Abstract
A universal gain-of-function approach for selective and temporal control of protein activity in living systems is crucial to understanding dynamic cellular processes. Here we report development of a computationally aided and genetically encoded proximal decaging (hereafter, CAGE-prox) strategy that enables time-resolved activation of a broad range of proteins in living cells and mice. Temporal blockage of protein activity was computationally designed and realized by genetic incorporation of a photo-caged amino acid in proximity to the functional site of the protein, which can be rapidly removed upon decaging, resulting in protein re-activation. We demonstrate the wide applicability of our method on diverse protein families, which enabled orthogonal tuning of cell signalling and immune responses, temporal profiling of proteolytic substrates upon caspase activation as well as the development of protein-based pro-drug therapy. We envision that CAGE-prox will open opportunities for the gain-of-function study of proteins and dynamic biological processes with high precision and temporal resolution.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All relevant data are included in the Article or its Supplementary Information. More details are available from the corresponding authors upon request.
Code availability
All relevant code is available on GitHub at https://github.com/wendao/CAGE-prox. All other code is available from the corresponding authors upon reasonable request.
References
Bishop, A. C. et al. A chemical switch for inhibitor-sensitive alleles of any protein kinase. Nature 407, 395–401 (2000).
Baud, M. G. J. et al. A bump-and-hole approach to engineer controlled selectivity of BET bromodomain chemical probes. Science 346, 638–641 (2014).
Winter, G. E. et al. Drug development. Phthalimide conjugation as a strategy for in vivo target protein degradation. Science 348, 1376–1381 (2015).
Ulman, V. et al. An objective comparison of cell-tracking algorithms. Nature Methods 14, 1141–1152 (2017).
Clift, D. et al. A method for the acute and rapid degradation of endogenous proteins. Cell 171, 1692–1706.e18 (2017).
Dagliyan, O. et al. Engineering extrinsic disorder to control protein activity in living cells. Science 354, 1441–1444 (2016).
Davis, K. M., Pattanayak, V., Thompson, D. B., Zuris, J. A. & Liu, D. R. Small molecule-triggered Cas9 protein with improved genome-editing specificity. Nature Chem. Biol. 11, 316–318 (2015).
Wu, Y. I. et al. A genetically encoded photoactivatable Rac controls the motility of living cells. Nature 461, 104–108 (2009).
Karginov, A. V., Ding, F., Kota, P., Dokholyan, N. V. & Hahn, K. M. Engineered allosteric activation of kinases in living cells. Nature Biotechnol. 28, 743–747 (2010).
Lungu, O. I. et al. Designing photoswitchable peptides using the AsLOV2 domain. Chem. Biol. 19, 507–517 (2012).
Niopek, D., Wehler, P., Roensch, J., Eils, R. & Di Ventura, B. Optogenetic control of nuclear protein export. Nature Commun. 7, 10624 (2016).
Pratt, M. R., Schwartz, E. C. & Muir, T. W. Small-molecule-mediated rescue of protein function by an inducible proteolytic shunt. Proc. Natl Acad. Sci. USA 104, 11209–11214 (2007).
Taslimi, A. et al. An optimized optogenetic clustering tool for probing protein interaction and function. Nature Commun. 5, 4925 (2014).
Wang, H. et al. LOVTRAP: an optogenetic system for photoinduced protein dissociation. Nature Methods 13, 755–758 (2016).
Lee, S. et al. Reversible protein inactivation by optogenetic trapping in cells. Nature Methods 11, 633–636 (2014).
Zhou, X. X., Fan, L. Z., Li, P., Shen, K. & Lin, M. Z. Optical control of cell signaling by single-chain photoswitchable kinases. Science 355, 836–842 (2017).
Toettcher, J. E., Weiner, O. D. & Lim, W. A. Using optogenetics to interrogate the dynamic control of signal transmission by the Ras/Erk module. Cell 155, 1422–1434 (2013).
Guntas, G. et al. Engineering an improved light-induced dimer (iLID) for controlling the localization and activity of signaling proteins. Proc. Natl Acad. Sci. USA 112, 112–117 (2015).
