Abstract
Much of the American Arctic was first settled 5,000 years ago, by groups of people known as Palaeo-Eskimos. They were subsequently joined and largely displaced around 1,000 years ago by ancestors of the present-day Inuit and Yup’ik1,2,3. The genetic relationship between Palaeo-Eskimos and Native American, Inuit, Yup’ik and Aleut populations remains uncertain4,5,6. Here we present genomic data for 48 ancient individuals from Chukotka, East Siberia, the Aleutian Islands, Alaska, and the Canadian Arctic. We co-analyse these data with data from present-day Alaskan Iñupiat and West Siberian populations and published genomes. Using methods based on rare-allele and haplotype sharing, as well as established techniques4,7,8,9, we show that Palaeo-Eskimo-related ancestry is ubiquitous among people who speak Na-Dene and Eskimo–Aleut languages. We develop a comprehensive model for the Holocene peopling events of Chukotka and North America, and show that Na-Dene-speaking peoples, people of the Aleutian Islands, and Yup’ik and Inuit across the Arctic region all share ancestry from a single Palaeo-Eskimo-related Siberian source.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
Raw sequence data (.bam files) from the 48 ancient individuals that we studied here are available from the European Nucleotide Archive under accession number PRJEB30575. The genotype data for the Iñupiat were obtained through informed consent, which does not allow us to provide the data through public or controlled-access data repositories; it also does not allow analyses of phenotypic traits, or commercial use of the data. To protect the privacy of participants and ensure that their wishes with respect to data usage are followed, researchers wishing to use data from the Iñupiat samples should contact M.G.H. (ghayes@northwestern.edu) and D.A.B. (deborah.bolnick@uconn.edu), who can then arrange to share the data with researchers who can affirm that they will abide by the relevant conditions through a signed data-sharing agreement. The SNP genotyping data for West Siberians (Enets, Kets, Nganasans, and Selkups) are publicly available at the Edmond database, under the permalink https://doi.org/10.17617/3.1z.
Code availability
Custom code used in this manuscript is available at dedicated GitHub repositories: rarecoal (https://github.com/stschiff/rarecoal), rarecoal-tools (https://github.com/stschiff/rarecoal-tools), and RAS-tools (https://github.com/TCLamnidis/RAStools).
References
Rasmussen, M. et al. Ancient human genome sequence of an extinct Palaeo-Eskimo. Nature 463, 757–762 (2010).
Raghavan, M. et al. The genetic prehistory of the New World Arctic. Science 345, 1255832 (2014).
Friesen, T. M. in The Oxford Handbook of the Prehistoric Arctic (eds Friesen, T. M. & Mason, O. K.) 673–692 (Oxford Univ. Press, New York, 2016).
Reich, D. et al. Reconstructing Native American population history. Nature 488, 370–374 (2012).
Raghavan, M. et al. Genomic evidence for the Pleistocene and recent population history of Native Americans. Science 349, aab3884 (2015).
Moreno-Mayar, J. V. et al. Terminal Pleistocene Alaskan genome reveals first founding population of Native Americans. Nature 553, 203–207 (2018).
Haak, W. et al. Massive migration from the steppe was a source for Indo-European languages in Europe. Nature 522, 207–211 (2015).
Patterson, N. et al. Ancient admixture in human history. Genetics 192, 1065–1093 (2012).
Schiffels, S. et al. Iron Age and Anglo-Saxon genomes from East England reveal British migration history. Nat. Commun. 7, 10408 (2016).
Skoglund, P. et al. Genetic evidence for two founding populations of the Americas. Nature 525, 104–108 (2015).
Moreno-Mayar, J. V. et al. Early human dispersals within the Americas. Science 362, eaav2621 (2018).
Posth, C. et al. Reconstructing the deep population history of Central and South America. Cell 175, 1185–1197.e22 (2018).
Potter, B. A. et al. Early colonization of Beringia and Northern North America: chronology, routes, and adaptive strategies. Quat. Int. 444, 36–55 (2017).
Llamas, B. et al. Ancient mitochondrial DNA provides high-resolution time scale of the peopling of the Americas. Sci. Adv. 2, e1501385 (2016).
Raff, J. A., Rzhetskaya, M., Tackney, J. & Hayes, M. G. Mitochondrial diversity of Iñupiat people from the Alaskan North Slope provides evidence for the origins of the Paleo- and Neo-Eskimo peoples. Am. J. Phys. Anthropol. 157, 603–614 (2015).
