dataset. b, F1 accuracy, precision and sensitivity estimates for somatic indels (n = 50 samples). SVM, support vector machine; union, calls made by all variant-calling algorithms; intersect2, calls made by any combination of two variant-calling algorithms; intersect3, calls made by any three variant-calling algorithms.
Extended Data Fig. 3 Distribution of numbers of somatic mutations of different classes across tumour types.
The y axis is on a log scale. The 2,583 donors with the highest quality metrics (white-listed donors) are plotted. SNVs indicate substitutions; indels are taken as insertions or deletions <100 bp in size; retrotranspositions are the combined counts of somatic retrotransposon insertions, transductions and somatic pseudogene insertions.
Extended Data Fig. 4 Patients with no detected driver mutations in PCAWG.
a, Number (red) of patients without detected driver mutations distributed across the different tumour types studied. b, Estimated sensitivity for detecting somatic point mutations genome-wide across tumour types (total sample size: n = 2,583 patients). Each point represents the estimate for a single patient, layered on violin plots that show the estimated density distribution of sensitivity values for that tumour type (the width proportional is to density). c, SETD2 expression levels across different medulloblastoma subtypes. Points represent individual patients, coloured by whether the gene exhibited focal copy number (CN) loss or a truncating point mutation, or was the wild-type gene. The coloured areas are violin plots showing the estimated density distribution of expression values for that medulloblastoma subtype.
Extended Data Fig. 5 Examples of clustered mutational processes.
a, Chromoplexy example in a thyroid adenocarcinoma. Genes at the breakpoints are schematically depicted in their normal genomic context and again in the reconstructed derivative chromosomes below. b, Distinct kataegis signatures in the genome of a pancreatic adenocarcinoma sample. SVs and their classification are shown above the main rainfall plot, as well as the total and minor allele copy number. Tra, translocation; del, deletion; dup, duplication; t2tInv, tail-to-tail inversion; h2hInv, head-to-head inversion. Magnifications of the three foci on chromosomes 1, 8 and 12, respectively, highlight distinct manifestations of kataegis. Left, a novel process similar to signature 17 with T > N mutations at CT or TT dinucleotides. Middle, the prototypical APOBEC3A/B type with C > T (signature 2) and/or C>G/A (signature 13) substitutions at TpC. Right, an alternative cytidine deaminase(s) with a preference for substitutions at C/GpC. Most of the SNVs in each of these foci can be phased to the same allele and no evidence of anti-phasing is observed. c, Example of a chromothripsis event in a melanoma. The black points (top) represent copy-number estimates from individual genomic bins, with SVs shown as coloured arcs (translocation in black, deletion in purple, duplication in brown, tail-to-tail inversion in cyan, head-to-head inversion in green) that mostly demarcate copy-number changes. The mate chromosomes are displayed above translocations. Bottom, the variant allele fractions of somatic mutations distributed along the relevant chromosomal region.
Extended Data Fig. 6 Patterns of intense kataegis.
a, Distribution of the tumour types (colour-coded as in Extended Data Fig. 3) of the samples in the top 5% of kataegis intensity in each of the four identified genome-wide patterns: non-APOBEC, replication stress, rearrangement-associated and the combination of the last two. b, c, Distribution of leading/lagging strand (b) and replication timing bias (c) for rearrangement-(in)dependent APOBEC kataegis, based on n = 2,583 tumours. P values were derived using a two-sided Mann–Whittney U-test. d, Example rainfall plots for each of the four identified kataegis patterns.
Extended Data Fig. 7 Association of chromothripsis with covariates and driver events.