Li, J. & Chen, P. R. Development and application of bond cleavage reactions in bioorthogonal chemistry. Nature Chem. Biol. 12, 129–137 (2016).
Deiters, A., Groff, D., Ryu, Y., Xie, J. & Schultz, P. G. A genetically encoded photocaged tyrosine. Angew. Chem. Int. Edn Engl. 45, 2728–2731 (2006).
Arbely, E., Torres-Kolbus, J., Deiters, A. & Chin, J. W. Photocontrol of tyrosine phosphorylation in mammalian cells via genetic encoding of photocaged tyrosine. J. Am. Chem. Soc. 134, 11912–11915 (2012).
Hartshorn, M. J. et al. Diverse, high-quality test set for the validation of protein-ligand docking performance. J. Med. Chem. 50, 726–741 (2007).
Park, H. et al. Simultaneous optimization of biomolecular energy function on features from small molecules and macromolecules. J. Chem. Theory Comput. 12, 6201–6212 (2016).
Simanshu, D. K., Nissley, D. V. & McCormick, F. RAS proteins and their regulators in human disease. Cell 170, 17–33 (2017).
Cox, A. D., Fesik, S. W., Kimmelman, A. C., Luo, J. & Der, C. J. Drugging the undruggable RAS: mission possible? Nature Rev. Drug Discov. 13, 828–851 (2014).
Zhao, B. S., Roundtree, I. A. & He, C. Post-transcriptional gene regulation by mRNA modifications. Nature Rev. Mol. Cell Biol. 18, 31–42 (2017).
Tomabechi, Y. et al. Crystal structure of nanoKAZ: The mutated 19 kDa component of Oplophorus luciferase catalyzing the bioluminescent reaction with coelenterazine. Biochem. Biophys. Res. Commun. 470, 88–93 (2016).
Zhang, G. et al. Bioorthogonal chemical activation of kinases in living systems. ACS Cent. Sci. 2, 325–331 (2016).
Holohan, C., Van Schaeybroeck, S., Longley, D. B. & Johnston, P. G. Cancer drug resistance: an evolving paradigm. Nature Rev. Cancer 13, 714–726 (2013).
Diaz, J. E. et al. A Split–Abl kinase for direct activation in cells. Cell Chem. Biol. 24, 1250–1258.e4 (2017).
Zheng, S., Fan, X., Wang, J., Zhao, J. & Chen, P. R. Dissection of kinase isoforms through orthogonal and chemical inducible signaling cascades. ChemBioChem 18, 1593–1598 (2017).
Chang, L. & Karin, M. Mammalian MAP kinase signalling cascades. Nature 410, 37–40 (2001).
Albeck, J. G. et al. Quantitative analysis of pathways controlling extrinsic apoptosis in single cells. Mol. Cell 30, 11–25 (2008).
Riedl, S. J. & Shi, Y. Molecular mechanisms of caspase regulation during apoptosis. Nature Rev. Mol. Cell Biol. 5, 897–907 (2004).
Mahrus, S. et al. Global sequencing of proteolytic cleavage sites in apoptosis by specific labeling of protein N termini. Cell 134, 866–876 (2008).
Agard, N. J. et al. Global kinetic analysis of proteolysis via quantitative targeted proteomics. Proc. Natl Acad. Sci. USA 109, 1913–1918 (2012).
Putt, K. S. et al. Small-molecule activation of procaspase-3 to caspase-3 as a personalized anticancer strategy. Nature Chem. Biol. 2, 543–550 (2006).
Liu, S. et al. Solid tumor therapy by selectively targeting stromal endothelial cells. Proc. Natl Acad. Sci. USA 113, E4079–E4087 (2016).
Shimbo, K. et al. Quantitative profiling of caspase-cleaved substrates reveals different drug-induced and cell-type patterns in apoptosis. Proc. Natl Acad. Sci. USA 109, 12432–12437 (2012).
