Abstract
In the adult hippocampus, synapses are constantly formed and eliminated1,2. However, the exact function of synapse elimination in the adult brain, and how it is regulated, are largely unknown. Here we show that astrocytic phagocytosis3 is important for maintaining proper hippocampal synaptic connectivity and plasticity. By using fluorescent phagocytosis reporters, we find that excitatory and inhibitory synapses are eliminated by glial phagocytosis in the CA1 region of the adult mouse hippocampus. Unexpectedly, we found that astrocytes have a major role in the neuronal activity-dependent elimination of excitatory synapses. Furthermore, mice in which astrocytes lack the phagocytic receptor MEGF10 show a reduction in the elimination of excitatory synapses; as a result, excessive but functionally impaired synapses accumulate. Finally, Megf10-knockout mice show defective long-term synaptic plasticity and impaired formation of hippocampal memories. Together, our data provide strong evidence that astrocytes eliminate unnecessary excitatory synaptic connections in the adult hippocampus through MEGF10, and that this astrocytic function is crucial for maintaining circuit connectivity and thereby supporting cognitive function.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All primary antibodies used in this study are listed in the Methods. Recipes for reporters used in this study are provided in the Methods. All data are available upon reasonable request. For further inquiries, please contact the corresponding author. Source data are provided with this paper.
Code availability
All codes used are available upon reasonable request.
References
Toni, N., Buchs, P. A., Nikonenko, I., Bron, C. R. & Muller, D. LTP promotes formation of multiple spine synapses between a single axon terminal and a dendrite. Nature 402, 421–425 (1999).
Attardo, A., Fitzgerald, J. E. & Schnitzer, M. J. Impermanence of dendritic spines in live adult CA1 hippocampus. Nature 523, 592–596 (2015).
Chung, W. S. et al. Astrocytes mediate synapse elimination through MEGF10 and MERTK pathways. Nature 504, 394–400 (2013).
Maletic-Savatic, M., Malinow, R. & Svoboda, K. Rapid dendritic morphogenesis in CA1 hippocampal dendrites induced by synaptic activity. Science 283, 1923–1927 (1999).
Okabe, S., Kim, H. D., Miwa, A., Kuriu, T. & Okado, H. Continual remodeling of postsynaptic density and its regulation by synaptic activity. Nat. Neurosci. 2, 804–811 (1999).
Xu, T. et al. Rapid formation and selective stabilization of synapses for enduring motor memories. Nature 462, 915–919 (2009).
Stevens, B. et al. The classical complement cascade mediates CNS synapse elimination. Cell 131, 1164–1178 (2007).
Schafer, D. P. et al. Microglia sculpt postnatal neural circuits in an activity and complement-dependent manner. Neuron 74, 691–705 (2012).
Paolicelli, R. C. et al. Synaptic pruning by microglia is necessary for normal brain development. Science 333, 1456–1458 (2011).
Leeman, D. S. et al. Lysosome activation clears aggregates and enhances quiescent neural stem cell activation during aging. Science 359, 1277–1283 (2018).
Doherty, G. P., Bailey, K. & Lewis, P. J. Stage-specific fluorescence intensity of GFP and mCherry during sporulation in Bacillus subtilis. BMC Res. Notes 3, 303 (2010).
Wang, C. et al. Microglia mediate forgetting via complement-dependent synaptic elimination. Science 367, 688–694 (2020).
Hong, S. et al. Complement and microglia mediate early synapse loss in Alzheimer mouse models. Science 352, 712–716 (2016).
Steward, O., Torre, E. R., Tomasulo, R. & Lothman, E. Neuronal activity up-regulates astroglial gene expression. Proc. Natl Acad. Sci. USA 88, 6819–6823 (1991).
Srinivasan, R. et al. New transgenic mouse lines for selectively targeting astrocytes and studying calcium signals in astrocyte processes in situ and in vivo. Neuron 92, 1181–1195 (2016).
Howard, C. V. & Reed, M. G. Unbiased Stereology: Three-Dimensional Measurement in Microscopy 1st edition, 69–105 (Springer, 1999)
Elmore, M. R. et al. Colony-stimulating factor 1 receptor signaling is necessary for microglia viability, unmasking a microglia progenitor cell in the adult brain. Neuron 82, 380–397 (2014).