a, Odds ratios per cancer type of containing chromothripsis in whole-genome duplicated versus diploid samples (n = 2,583 patients). ***q < 0.001; **q < 0.01; *q < 0.05. Two-sided hypothesis testing was performed using Fisher–Boschloo tests, corrected for multiple-hypothesis testing. b, Same as a for female versus male. c, Proportion of mutations explained by single-base substitution signature 1 and age at diagnosis in prostate cancer samples (n = 210 patients) with or without chromothripsis (q < 0.05). The early-onset prostate cancer project drives the signal and was sequenced at lower depth. For the box-and-whisker plots, the box denotes the interquartile range, with the median marked as a horizontal line. The whiskers extend as far as the range or 1.5× the interquartile range, whichever is less. Two-sided hypothesis testing was performed using Mann–Whitney U-tests. d, Counts of co-occurrence of chromothripsis with amplification (blue) and homozygous deletions (red) in driver regions: observed (thick line) versus randomized (shaded area and thin line). The cumulative number of drivers that were hit is plotted as a function of the number of times those drivers were hit. e, For each sample in which chromothripsis coincided with a driver event in those genes, we show the fold change in gene expression compared to the median expression of the gene in non-chromothripsis samples of the same cancer type, coloured by cancer type and shaped by the type of driver event. We show with added transparency the fold changes calculated the same way for samples with driver mutations hitting the same driver genes, but that had no evidence of chromothripsis. Analysis is based on n = 1,222 patients with RNA-sequencing data. f, Enrichment of co-occurrence of chromothripsis with driver events. The x axis shows the association of chromothripsis with a driver in a given cancer type compared with its rate of association with that driver in all other cancer types. The y axis shows the association of chromothripsis with a driver in a given cancer type compared with its rate of association with all other drivers in that type. Exact binomial tests are used and P values are corrected for multiple testing according to the Benjamini–Hochberg method.
Extended Data Fig. 8 Further examples of chromothripsis-induced amplification targeting multiple cancer-associated genes simultaneously in melanoma.
a, Examples of amplifications that occurred early in the development of melanoma. The black points (top) represent copy-number estimates from individual genomic bins, with SVs shown as coloured arcs (translocation in black, deletion in purple, duplication in brown, tail-to-tail inversion in cyan and head-to-head inversion in green) that mostly demarcate copy-number changes. Bottom, the variant allele fractions of SNVs distributed along the relevant chromosomal region. The paucity of somatic mutations at high variant allele fractions in the most-heavily amplified regions indicates that these amplifications began very early in tumour evolution, before the lineage had had opportunity to acquire many SNVs. b, Example of an amplification that occurred late in melanoma development. The large numbers of somatic mutations at high variant allele fractions in the most-heavily amplified regions indicate that these amplifications began late in tumour evolution, after the lineage had already acquired many SNVs.
Extended Data Fig. 9 Timing the amplifications after chromothripsis in molecular time for 10 representative cases.
a, Copy-number plot of chromothriptic regions categorized as ‘liposarc-like’ in five acral melanomas with CCND1 amplification. Segments indicate the copy number of the major allele. Points represent SNV multiplicities, that is, the estimated number of copies carrying each SNV, coloured by base change and shaped by strand. Small vertical arrows link SNVs to their corresponding copy-number segment. Kataegis foci are shown within black boxes and show typical strand specificities (all triangles or all circles), similar multiplicities and base changes of signatures 2 and 13 (red and black, respectively). A coloured bar (top right) represents the molecular timing of the amplification (red bar; high is early, low is late) and is coloured by the fraction of total SNVs assigned to the following timing categories: clonal [early], clonal mutations that occurred before duplications involving the relevant chromosome (including whole-genome duplications); clonal [late], clonal mutations that occurred after such duplications; and clonal [NA], mutations that occurred when no duplication was observed. b, Same as a in two cutaneous melanomas, one shows early amplification, the other late amplification. c, Same as a, b, for three lung squamous cell carcinomas and late amplification of SOX2.
Extended Data Fig. 10 Association between common germline variants and endogenous mutational processes.
Genome-wide association of somatic CpG mutagenesis in individuals of European ancestry (n = 1,201 patients) based on mutational signature analysis (a) and NpCpG motif analysis (b). Two-sided hypothesis testing was performed using PLINK v.1.9. To mitigate multiple-hypothesis testing, the significance threshold was set to genome-wide significance (P < 5 × 10−8). c, d, Locuszoom plot for somatic APOBEC3B-like mutagenesis association results, linkage disequilibrium and recombination rates around the genome-wide significant 22q13.1 locus in individuals with European (c) and East Asian (d) ancestry (n = 1,201 and 318 patients, respectively). Locuszoom plot for somatic APOBEC3B-like mutagenesis association results around the 22q13.1 locus in individuals with European (e) and East Asian (f) ancestry after conditioning on rs12628403. g, h, Association between rs2142833 and expression of APOBEC3 genes in PCAWG tumour samples (adjusted for sex, age at diagnosis, histology and population structure in linear-regression models with two-sided hypothesis testing not corrected for multiple tests). For the box-and-whisker plot, the box denotes the interquartile range, with the median marked as a horizontal line. The whiskers extend as far as the range or 1.5× the interquartile range, whichever is less. Outliers are shown as points.