Friesen, T. M. On the naming of Arctic archaeological traditions: the case for Paleo-Inuit. Arctic 68, iii–iv (2015).
Park, R. W. in The Oxford Handbook of the Prehistoric Arctic (eds Friesen, T. M. & Mason, O. K.) 417–442 (Oxford Univ. Press, New York, 2016).
Prentiss, A. M., Walsh, M. J., Foor, T. A. & Barnett, K. D. Cultural macroevolution among high latitude hunter–gatherers: a phylogenetic study of the Arctic Small Tool tradition. J. Archaeol. Sci. 59, 64–79 (2015).
Tremayne, A. H. & Rasic, J. T. in The Oxford Handbook of the Prehistoric Arctic (eds Friesen, T. M. & Mason, O. K.) 303–322 (Oxford Univ. Press, New York, 2016).
Friesen, T. M. Contemporaneity of Dorset and Thule cultures in the North American Arctic: new radiocarbon dates from Victoria Island, Nunavut. Curr. Anthropol. 45, 685–691 (2004).
Dabney, J. et al. Complete mitochondrial genome sequence of a Middle Pleistocene cave bear reconstructed from ultrashort DNA fragments. Proc. Natl Acad. Sci. USA 110, 15758–15763 (2013).
Rohland, N., Harney, E., Mallick, S., Nordenfelt, S. & Reich, D. Partial uracil-DNA-glycosylase treatment for screening of ancient DNA. Phil. Trans. R. Soc. Lond. B 370, 20130624 (2015).
Fu, Q. et al. An early modern human from Romania with a recent Neanderthal ancestor. Nature 524, 216–219 (2015).
Bardill, J. et al. Advancing the ethics of paleogenomics. Science 360, 384–385 (2018).
Lawson, D. J., Hellenthal, G., Myers, S. & Falush, D. Inference of population structure using dense haplotype data. PLoS Genet. 8, e1002453 (2012).
Hellenthal, G. et al. A genetic atlas of human admixture history. Science 343, 747–751 (2014).
Smith, S. E. et al. Inferring population continuity versus replacement with aDNA: a cautionary tale from the Aleutian Islands. Human Biol. 81 407–426 (2009).
Scally, A. & Durbin, R. Revising the human mutation rate: implications for understanding human evolution. Nat. Rev. Genet. 13, 745–753 (2012).
Fenner, J. N. Cross-cultural estimation of the human generation interval for use in genetics-based population divergence studies. Am. J. Phys. Anthropol. 128, 415–423 (2005).
Kari, J. in The Dene-Yeniseian Connection, Anthropological Papers of the University of Alaska: New Series, vol. 5 (eds Kari, J. & Potter, B. A.) 194–222 (Univ. of Alaska and Alaska Native Language Centre, Fairbanks, Alaska, 2010).
Reynolds, A. W. et al. Comparing signals of natural selection between three Indigenous North American populations. Proc. Natl Acad. Sci. USA 116, 9312–9317.
Flegontov, P. et al. Genomic study of the Ket: a Paleo-Eskimo-related ethnic group with significant ancient North Eurasian ancestry. Sci. Rep. 6, 20768 (2016).
Mallick, S. et al. The Simons Genome Diversity Project: 300 genomes from 142 diverse populations. Nature 538, 201–206 (2016).
Li, H. & Durbin, R. Inference of human population history from individual whole-genome sequences. Nature 475, 493–496 (2011).
Rasmussen, M. et al. The genome of a Late Pleistocene human from a Clovis burial site in western Montana. Nature 506, 225–229 (2014).
Raghavan, M. et al. Upper Palaeolithic Siberian genome reveals dual ancestry of Native Americans. Nature 505, 87–91 (2014).
Lazaridis, I. et al. Ancient human genomes suggest three ancestral populations for present-day Europeans. Nature 513, 409–413 (2014).
O’Connell, J. et al. A general approach for haplotype phasing across the full spectrum of relatedness. PLoS Genet. 10, e1004234 (2014).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
Alexander, D. H., Novembre, J. & Lange, K. Fast model-based estimation of ancestry in unrelated individuals. Genome Res. 19, 1655–1664 (2009).
Loh, P. R. et al. Inferring admixture histories of human populations using linkage disequilibrium. Genetics 193, 1233–1254 (2013).
Lazaridis, I. et al. Genomic insights into the origin of farming in the ancient Near East. Nature 536, 419–424 (2016).