Rice, R. A. et al. Elimination of microglia improves functional outcomes following extensive neuronal loss in the hippocampus. J. Neurosci. 35, 9977–9989 (2015).
Rhee, J.-S. et al. Augmenting neurotransmitter release by enhancing the apparent Ca2+ affinity of synaptotagmin 1. Proc. Natl Acad. Sci. USA 102, 18664–18669 (2005).
Schneggenburger, R., Meyer, A. C. & Neher, E. Released fraction and total size of a pool of immediately available transmitter quanta at a calyx synapse. Neuron 23, 399–409 (1999).
Elmqvist, D. & Quastel, D. M. J. A quantitative study of end-plate potentials in isolated human muscle. J. Physiol. (Lond.) 178, 505–529 (1965).
Caroni, P., Donato, F. & Muller, D. Structural plasticity upon learning: regulation and functions. Nat. Rev. Neurosci. 13, 478–490 (2012).
Kandel, E. R. The molecular biology of memory storage: a dialogue between genes and synapses. Science 294, 1030–1038 (2001).
Vogel-Ciernia, A. & Wood, M. A. Examining object location and object recognition memory in mice. Curr. Protoc. Neurosci. 69, 1–17 (2014).
Tonegawa, S., Morrissey, M. D. & Kitamura, T. The role of engram cells in the systems consolidation of memory. Nat. Rev. Neurosci. 19, 485–498 (2018).
Skelton, P. D., Frazel, P. W., Lee, D., Suh, H. & Luikart, B. W. Pten loss results in inappropriate excitatory connectivity. Mol. Psychiatry 24, 1627–1640 (2019).
Guang, S. et al. Synaptopathology involved in autism spectrum disorder. Front. Cell. Neurosci. 12, 470 (2018).
Fischer, D., Bieber, T., Li, Y., Elsässer, H. P. & Kissel, T. A novel non-viral vector for DNA delivery based on low molecular weight, branched polyethylenimine: effect of molecular weight on transfection efficiency and cytotoxicity. Pharm. Res. 16, 1273–1279 (1999).
Guo, P. et al. A simplified purification method for AAV variant by polyethylene glycol aqueous two-phase partitioning. Bioengineered 4, 103–106 (2013).
Gibson, D. G. et al. Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat. Methods 6, 343–345 (2009).
Choi, J. H. et al. Optimization of AAV expression cassettes to improve packaging capacity and transgene expression in neurons. Mol. Brain 7, 17 (2014).
Gilles, J. F., Dos Santos, M., Boudier, T., Bolte, S. & Heck, N. DiAna, an ImageJ tool for object-based 3D co-localization and distance analysis. Methods 115, 55–64 (2017).
Bloss, E. B., Cembrowski, M. S., Karsh, B., Fetter, R. D. & Spruston, N. Single excitatory axons form clustered synapses onto CA1 pyramidal cell dendrites. Nat. Neurosci. 21, 353–363 (2018).
Vandael, D., Borges-Merjane, C., Zhang, X. & Jonas, P. Short-term plasticity at hippocampal mossy fiber synapses is induced by natural activity patterns and associated with vesicle pool engram formation. Neuron 107, 509–521.e7 (2020).
Rosenmund, C. & Stevens, C. F. Definition of the readily releasable pool of vesicles at hippocampal synapses. Neuron 16, 1197–1207 (1996).
Acknowledgements
We thank all members of the Chung, Mun, and Park laboratories for helpful discussions, and C. Cho for reading the paper. This work was supported by grants from the Samsung Science & Technology Foundation (SSTF-BA1701-18, W.-S.C.), the National Research Foundation of Korea (NRF) (2019R1A2C1010634 (J.Y.M.), 2016M3C7A1905391), and the KBRI basic research program (20-BR-01-04 (H.P.), 20-BR-01-09 (J.Y.M)) funded by the Ministry of Science and ICT. J.-H.L is partly supported by the Global PhD Fellowship Program through the NRF funded by the Ministry of Education (2017H1A2A1042287). Instruments (SEM and TEM) and whole-cell patch clamp data were acquired at the Brain Research Core Facilities in KBRI. Imaris software was supported by Bio Core facilities in KAIST.