Extended Data Fig. 11 Association between rare germline PTVs in protein-coding genes and somatic mutational phenotypes.
a–d, f, Data are based on two-sided rare-variant association testing across n = 2,583 patients, with a stringent P value threshold of P < 2.5 × 10−6 used to mitigate multiple-hypothesis testing (significant genes marked with coloured circles). Blue/red circles mark genes that decrease/increase somatic mutation rates. The black line represents the identity line that would be followed if the observed P values followed the null expectation, with the shaded area showing the 95% confidence intervals. a, QQ plots for the proportion of somatic SV deletions, tandem duplications, inversions and translocation in cancer genomes. b, QQ plots for the proportion of somatic SV deletions in cancer genomes stratified by four size groups (1–10 kb, 10–100 kb, 100–1,000 kb and >1,000 kb). c, QQ plots for the proportion of somatic SV tandem duplications in cancer genomes stratified by four size groups (1–10 kb, 10–100 kb, 100–1,000 kb and >1,000 kb). d, QQ plot for the presence or absence of somatic SV templated insertion (cycles) in cancer genomes. e, Number of SV-templated insertion cycles in PCAWG tumours with germline BRCA1 PTVs. Only histological samples with at least one germline BRCA1 PTV carrier are shown (n = 1,095 patients combined). The box denotes the interquartile range, with the median marked as a horizontal line. The whiskers extend as far as the range or 1.5× the interquartile range, whichever is less. Outliers are shown as points. f, QQ plot for somatic CpG mutagenesis in cancer genomes based on NpCpG motif analysis. g, Violin plots show estimated densities of the proportion of somatic CpG mutations in PCAWG donors with germline MBD4 and BRCA2 PTVs. The box denotes the interquartile range, with the median marked as a white point. The whiskers extend as far as the range or 1.5× the interquartile range, whichever is less. Two-sided hypothesis testing, not corrected for multiple testing, was performed using linear regression models. h, Replication of germline MBD4 and BRCA2 PTV associations with somatic CpG mutagenesis in TCGA whole-exome sequencing donors. Violin plots show the estimated density of the proportion of somatic CpG mutations in TCGA exomes with germline MBD4 and BRCA2 PTVs. The box denotes the interquartile range, with the median marked as a white point. The whiskers extend as far as the range or 1.5× the interquartile range, whichever is less. Two-sided hypothesis testing, not corrected for multiple testing, was performed using linear-regression models. i, Correlation between MBD4 expression and somatic CpG mutagenesis in primary solid PCAWG tumours. Hypothesis testing was two-sided and not corrected for multiple testing, using linear-regression models. The box denotes the interquartile range, with the median marked as a horizontal line. The whiskers extend as far as the range or 1.5× the interquartile range, whichever is less. j, Data are mean ± s.e.m. across n = 20 tumour types. The dashed black line shows the fitted line to the data, estimated using linear-regression models. Hypothesis testing was two-sided and not corrected for multiple testing, using Spearman’s rank correlations. k, MBD4 effect sizes (open circles) with 95% confidence intervals (error bars) for individual cancer types were estimated using linear-regression analysis after (if available) accounting for sex, age at diagnosis (young/old) and ICGC project. Hypothesis testing was two-sided and not corrected for multiple testing.
Extended Data Fig. 12 Germline MEI call set.