Wang, Y. et al. Chemotherapy drugs induce pyroptosis through caspase-3 cleavage of a gasdermin. Nature 547, 99–103 (2017).
Miller, J. C., Silverman, S. K., England, P. M., Dougherty, D. A. & Lester, H. A. Flash decaging of tyrosine sidechains in an ion channel. Neuron 20, 619–624 (1998).
Kim, D. E., Chivian, D. & Baker, D. Protein structure prediction and analysis using the Robetta server. Nucleic Acids Res. 32, W526-31 (2004).
Meiler, J. & Baker, D. ROSETTALIGAND: protein-small molecule docking with full side-chain flexibility. Proteins 65, 538–548 (2006).
Chin, J. W. Expanding and reprogramming the genetic code. Nature 550, 53–60 (2017).
Li, J., Jia, S. & Chen, P. R. Diels-Alder reaction-triggered bioorthogonal protein decaging in living cells. Nature Chem. Biol. 10, 1003–1005 (2014).
Luo, J. et al. Genetically encoded optochemical probes for simultaneous fluorescence reporting and light activation of protein function with two-photon excitation. J. Am. Chem. Soc. 136, 15551–15558 (2014).
Trott, O. & Olson, A. J. AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J. Comput. Chem. 31, 455–461 (2010).
Salomon-Ferrer, R., Case, D. A. & Walker, R. C. An overview of the amber biomolecular simulation package. Wires. Comput. Mol. Sci. 3, 198–210 (2013).
Sousa da Silva, A. W. & Vranken, W. F. ACPYPE - AnteChamber Python parser interface. BMC Res. Notes 5, 367 (2012).
Jia, G. et al.; J. G et al. N6-methyladenosine in nuclear RNA is a major substrate of the obesity-associated FTO. Nature Chem. Biol. 7, 885–887 (2011).
Fu, Y. et al. FTO-mediated formation of N6-hydroxymethyladenosine and N6-formyladenosine in mammalian RNA. Nature Commun. 4, 1798 (2013).
Wallace, E. M., Lyssikatos, J. P., Yeh, T., Winkler, J. D. & Koch, K. Progress towards therapeutic small molecule MEK inhibitors for use in cancer therapy. Curr. Top. Med. Chem. 5, 215–229 (2005).
Duncia, J. V. et al. MEK inhibitors: the chemistry and biological activity of U0126, its analogs, and cyclization products. Bioorg. Med. Chem. Lett. 8, 2839–2844 (1998).
Yamaguchi, T., Kakefuda, R., Tajima, N., Sowa, Y. & Sakai, T. Antitumor activities of JTP-74057 (GSK1120212), a novel MEK1/2 inhibitor, on colorectal cancer cell lines in vitro and in vivo. Int. J. Oncol. 39, 23–31 (2011).
Dong, Q. et al. Discovery of TAK-733, a potent and selective MEK allosteric site inhibitor for the treatment of cancer. Bioorg. Med. Chem. Lett. 21, 1315–1319 (2011).
Hatzivassiliou, G. et al.; H. G et al. Mechanism of MEK inhibition determines efficacy in mutant KRAS- versus BRAF-driven cancers. Nature 501, 232–236 (2013).
Phadke, M. S., Sini, P. & Smalley, K. S. The novel ATP-competitive MEK/Aurora kinase inhibitor BI-847325 overcomes acquired BRAF inhibitor resistance through suppression of Mcl-1 and MEK expression. Mol. Cancer Ther. 14, 1354–1364 (2015).
Hoeflich, K. P. et al. Intermittent administration of MEK inhibitor GDC-0973 plus PI3K inhibitor GDC-0941 triggers robust apoptosis and tumor growth inhibition. Cancer Res. 72, 210–219 (2012).
Ohren, J. F. et al. Structures of human MAP kinase kinase 1 (MEK1) and MEK2 describe novel noncompetitive kinase inhibition. Nature Struct. Mol. Biol. 11, 1192–1197 (2004).
Whitehead, T. A. et al. Optimization of affinity, specificity and function of designed influenza inhibitors using deep sequencing. Nature Biotechnol. 30, 543–548 (2012).