Author information
Authors and Affiliations
Contributions
W.-S.C. and H.P. designed projects. J.-H.L. designed DNA constructs used in this paper, produced AAV-based reporters, performed all imaging and western blot experiments, and analysed data. J.-Y.K. performed and analysed electrophysiology and behavioral experiments. H.L. performed and analysed electrophysiology experiments. J.-H.L. and S.N. prepared brain samples for TEM and SEM experiments. S.N. and J.Y.M. performed and analysed TEM, ssTEM and SEM experiments. S.Y.L. performed stereotaxic surgeries and analysed the phagocytic capacity of aged astrocytes. W.-S.C. and H.P. supervised the project and wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature thanks Marc Freeman, Jeremy Kay and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
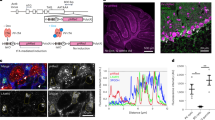
Extended Data Fig. 1 Development of the mCherry-eGFP reporter system for monitoring glial phagocytosis.
a, Schematic diagram of the mCherry-eGFP reporter system for monitoring glial phagocytosis of neuronal material. b, AAV9::hSyn-lyn-mCherry-eGFP was injected into hippocampal CA3 of 4-week-old mice. c, d, Representative confocal single-plane images of CA3 (c) and CA1 (d) of AAV9::hSyn-lyn-mCherry-eGFP injected mice. The white box in d highlights mCherry-alone puncta region. Scale bars = 100μm (left and middle) and 10μm (right). e, Distribution of mCherry-alone puncta in CA1 of AAV9::hSyn-lyn-mCherry-eGFP injected mice. n = 8 individual experiments from 3 mice for each group. Data are mean ± s.e.m. Astrocytes versus microglia, **P = 0.0019. Mann–Whitney test. f, Representative confocal single-plane images of the primary visual cortex (V1) of the mice injected with AAV9::hSyn-lyn-mCherry-eGFP into the lateral geniculate nucleus (LGN) which projects their axons to V1. g, Representative confocal single-plane images of CA1 of the mice injected with AAV9::hSyn-lyn-mCherry-eGFP into CA3. Scale bars = 10 μm (f, g). h, Quantification of the area of mCherry-alone puncta in V1 and CA1. n = 16 individual experiments from 4 mice per each group. CA1 versus V1, ****P < 0.0001. For all quantified data, Mann–Whitney test. All data are mean ± s.e.m.
Extended Data Fig. 2 AAV-based synapse phagocytosis reporters accurately incorporate into excitatory and inhibitory synapses.
a, c, e, g, Representative confocal single-plane images of the CA1 hippocampus injected with AAV9::ExPre (a), AAV9::InhiPre (c), AAV9::ExPost (e) or AAV9::InhiPost (g) reporters. Synaptic proteins (a: VGluT1, c: VGAT, e: PSD95, g: Gephyrin, cyan) with mCherry-alone (red) and mCherry-eGFP (yellow) (upper left). mCherry-alone and mCherry-eGFP without synaptic proteins (upper right). White dotted circles highlight the mCherry-alone puncta. Zoom-in panels of the upper left panels (lower left). White arrowheads in boxes indicate the mCherry-eGFP puncta co-localized with synaptic proteins. Zoom-in panels of the upper left panels without mCherry-eGFP puncta (lower right). Scale bars = 10 μm (Histogram) Fluorescence trajectory in white boxes showing co-localization of mCherry-eGFP and synaptic proteins. b, d, f, h, Representative single-plane confocal images of mCherry-alone puncta (red) and synaptic proteins (b: VGluT1. d: VGAT, f: PSD95, h: Gephyrin, white). White lines indicate the synaptic protein puncta co-localized with mCherry-alone puncta.