Verdu, P. et al. Patterns of admixture and population structure in native populations of Northwest North America. PLoS Genet. 10, e1004530 (2014).
Acknowledgements
We acknowledge the ancient people whose skeletal samples were studied, the Aleut Corporation, the Aleutians Pribilof Islands Association, and the Chaluka Corporation for granting permissions to conduct genetic analyses on the eastern Aleutians. We thank the staff at the Smithsonian Institution’s National Museum of Natural History for facilitating the sample collection; the McGrath Native Village Council and MTNT Ltd for granting permissions to conduct genetic analyses on the Tochak McGrath remains; J. Clark, who performed biological age estimates on these remains; the research participants in Alaska (Genetics of Alaskan North Slope (GeANS) project funded by NSF OPP-0732857) and West Siberia who donated samples for genome-wide analysis; J. B. Coltrain for sharing data on stable isotopes; and J. W. Ives, J. Tackney, L. Norman, and K. TallBear for comments on earlier drafts of this paper. Sample collection and the initial molecular, isotopic, and accelerator mass spectrometry (AMS) 14C dating of the samples described here were funded by National Science Foundation Office of Polar Program grants OPP-9726126, OPP-9974623, and OPP-0327641; the Natural Sciences and Engineering Research Council of Canada; and the Wenner-Gren Foundation for Anthropological Research (6364). This work was supported by the Czech Ministry of Education, Youth and Sports from the project ‘IT4Innovations National Supercomputing Center – LM2015070’. P.F., P.C., O.F., and N.E.A. were supported by the Institutional Development Program of the University of Ostrava; P.F. and P.C. were supported by the EU Operational Programme ‘Research and Development for Innovations’ (CZ.1.05/2.1.00/19.0388) and P.C. was also supported by the Statutory City of Ostrava (0924/2016/ŠaS) and the Moravian-Silesian Region (01211/2016/RRC); P.S. was funded by the Francis Crick Institute, which receives its core funding from Cancer Research UK (FC001595), the UK Medical Research Council (FC001595), and the Wellcome Trust (FC001595); D.R. was funded by NSF HOMINID (grant BCS-1032255), NIH (NIGMS), the Allen Discovery Center of the Paul Allen Foundation (grant GM100233), and is an Investigator of the Howard Hughes Medical Institute; D.A.B. was supported by a Norman Hackerman Advanced Research Program grant from the Texas Higher Education Coordinating Board; AMS 14C work at Pennsylvania State University by D.J.K. and B.J.C was funded by the NSF Archaeometry programme (BCS-1460369); and C.J., T.C.L., J.K., and S.S. were supported by the Max Planck Society.
Reviewer information
Nature thanks Carles Lalueza-Fox, John Lindo and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
S.S., P.F., and D.R. supervised the study. B.A.P., T.M.F., A.M.K., R.A.S., S.V., E.V., D.H.O’R., R.P., and D.R. assembled the collection of archaeological samples. D.A.B., O.F., J.R., M.G.H., and J.K. assembled the sample collection from present-day populations. T.K.H., D.J.K., B.J.C., and T.M.F. were responsible for radiocarbon dating and calibration. N.R., N.A., N.B., F.C., D.K., A.M.L., J.O., and K.S. performed laboratory work and supervised ancient DNA sequencing. P.F., N.E.A., P.C., S.M., C.J., T.C.L., I.O., P.S., and S.S., analysed genetic data. E.J.V. wrote the supplemental section on linguistics. P.F., D.R., and S.S. wrote the manuscript with additional input from all other co-authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
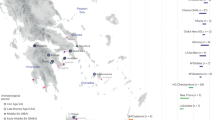
Extended Data Fig. 1 Geographic locations of Siberian and North American populations used in this study.