Lakhani, S. A. et al. Caspases 3 and 7: key mediators of mitochondrial events of apoptosis. Science 311, 847–851 (2006).
Perinpanayagam, M. A. et al. Regulation of co- and post-translational myristoylation of proteins during apoptosis: interplay of N-myristoyltransferases and caspases. FASEB J. 27, 811–821 (2013).
Dix, M. M., Simon, G. M. & Cravatt, B. F. Global mapping of the topography and magnitude of proteolytic events in apoptosis. Cell 134, 679–691 (2008).
Dix, M. M. et al. Functional interplay between caspase cleavage and phosphorylation sculpts the apoptotic proteome. Cell 150, 426–440 (2012).
Tabb, D. L., McDonald, W. H. & Yates, J. R., III. DTASelect and Contrast: tools for assembling and comparing protein identifications from shotgun proteomics. J. Proteome Res. 1, 21–26 (2002).
Weerapana, E. et al. Quantitative reactivity profiling predicts functional cysteines in proteomes. Nature 468, 790–795 (2010).
Crawford, E. D. et al. The DegraBase: a database of proteolysis in healthy and apoptotic human cells. Mol. Cell. Proteomics 12, 813–824 (2013).
Wang, C. et al. A bead-based cleavage method for large-scale identification of protease substrates. Sci. Rep. 6, 22645 (2016).
Stoehr, G., Schaab, C., Graumann, J. & Mann, M. A SILAC-based approach identifies substrates of caspase-dependent cleavage upon TRAIL-induced apoptosis. Mol. Cell. Proteomics 12, 1436–1450 (2013).
Lüthi, A. U. & Martin, S. J. The CASBAH: a searchable database of caspase substrates. Cell Death Differ. 14, 641–650 (2007).
Young, L. C., Thulien, K. J., Campbell, M. R., Tron, V. A. & Andrew, S. E. DNA mismatch repair proteins promote apoptosis and suppress tumorigenesis in response to UVB irradiation: an in vivo study. Carcinogenesis 25, 1821–1827 (2004).
Nakashima, S. et al. Vacuolar H+-ATPase inhibitor induces apoptosis via lysosomal dysfunction in the human gastric cancer cell line MKN-1. J. Biochem. 134, 359–364 (2003).
Victor, K. G. et al. Proteomic identification of synaptic caspase substrates. Synapse 72, e22014 (2018).
Miller, C. J., Elliott, J. L. & Collier, R. J. Anthrax protective antigen: prepore-to-pore conversion. Biochemistry 38, 10432–10441 (1999).
Liu, S. et al. Matrix metalloproteinase-activated anthrax lethal toxin demonstrates high potency in targeting tumor vasculature. J. Biol. Chem. 283, 529–540 (2008).
Acknowledgements
We thank G. Jia, X. Zhang, W. Wang, Y. Wang and H. Song for providing the RNA oligonucleotide, expert technical assistance and helpful discussions. We thank the Computing Platform of the Center for Life Science for supporting the proteomic data analysis. This work was supported by research grants from the National Key Research and Development Program of China (2016YFA0501500 to P.R.C. and C.W.), the National Natural Science Foundation of China (21521003 to P.R.C. and C.W., 21740001 and 91753000 to P.R.C., 81490741 and 21778004 to C.W.), and a ‘Young 1000-Talent Plan’ Award to C.W.
Reviewer information
Nature thanks Martin Schnermann, Klaus Michael Hahn, Nanda Vikas and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
P.R.C. and C.W. conceived the study. J.W. conducted most of the experiments unless otherwise specified. Y.L. developed the computational pipeline and Y.-J.L. conducted the LF-based pro-drug experiments in mice. S.-Q.Z., J.-Y.Z. and G.Z. contributed to the biochemical experiments. X.W. contributed to the chemical synthesis. F.Y. contributed to the proteomic experiments. J.W., Y.L., Y-J.L., C.W. and P.R.C. wrote the paper with inputs from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Computational analysis and validation of Tyr as an optimal anchor residue.