Extended Data Fig. 3 Cellular localization of mCherry-eGFP and mCherry-alone puncta inside of glial cells.
a, b, Representative 3D-rendered images showing ExPre-derived mCherry-alone puncta (red) and mCherry-eGFP synaptic puncta (green) associated with astrocytes (a, white) or microglia (b, white). Scale bars = 10 μm. c, f, Representative confocal single plane images showing co-localization of ExPre-derived mCherry-alone (red) or mCherry-eGFP (yellow) puncta with Cathepsin D-positive lysosomes (c, blue) or Rab5-positive endosomes (f, blue) in astrocytes (S100B, white). White lines mark the surface of astrocytes. In divided images, red and yellow dotted circles mark mCherry-alone and mCherry-eGFP puncta, respectively. White arrows indicate co-localized puncta with Cathepsin D-positive lysosomes (c) or Rab5-positive endosomes (f). Scale bars = 10 μm. d, e, g, h, Percentages of ExPre-derived mCherry-eGFP (d, g) or mCherry-alone (e, h) in either inside or outside of Cathepsin D-positive lysosomes (d, e) or Rab5-positive endosomes (g, h) in astrocytes. n = 3 mice per each group.
Extended Data Fig. 4 mCherry-alone puncta derived from glial synapse elimination reporters.
a, b, Representative confocal z projection images of glial synapse elimination reporters (mCherry (red) and eGFP (green) derived from ExPre, InhiPre, ExPost and InhiPost), lysosome marker Cathepsin D (blue) and glial cell markers (a, S100B for astrocytes. b, IBA1 for microglia, white). Scale bars = 10 μm. White dotted circles mark mCherry-alone puncta. White arrows indicate mCherry-alone puncta co-localized with Cathepsin D inside either astrocytes (a) or microglia (b). c, Quantification of individual synaptic reporter puncta size (mCherry-eGFP puncta, area). n = 12 individual experiments per each group. d, Representative confocal single plane images of CA1 showing mCherry-eGFP (green) and mCherry-alone (red) puncta derived from each reporters. Scale bars = 5 μm. e, f, Detailed procedures explaining how the area of engulfed synapses (mCherry-alone) by astrocytes (e, n = 12, 10, 12, 12) or microglia (f, n = 10, 10, 10, 12) was normalized. For description, please see the Method section. For all comparisons, statistics were not applied in these graphs. For all data, each n represent individual experiment from 3 mice. All data are mean ± s.e.m.
Extended Data Fig. 5 Both astrocytes and microglia engulf more excitatory than inhibitory synapses.
a, b, Representative confocal single plane images of CA1 astrocytes (a, S100B, red) or microglia (b, IBA1, red) with synaptic proteins (VGluT1, PSD95, VGAT or Gephyrin, green). White arrows indicate engulfed synaptic proteins in glial cells. Scale bars = 5 μm. c, d, Quantification of engulfed synaptic proteins per unit astrocytes (c) or microglia (d). n = 9 individual experiments from 3 mice per each groups. (c) Excitatory Pre versus Inhibitory Pre/Post, ****P < 0.0001. Excitatory Pre versus Excitatory Post, ***P = 0.0004. Excitatory Post versus Inhibitory Pre/Post, **P = 0.0015/0.0048. (d) Excitatory Pre versus Inhibitory Pre/Post, ****P < 0.0001. Excitatory Pre versus Excitatory Post, ***P = 0.0006. Excitatory Post versus Inhibitory Post, **P = 0.0016. One-way ANOVA followed by Tukey’s multiple comparisons. e, Comparison between the quantified engulfed synapses by astrocytes and microglia. Excitatory Pre, *P = 0.0151. Excitatory Post, *P = 0.0256. Inhibitory Pre, *P = 0.0142. Inhibitory Post, *P = 0.0428. Mann–Whitney test. n = 3 mice per group. All data are mean ± s.e.m.
Extended Data Fig. 6 AAV-based synapse phagocytosis reporters do not induce changes in synaptic properties or reactive gliosis.