The three main datasets are as follows (Supplementary Tables 4, 5): (1) a set based on the Affymetrix Human Origins genotyping array, including alternatively pseudo-haploid or diploid genotypes for the ancient Saqqaq individual1; diploid genotypes for the ancient Clovis35 individual, together with 1240K SNP capture pseudo-haploid data from 6 ancient Aleuts who had the highest coverage; 2 unrelated ancient Athabaskans; 19 ancient Old Bering Sea individuals from the Ekven and Uelen sites; the Middle Dorset and Late Dorset Palaeo-Eskimo individuals; and the ancient Ust’-Belaya Angara population of 9 individuals (Supplementary Table 1); (2) a set based on various Illumina arrays, including Saqqaq and the other ancient samples; and (3) a whole-genome dataset of 190 individuals from 87 populations, including the Saqqaq individual, 1 ancient Athabaskan individual (I5319), and 1 ancient Aleut individual (I0719), for whom we generated complete genomes with 6.1× and 2.3× coverage, respectively (Supplementary Table 1). The dataset composition—that is, the number of individuals in each meta-population—is shown in the table on the right. Locations of individuals with whole-genome sequencing data (SEQ) are shown with circles, and those of Illumina (ILL) and HumanOrigins (HO) SNP-array samples with triangles and diamonds, respectively. Meta-populations are colour-coded in a similar way throughout all figures and designated as follows: Na-Dene speakers (abbreviated as ATH), other northern Native Americans (NAM) (alternatively known as Northern First Peoples), Southern First Peoples (SAM), Ancient Beringians (BER), Eskimo–Aleut speakers (E-A), Chukotko-Kamchatkan speakers (C-K), Palaeo-Eskimos (P-E), West and East Siberians (WSIB and ESIB), Southeast Asians (SEA), Europeans (EUR), and Africans (AFR). The locations of the Saqqaq, Dorset, and other ancient individuals are shown as stars that are coloured to reflect their meta-population affiliation.
Extended Data Fig. 2 PCA based on the Illumina dataset.
A plot of two principal components (PC1 versus PC2) calculated using PLINK2 is shown (linkage disequilibrium pruning was not applied). No outliers were excluded for this analysis, which was based on 642 individuals and 524,830 loci. The following meta-populations most relevant for our study are plotted: present-day Eskimo–Aleut and Chukotko-Kamchatkan speakers, ancient Chukotkan Neo-Eskimos (Ekven and Uelen sites), ancient Aleuts, Palaeo-Eskimos (the Saqqaq, Middle Dorset, and Late Dorset individuals), ancient Northern Athabaskans, present-day Na-Dene speakers, Northern and Southern First Peoples, West and East Siberians, the Ust’-Belaya Angara ancient Siberian group, Southeast Asians, and Europeans. Radiocarbon dates in cal. yr bp are shown for ancient samples. For individuals, 95% confidence intervals are shown, and for groups of individuals, minimal and maximal median dates among individuals are shown.
Extended Data Fig. 3 Ancestry proportions in American, Chukotkan, and Kamchatkan populations.
a–j, The HumanOrigins (a–e) and Illumina (f–j) datasets without transition polymorphisms are shown. Five alternative outgroup sets are indicated below the plots and described in detail in the Methods and Supplementary Information section 5. Bold formatting denotes ancient target groups. Saqqaq (pseudo-haploid genotype calls) was considered as a Palaeo-Eskimo source for all populations apart from Saqqaq itself (for which Late Dorset was used as a source) and alternative First Peoples sources were as follows: Mixe, Guarani, or Karitiana for the HumanOrigins dataset; Nisga’a, Mixtec, Pima, or Karitiana for the Illumina dataset. To visualize both systematic and statistical errors, ancestry proportions inferred by qpAdm and their standard errors are shown for all triplets including these different First Peoples sources, or for many alternative target groups in the case of Southern First Peoples (single standard error intervals are plotted here). Asterisks indicate ancestry proportions greater than 150% (inappropriate models). Meta-populations are colour-coded according to the legend and abbreviated as before (N-D, Na-Dene speakers). Target group sizes in the HumanOrigins dataset ranged from 1 to 23 individuals (average 5.6), and in the Illumina dataset they ranged from 1 to 16 individuals (average 5.1).
Extended Data Fig. 4 Ancestry proportions in American, Chukotkan, and Kamchatkan populations.
a–j, Similar analysis as in Extended Data Fig. 3, but including transition polymorphisms. Target population sizes in the HumanOrigins dataset (a–e) ranged from 1–23 individuals (average 5.6), and in the Illumina dataset (f–j) they ranged from 1–16 individuals (average 5.1).