a, Left, dots show the number of transductions promoted by each hot element in individual samples. Arrows highlight retrotransposition burst. Right, the contribution of each hot locus is represented. The total number of transductions mediated by each source element is shown on the right. b, Source L1 activity rate (that is, measured as the average number of transductions mediated by an element) versus the percentage of samples with retrotransposition activity in which the germline element is active. For visualization purposes, extreme points observed for a source L1 with an activity rate of 49 and for a L1 active in 31% of the samples are shown at ≥20 and ≥10, respectively. c, Contrasting allele frequencies for Strombolian and Plinian source loci (sample sizes shown under each axis label). The box denotes the interquartile range, with the median marked as a white point. The whiskers extend as far as the range or 1.5× the interquartile range, whichever is less. Hypothesis testing was performed using two-sided Mann–Whitney U-tests without correction for multiple tests. d, Numbers of active and hot source L1 elements per donor. Data are mean ± s.d. number of elements per donor. e, The novel Plinian source element on 7p12.3 mediates 72 transductions among only 6 cancer samples. This generates a transduction that induces the deletion of the tumour-suppressor gene CDKN2A. f, Violin plots show the estimated number of distinct germline MEI alleles per PCAWG donor. The box denotes the interquartile range, with the median marked as a white point. The whiskers extend as far as the range or 1.5× the interquartile range, whichever is less. Donors are grouped according to their genetic ancestry: AFR, African; AMR, admixed American; EAS, East Asian; EUR, European; SAS, South Asian. Sample sizes are shown under each axis label. g, For each type of MEI (L1, Alu and SVA) identified both in PCAWG and in the 1000 Genomes Project (1KGP), the correlations between allele frequency estimates per ancestry derived from both projects are displayed in a blue (0) to red (1) coloured gradient. n = 2,583 PCAWG patients. Two-sided hypothesis testing was performed using Spearman’s rank correlations without correction for multiple tests. h, Example correlation between MEI allele frequencies derived from PCAWG and the 1000 Genomes Project for individuals with European ancestry (n = 1,201 patients in PCAWG). Two-sided hypothesis testing was performed using Spearman’s rank correlations without correction for multiple tests. i, Evaluation of TraFiC-mem false-discovery rate on a liver hepatocellular carcinoma sample (DO50807) and a cell line (NCI-BL2087) sequenced using single-molecule sequencing with MinION (Oxford Nanopore). For each allele frequency bin (common, >5%; low frequency, 1–5%; rare, <1%), the percentage of events supported by N long reads is represented (N ranges from 0–1 to more than 5). MEIs supported by at least two Nanopore reads were considered to be true positives (blue palette) and were classified as false positives (red) otherwise. The total number of germline MEIs per allele frequency bin is shown on the right. j, Correlation between predicted MEI lengths from Illumina and Nanopore data. Two-sided hypothesis testing was performed using Spearman’s rank correlations without correction for multiple testing.
Extended Data Fig. 13 Different mechanisms of telomere lengthening in cancer.
a, Scatter plot showing the four clusters of tumour-specific telomere patterns identified across PCAWG samples, together with the clusters of matched normal samples, generated by t-distributed stochastic neighbour embedding. Circles represent tumour samples and triangles represent matched normal samples. Points are coloured by tissue of origin. Data are based on n = 2,518 tumour samples and their matched normal samples. b, Patterns of comutation of the relevant driver mutations across individual patients. Columns in plot represent individual patients, coloured by type of abnormality observed. c, Distribution of clonality of driver mutations in genes relevant to telomere maintenance across clusters. Clonal [early], clonal mutations that occurred before duplications involving the relevant chromosome (including whole-genome duplications); clonal [late], clonal mutations that occurred after such duplications; and clonal [NA], mutations that occurred when no duplication was observed. d, Relationship between the estimated number of stem cell divisions per year and rate of telomere maintenance abnormalities across tumour types. The analysis uses data on estimated rates of stem cell division per year across n = 19 tissue types previously collated from the literature82. Tumour types are coloured according to the scheme shown in Extended Data Fig. 3. Two-sided hypothesis testing was performed using likelihood ratio tests on Poisson regression models with no correction for multiple tests.
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-19, Supplementary Methods and Supplementary Notes 1-6
Supplementary Tables
This zipped file contains Supplementary Tables 1-21 and a Supplementary Table Guide
Supplementary Information
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
The ICGC/TCGA Pan-Cancer Analysis of Whole Genomes Consortium. Pan-cancer analysis of whole genomes. Nature 578, 82–93 (2020). https://doi.org/10.1038/s41586-020-1969-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-1969-6
This article is cited by
-
Identification and verification of PTPN3 as a novel biomarker in predicting cancer prognosis, immunity, and immunotherapeutic efficacy
European Journal of Medical Research (2024)
-
Evolutionary dependency of cancer mutations in gene pairs inferred by nonsynonymous-synonymous mutation ratios
Genome Medicine (2024)
-
Large-scale copy number alterations are enriched for synthetic viability in BRCA1/BRCA2 tumors
Genome Medicine (2024)
-
Unveiling the unique role of TSPAN7 across tumors: a pan-cancer study incorporating retrospective clinical research and bioinformatic analysis
Biology Direct (2024)
-
GPX8+ cancer-associated fibroblast, as a cancer-promoting factor in lung adenocarcinoma, is related to the immunosuppressive microenvironment
BMC Medical Genomics (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.