a, Schematic diagram of experimental schedule for measuring sEPSCs and sIPSCs of reporter-expressing neurons. b–d, Representative images that depict experimental and control groups. (b) AAV9::ExPre and AAV9::ExPost were injected into CA3 and CA1, respectively. (c) PBS was injected into CA3 and CA1. (d) AAV9::InhiPre and AAV9::InhiPost were injected into CA1. e, f, Quantifications of mean frequencies (e) or amplitudes (f) of sEPSC/sIPSCs from ExPre+ExPost injected (b; sEPSC: 39 cells from 4 mice, sIPSC: 36 cells from 4 mice), PBS injected (c; sEPSC: 46 cells from 4 mice, sIPSC: 38 cells from 4 mice) or InhiPre+InhiPost injected (d; sEPSC: 37 cells from 4 mice, sIPSC: 29 cells from 4 mice) groups. g, Quantifications of mean voltage thresholds for action potential initiation (AP threshold) from each group. ExPre+ExPost injected: 46 cells from 2 mice, PBS injected: 36 cells from 3 mice, InhiPre+InhiPost injected: 51 cells from 3 mice. n.s. h, Quantifications of mean number of action potential induced by current injection in each group. ExPre+ExPost injected: 46 cells from 2 mice, PBS injected: 36 cells from 3 mice, InhiPre+InhiPost injected: 51 cells from 3 mice. One-way ANOVA followed by Tukey’s multiple comparisons test (e–h). i, AAV9::ExPre was injected into CA3 unilaterally. After 4 weeks of recovery, ipsilateral CA3, CA1 (Ipsi-CA3, Ipsi-CA1) and contralateral CA3, CA1 (Contra-CA3, Contra-CA1) were examined. j, Representative confocal single plane images of GFAP expression (red) in Ipsi-CA1 (upper) or Contra-CA1 (lower) of AAV9::Expre-CA3 injected brains. k, l, Quantification of GFAP immunoreactivity comparing Ipsi- and Contra-CA1 (k) or CA3 (l). n = 9 individual experiments from 3 mice per each groups. Mann–Whitney test. m, Representative confocal single plane images of CD68 (green) expression with microglia (IBA1, red) in Ipsi-CA1 (upper) or Contra-CA1 (lower) of AAV9::Expre-CA3 injected brains. n, o, Quantification of microglial CD68 area comparing Ipsi- and Contra-CA1 (n) or CA3 (o). n = 9 individual experiments from 3mice per each groups. Mann–Whitney test. p, AAV9::InhiPre was injected into CA1 unilaterally. After 4 weeks of recovery, ipsilateral CA1 (Ipsi-CA1) and contralateral CA1 (Contra-CA1) were examined. q, Representative confocal single plane images of GFAP expression (red) in Ipsi-CA1 (upper) or Contra-CA1 (lower) of AAV9::Inhipre-CA1 injected brains. r, Quantification of GFAP immunoreactivity comparing Ipsi- and Contra-CA1. n = 9 individual experiments from 3 mice per each groups. Mann–Whitney test. s, Representative images of CD68 (green) expression with microglia (IBA1, red) in Ipsi-CA1 (upper) or Contra-CA1 (lower) of AAV9::Inhipre-CA1 injected brains. t, Quantification of microglial CD68 area comparing Ipsi- and Contra-CA1. n = 9 individual experiments from 3 mice per each groups. Mann–Whitney test. u, Quantification of Cathepsin D-positive lysosome area in astrocytes of control (Ctrl), AAV9::ExPre, InhiPre, ExPost and InhiPost injected brain. n = 9 individual experiments from 3 mice per each groups. One-way ANOVA followed by Tukey’s multiple comparisons. v, Schematic diagram of experimental schedule for comparing the amount of engulfed excitatory pre-synapses by CA1 astrocytes in 3- and 9-month-old mice. w, Representative confocal single plane images of astrocytes (S100B, red) containing mCherry-alone puncta (green) in CA1 of 3- and 9-month-old mice. x, Quantification of the area of mCherry-alone puncta in astrocytes normalized with the area of mCherry-eGFP and astrocytes in CA1 of ExPre injected 3- and 9-month-old mice. n = 21, 23 from 4 mice for each group. Mann–Whitney test. For all analyses, n.s., not significant. Scale bars = 10 μm. For all comparisons, n.s., not significant. All data are mean ± s.e.m.