Extended Data Fig. 5 Relative Saqqaq, Arctic, and European haplotype-sharing statistics for American individuals.
a, b, Results are shown for the Human Origins (a) and Illumina (b) datasets, normalized using the African meta-population. Both Eskimo–Aleut- and Chukotko-Kamchatkan-speaking groups contributed to the Arctic HSS. The same statistics and statistics with other normalizers are shown in the form of two-dimensional plots in Supplementary Information section 6. Two Dakelh (Northern Athabaskan) individuals with whole-genome sequencing data5 were included in both datasets and are marked with asterisks. The plots based on both datasets demonstrate that Na-Dene speakers have the highest relative Saqqaq HSS. One Haida and three Splatsin individuals also demonstrate outlying Saqqaq HSSs (b); however, these individuals contrast with a majority of non-Na-Dene-speaking Northern First Peoples, and Palaeo-Eskimo ancestry in these individuals may be explained by recent interaction with Na-Dene speakers living in close proximity43. The Haida outlier demonstrates the maximal Arctic HSS among all First Peoples, and their Arctic ancestry has contributed to their elevated Saqqaq HSS. Saqqaq, Arctic, and European statistics are largely uncorrelated in First Peoples: Pearson’s correlation coefficients for Saqqaq versus Arctic relative HSSs are 0.56 among all First Peoples and 0.64 among Northern First Peoples in the case of the Illumina dataset, and 0.66 and 0.72, respectively, in the case of the HumanOrigins dataset. h.s., haplotype sharing.
Extended Data Fig. 6 Rare-allele-sharing analysis.
A two-dimensional plot of Chukotko-Kamchatkan and Siberian rare-allele-sharing statistics for First Peoples, Na-Dene-speaking, Eskimo–Aleut-speaking, and Palaeo-Eskimo individuals. Rare alleles occurring from 2 to 5 times in the reference set of 238 haploid genomes (0.8–2.1% frequency) contributed to the statistics; the Chukchi individual was dropped from the Chukotko-Kamchatkan reference group, and the transversion-only dataset was used. Thus, this analysis was based on 918,474 loci. The sample size for this analysis equals 238 + 2 haploid genomes in a target individual, as individuals were analysed separately. Standard deviations were calculated using a jackknife approach, with chromosomes used as resampling blocks. Single standard error intervals and means are plotted. Populations and meta-populations are colour-coded according to the legend. Rare-allele-sharing statistics for simulated mixtures of any present-day southern Native American individual and the Saqqaq individual (from 5–75% Saqqaq ancestry, with 5% increments) are plotted as semi-transparent pink circles. Plots for the 2–10 allele frequency range and other versions are shown in Supplementary Information section 8.
Extended Data Fig. 7 An admixture graph connecting various modern meta-populations and ancient populations or individuals.
The graph (see Supplementary Information section 10) features a simplified three-component model for Europeans as previously suggested37, and two gene flows from a European lineage related to the ancient Siberian genome MA-136 into Native Americans and Siberians. The topology within the PPE clade was obtained by cycling through dozens of trees with all possible topologies of branches and admixture edges, and selecting the one with the highest support and no zero-length edges within the PPE clade.
Extended Data Fig. 8 ADMIXTURE analysis.
a, b, Results are shown for the HumanOrigins (a) and Illumina (b) SNP-array datasets. The number of source populations in ADMIXTURE is 14 and 11 for the 2 datasets, respectively. One hundred iterations were calculated for each value of K from 5–20 (in which K is the number of ancestral populations), and the optimal K values were selected based on 10-fold cross-validation. Contributions from hypothetical ancestral populations are colour-coded, and meta-populations used in this study are indicated above the plot (abbreviations as before). Chipewyan or Northern Athabaskan and Tlingit individuals with European admixture are plotted in separate bars, as are ancient individuals: Clovis, Northern Athabaskans, Aleuts, Chukotkan Neo-Eskimos (Ekven and Uelen sites), Saqqaq and Late Dorset Palaeo-Eskimos, and a genetically heterogeneous Ust’-Belaya Angara Siberian population (Ust’-Belaya WSIB, an individual I7760 who has a West Siberian genetic profile according to PCA and this ADMIXTURE analysis; Ust’-Belaya, the remaining eight individuals from the Ust’-Belaya Angara site who have a distinct genetic profile according to our PCA analysis). Outliers, including individuals admixed with Europeans and East Asians, were not removed from Na-Dene-speaking populations in the Illumina dataset (b) to preserve their maximal diversity. Outliers were removed for the purpose of other analyses that rely on pre-defined populations (for example, qpAdm and f4-statistics).