Extended Data Fig. 7 Hippocampal activity regulates astrocytic synapse elimination.
a, Representative confocal z projection images of CA1 neurons (NeuN, blue) with c-fos (green) from Ctrl, EE2d and EE7d mice. Scale bar = 50μm. b, Quantification of c-fos+ and NeuN+ neurons in CA1 from control, EE2d and EE7d mice, n = 6 individual experiments from 3 mice per each group. Ctrl versus EE2d, **P = 0.0046, Ctrl versus EE7d, ***P = 0.0002. One-way ANOVA followed by Tukey’s multiple comparisons test. c, Representative confocal z projection images of dentate gyrus (DG) cells (DAPI, red pseudo-colored) with newborn cells (DCX, green) from Ctrl, EE2d and EE7d mice. Scale bar = 10 μm. d, Quantification of DCX+ cells in DG from Ctrl, EE2d and EE7d mice, n = 6 individual experiments from 3 mice per each group. Ctrl or EE2d versus EE7d, ****P < 0.0001. One-way ANOVA followed by Tukey’s multiple comparisons test. e, f, Representative confocal single plane images of astrocytes (e, red) or microglia (f, red) containing mCherry-alone puncta (green) in CA1 of reporter-injected Ctrl and EE7d mice. Scale bars = 10 μm. g, A representative confocal z projection of an axon terminal labelled with ExPre reporter (synaptic boutons, yellow) and tagBFP (axon, blue) in CA1 of reporter-electroporated Ctrl mice. White dotted circles and arrows highlight mCherry-alone puncta (red) outside (engulfed) or inside (recycled) of tagBFP-positive axon terminal. Scale bar = 5 μm. h, Quantification of the number of mCherry-alone puncta outside (engulfed) or inside (recycled) of tagBFP-positive axon terminal in Ctrl or EE7d groups. n = 9 individual experiments from 5 mice per each group. EE7d engulfed versus Ctrl recycled or engulfed, ****P < 0.0001. EE7d engulfed versus EE7d recycled, **P = 0.0020. Two-way ANOVA followed by Tukey’s multiple comparisons test. All data are mean ± s.e.m.
Extended Data Fig. 8 Astrocytes regulate nearby synapses by MEGF10 while microglia do not participate in homeostatic synapse elimination.
a, Representative images of MEGF10 (green) with astrocytes (S100B, red) in control (upper) and acMegf10 KO (lower) mice. Scale bars = 20 μm. White arrowheads indicate co-localization of MEGF10 and astrocytes. b, western blotting of hippocampal homogenates from Ctrl, acMegf10 KO and straight Megf10 KO mice. c, Representative confocal z projection images of astrocytes (S100B, green) in CA1 of Ctrl and acMegf10 KO mice. Scale bars = 60 μm. d, e, Quantification of the number (d) and territory area (e) of astrocytes in CA1 of Ctrl and acMegf10 KO mice. n = 17 individual experiments from 3 mice per each group. n.s., not significant. Mann–Whitney test. f, Schematic illustrations that depict the experimental and control groups. To examine potential effects in the number of excitatory synapses or glial phagocytosis, 4-hydroxytamoxifen or tamoxifen was consecutively injected into Megf10fl/fl mice without Aldh1l1-creERT2 allele for 5 times, once per a day (Tam group). For control, vehicle (see Method) was consecutively injected into Megf10fl/fl mice with Aldh1l1-creERT2 allele for 5 times, once per a day (Vehi group). g, h, Quantification of the area of Cathepsin D-positive lysosomes in astrocytes (g) or microglial CD68 (h) of Tam and Vehi groups. n = 10 individual experiments from 3 mice per each groups. P > 0.05. n.s., not significant. Mann–Whitney test. i, Quantification of the number of VGLUT1- and PSD95-double positive (left), VGLUT1-positive pre- (middle), and