Extended Data Fig. 9 Clustering trees of individuals, computed by fineSTRUCTURE.
a, b, The trees are based on co-ancestry matrices of counts of shared haplotypes. Reduced versions of the HumanOrigins (a) and Illumina (b) SNP-array datasets were used (Supplementary Table 5), including only the following meta-populations that were most relevant for our study: Eskimo–Aleut speakers, Chukotko-Kamchatkan speakers, Na-Dene speakers, Northern First Peoples, Southern First Peoples, West Siberians, East Siberians, Southeast Asians, and Europeans. Meta-population affiliation is colour-coded for individuals. Iñupiat individuals genotyped in this study are marked with a blue line. The two Dakelh (Northern Athabaskan) individuals with sequenced genomes are also indicated, as well as the ancient individuals—Clovis within the Southern First Peoples clade and Saqqaq within the Chukotko-Kamchatkan clade. Most members of each clade belong to the meta-populations indicated, with a few exceptions. First (a), Altaians fall into the ESIB clade, some Chilote fall into the NAM, and Aleuts fall into the WSIB clades (the two latter cases might be explained by extensive European ancestry in Chilote and in Aleuts (Extended Data Fig. 8a), which drives this clustering). Second (b), some Selkups fall into the ESIB clade, all four Southern Athabaskan speakers cluster with South Americans (reflecting their substantial South American ancestry (Extended Data Fig. 8b)), one Haida individual clusters with Na-Dene speakers, and five Northern Athabaskan speakers cluster with other Northern First Peoples.
Supplementary information
Supplementary Information
This file contains Supplementary Information Sections 1-13 and a Supplementary Discussion. Supplementary Figures are included within the respective sections. Additional tables referred to from within sections 4, 5, 10 and 12 are provided as separate Excel files named Supplementary Data 1 (containing Supplementary Tables for section 4), Supplementary Data 2 (containing Supplementary Tables for section 5), Supplementary Data 3 (containing Supplementary Tables for section10) and Supplementary Data 4 (containing Supplementary Tables for section 12).
Supplementary Table 1
Summary of genome-wide data from 48 newly reported ancient samples. A table summarising all newly reported ancient samples, including age estimates and sampling location.
Supplementary Table 2
Reservoir-adjusted radiocarbon calibrations and stable isotope data for ancient skeletal samples analyzed in this study. A table summarizing radiocarbon measurements and calibrations, as well as data from stable isotope analysis.
Supplementary Table 3
Information on newly genotyped present-day individuals. A table summarizing the newly genotyped West Siberian and Alaskan individuals, alongside gender, population and sampling location.
Supplementary Table 4
Composition of the genomic and SNP array datasets used in this study. A table summarizing the composition of datasets from publicly available and new data used for analyses in this study.
Supplementary Table 5
Details of datasets used in this study. A summary of key statistics for datasets used for different analyses, including the percentage of missing data and the number of sites.
Supplementary Table 6
Analysis results showing that the SNP panels used in this study provide sufficient power to distinguish Native American populations from each other. In order to test whether the datasets used in this study allow detecting substructure in the First Peoples and American Arctic populations, we divided each American population consisting of 2 or more individuals into two halves (equal, if possible) randomly and show that the following f4-statistics: (Americani Half A, Americanj; Americani Half B, Dai) are significantly different from zero in more than 90% of the cases.
Supplementary Data 1
This file contains Supplementary Tables related to Section 4 of the Supplementary Information.
Supplementary Data 2
This file contains Supplementary Tables related to Section 5 of the Supplementary Information.
Supplementary Data 3
This file contains Supplementary Tables related to Section 10 of the Supplementary Information.
Supplementary Data 4
This file contains Supplementary Tables related to Section 12 of the Supplementary Information.
Rights and permissions
About this article
Cite this article
Flegontov, P., Altınışık, N.E., Changmai, P. et al. Palaeo-Eskimo genetic ancestry and the peopling of Chukotka and North America. Nature 570, 236–240 (2019). https://doi.org/10.1038/s41586-019-1251-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-1251-y
This article is cited by
-
Differentiated genomic footprints suggest isolation and long-distance migration of Hmong-Mien populations
BMC Biology (2024)
-
The Allen Ancient DNA Resource (AADR) a curated compendium of ancient human genomes
Scientific Data (2024)
-
Medieval DNA from Soqotra points to Eurasian origins of an isolated population at the crossroads of Africa and Arabia
Nature Ecology & Evolution (2024)
-
Biological anthropology must reassess museum collections for a more ethical future
Nature Ecology & Evolution (2023)
-
Mitogenomics of the Koryaks and Evens of the northern coast of the Sea of Okhotsk
Journal of Human Genetics (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.