PSD95-positive post- (right) excitatory synapses in hippocampal CA1 of Tam and Vehi groups. n = 12 individual experiments from 3 mice per each group. For all comparisons, P > 0.05. n.s., not significant. Mann–Whitney test. j, k, Quantification of the area of mCherry-alone puncta in microglia normalized with the area of mCherry-eGFP and microglia in CA1 of AAV9::ExPre (j) or AAV9::ExPost (k) injected Ctrl and acMegf10 KO mice. n = 12 individual experiments from 4 mice for each group. n.s., not significant. Mann–Whitney test. l, To acquire mosaic Megf10 KO in CA1 astrocytes, low titer of AAV5-GFAP-Cre-eGFP (haMegf10 KO) or AAV5-GFAP-eGFP (Ctrl) was injected into Megf10-floxed CA1. m, Representative confocal single plane images of eGFP positive (blue-colored eGFP and red-colored S100B) and negative astrocytes (S100B only), along with MEGF10 staining (green). In haMegf10 KO, only eGFP-positive astrocytes that express Cre fail to express MEGF10. White dotted lines mark astrocytic territories. Scale bars = 10 μm. n–p, Quantification of the number of VGLUT1- and PSD95-double positive (n), VGLUT1-positive pre- (o), and PSD95-positive post- (p) excitatory synapses in the eGFP-positive and negative astrocytic territories of Ctrl and haMegf10 KO groups. n = 16 individual experiments from 4 mice per each group. (n) eGFP-positive of haMegf10 KO versus eGFP-positive/negative of Ctrl, ***P = 0.0004/0.0009. eGFP-positive of haMegf10 KO versus eGFP-negative of haMegf10 KO, **P = 0.0039. (o) eGFP-positive of haMegf10 KO versus eGFP-negative of Ctrl, ***P = 0.0006. eGFP-positive of haMegf10 KO versus eGFP-negative of haMegf10 KO, ****P < 0.0001. eGFP-positive of haMegf10 KO versus eGFP-positive of Ctrl, **P = 0.0013. (n) eGFP-positive of haMegf10 KO versus eGFP-positive/negative of Ctrl, ****P < 0.0001. eGFP-positive of haMegf10 KO versus eGFP-negative of haMegf10 KO, ***P = 0.0001. Two-way ANOVA followed by Tukey’s multiple comparisons. q, Schematic diagram of experimental schedule. r, Representative confocal z projection images of hippocampal microglia (IBA1, red) with astrocyte (S100B, green) and DAPI staining (blue), comparing control (Ctrl) and PLX3397/5622 treated (Microglia-depleted) groups. Scale bar = 100 μm. s, Quantification of the number of VGLUT1- and PSD95-double positive (left), VGLUT1-positive pre- (middle), and PSD95-positive post- (right) excitatory synapses in hippocampal CA1 of control and PLX3397/5622 treated mice, n = 18 individual experiments from 3 mice per each group. For all comparisons, P > 0.05. n.s., not significant. Mann–Whitney test. All data are mean ± s.e.m.
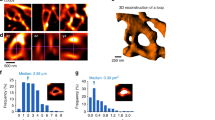
Extended Data Fig. 9 3-dimensional (3D) reconstruction of dendrites and astrocytes engulfing spine synapses.
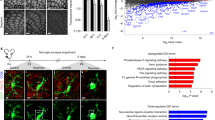
a–f, Representative single-plane SEM images (a, d) and 3D reconstruction of Ctrl astrocytes which are engulfing spine synapses (b, c, e, f). Scale bars = 500 nm. Red arrows indicate pre-synaptic vesicles inside of astrocytic acidic organelles. Dotted lines of 3D reconstructions highlight the synapses being engulfed and acidic organelles in astrocytes. g, Schematic diagram of experimental schedule. h, Representative images of dendrites and dendritic spines reconstructed by 3D tracing technique from Ctrl (upper), and 3-week-acMegf10 KO (lower) CA1 hippocampus.
Extended Data Fig. 10 Synaptic properties and basal locomotive activities affected by astrocytic Megf10 KO.
a, Representative mEPSC traces recorded from control (Ctrl) or haMegf10 KO slices. b, c, Scatter plots showing average mEPSC frequency (b) or amplitude (c). Control (Ctrl): n = 16 cells from 4 mice. haMegf10 KO: n = 11 cells from 4 mice. n.s., not significant. *P = 0.0242, unpaired Student t-test. d, Representative recording traces showing overlaid AMPAR-(red) or NMDAR-dependent evoked EPSC responses. e, Summary of AMPAR/NMDAR EPSC ratio (AMPA/NMDA ratio) from control (Ctrl) or haMegf10 KO slices. Control (Ctrl): n = 10 cells from 4 mice. haMegf10 KO: n = 7 cells from 3 mice. n.s., not significant. Unpaired Student t-test. f, Schematic diagram representing feed-forward inhibition circuits in the hippocampal CA1 area. g, Representative evoked EPSC or IPSC traces recorded from the same CA1 neuron. h, Bar graphs depict average EPSC/IPSC ratio from control (Ctrl) or haMegf10 KO slices. Control (Ctrl): n = 10 cells from 6 mice. haMegf10 KO: n = 9 cells from 6 mice. n.s., not significant. Unpaired Student t-test. i, Representative 3D-reconstructed EM images showing distributions of total synaptic vesicles in the hippocampus of control (Ctrl) or acMegf10 KO mice. Presynaptic bouton (cyan), postsynaptic spine (scarlet), postsynaptic density (purple) and synaptic vesicles (green). Scale bar = 500nm. j, Bar graphs depict average total vesicle numbers per μm3. Control (Ctrl): n = 109 boutons from 3 mice. acMegf10 KO: n = 114 boutons from 3 mice. ***P < 0.0001, unpaired student t-test. k, Representative cumulative EPSC responses recorded from CA1 neurons in control (blue) or haMegf10 KO (orange), in response to 20 Hz for 5 s stimulation. l, Plot showing cumulative evoked EPSC (eEPSC) amplitudes during 20 Hz stimulation. Data points in a range of 4 – 5 s were fitted by linear regression, and back-extrapolated (solid line) to time 0 to estimate the RRP size (SMN method). m, Bar graphs depicting summary of estimated RRP size by the SMN method. Ctrl (blue): n = 10 cells from 3 mice; haMegf10 KO (orange): n = 8 cells from 3 mice. *P = 0.0181, unpaired Student’s t-test. n, Plot showing eEPSC amplitudes against the amplitude of the cumulative eEPSC in Ctrl or haMegf10 KO mice. Plots were linearly fitted from the sixth to the twentieth cumulative eEPSCs and back-extrapolated linear fits to the x axis to estimate the RRP (EQ method). o, Bar graphs depicting summary of estimated RRP size by the EQ method. Ctrl (blue): n = 10 cells from 3 mice; haMegf10 KO (orange): n = 8 cells from 3 mice. *P = 0.0026, unpaired Student’s t-test. p, Summary of input/output relationships of fEPSP recorded from Ctrl or haMegf10 KO hippocampal slices, n = 10 and 11 slices in Ctrl and haMegf10 KO, respectively. *P = 0.0431 (150 μA), *P = 0.0203 (200 μA). Unpaired student’s t-test. q, r, Representative traces showing mouse-travelling in the open field chamber during the NOR habituation phase in Ctrl (q) and haMegf10 KO mice (r). s–u, Bar graphs depict total distance (s), mean speed (t), percentage of time in central zone (u) of Ctrl and haMegf10 KO mice, n = 19 and 20 mice in Ctrl and haMegf10 KO, respectively. No statistical significance was found. Unpaired Student’s t-test. v, w, Representative traces showing mouse-travelling in the open field chamber during the NOL habituation phase in Ctrl (v) and haMegf10 KO mice (w). x–z, Bar graphs depict total distance (x), mean speed (y), percentage of time in central zone (z) of Ctrl and haMegf10 KO mice, n = 12 and 13 mice in Ctrl and haMegf10 KO, respectively. No statistical significance was found. Unpaired Student’s t-test. All data are mean ± s.e.m.
Supplementary information
Supplementary Table
This file contains Supplementary Table 1: Electrophysiological properties of recorded CA1 neurons.
Supplementary Data
This file contains raw blot data for extended data figure 9b.
Source data
Rights and permissions
About this article
Cite this article
Lee, JH., Kim, Jy., Noh, S. et al. Astrocytes phagocytose adult hippocampal synapses for circuit homeostasis. Nature 590, 612–617 (2021). https://doi.org/10.1038/s41586-020-03060-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-03060-3
This article is cited by
-
Brain stars take the lead during critical periods of early postnatal brain development: relevance of astrocytes in health and mental disorders
Molecular Psychiatry (2024)
-
Astrocytes in the adult dentate gyrus—balance between adult and developmental tasks
Molecular Psychiatry (2024)
-
Targeting synapse function and loss for treatment of neurodegenerative diseases
Nature Reviews Drug Discovery (2024)
-
Lifelong absence of microglia alters hippocampal glutamatergic networks but not synapse and spine density
EMBO Reports (2024)
-
Ogt-mediated O-GlcNAcylation inhibits astrocytes activation through modulating NF-κB signaling pathway
Journal of Neuroinflammation (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.