a, Surveying the mutational stability of each of the 20 amino acids. The mutation stability of each residue was obtained by the in silico site-saturated mutagenesis of these 3,500 residues and energy calculation of each resulting mutant. Distribution of folding stability for each candidate amino acid is shown in the heat map. The blue and red colours represent the stable and deleterious mutations, respectively. b, Schematic of the validation workflow. According to our computational analysis and ranking, Tyr is a suitable anchor residue, whereas the introduced Lys mutations were more likely to affect the enzyme activity. To validate the computational results, a total of 30 randomly selected sites on each of the two model enzymes—FLuc and Nluc—was mutated to either Lys or Tyr, and the activities of the resulting mutants were systematically compared by measuring the resulting luminescence intensity. c, d, Results from tested mutations on FLuc and NLuc model proteins. Tyrosine mutations showed more stability in general than the lysine mutants. The randomly selected 30 mutation sites of FLuc: N84, F89, L194, N197, T214, R218, N229, H244, H245, F247, F250, Y255, S284, L286, E311, A313, Q338, Y340, I351, E354, V362, L411, L418, I434, R437, L441, Q448, T527, G528 and L530. The randomly selected 30 mutation sites of NLuc: F8, Q14, L20, V23, L24, S31, F33, Q34, P42, Q44, I46, I56, I58, V60, I62, I78, F79, V92, L94, Y96, I109, Y111, F112, Y116, V129, L133, R143, L151, F153 and I157. Results are the representative data of two biological replicates.
Extended Data Fig. 2 Optimizing the experimental and computational procedures of CAGE-prox.
a, Optimization of the UV irradiation time for protein activation in living cells. HEK293T cells expressing FLuc-Y255-ONBY were irradiated by UV light for different times and were further tested for FLuc activity. n = 3. b, Demonstration of the experimental variation from different batches. The expression level and photo-release efficiency are consistent in different batches of experiments. FLuc was used as an example. Mean ± s.d.; n = 3. c, Verification of the cell viability under the condition of ONBY incorporation. Mean ± s.d.; n = 3. d, e, The heterogeneity analysis of transient or stable transfection. FLuc-Y255-TAG was used as a model protein for evaluation and cells were stained using an anti-His tag antibody followed by an Alexa-Fluor-488-conjugated anti-mouse IgG. n = 2. The high transfection and expression efficiency were observed in HEK293T, HeLa and Jurkat cells (d) as well as in \({{\rm{ONBY\mbox{--}RS-tRNA}}}_{{\rm{CUA}}}^{\mathrm{pyl}}\) pair stable-expressed HEK293T cells expressing FLuc-Y255-TAG (e). f, Thermodynamic cycle of calculating protein stability (∆Gf) and binding affinity (∆∆Gb) as introduced by single point mutation. g, Equations for calculating protein stability (∆Gf) and binding affinity (∆∆Gb). The ∆Gf and ∆∆Gb of a given protein are approximated by Rosetta energy scores. h, Relative activity of the positive variants of FLuc before and after photo-decaging. Mean ± s.d.; n = 3; two-tailed t-test. i, Distribution of the distance (Å) and angular direction (degree) from the 60 anchor sites to the FLuc substrate. j, Distribution of the mutation stability (∆Gf) and substrate binding energy change (∆∆Gb) of all the 60 variants calculated by Rosetta. All above-mentioned samples are biological replicates. P values are shown in the figure.
Extended Data Fig. 3 Detailed workflow of the CAGE-prox strategy.
To prepare the initial structure for CAGE-prox calculation, an optimized protein–ligand complex structure was obtained from the PDB, and comparative modelling and/or ligand-docking were used when necessary. Geometry and energy parameters (D, A, ∆Gf and ∆∆Gb) were calculated by Rosetta and used as filters to exclude undesired sites. The sites that successfully passed the filters are considered as the ‘recommended sites’. In the next experimental validation step, ONBY was incorporated into the recommended sites followed by protein activity characterization with and without photo-activation.
Extended Data Fig. 4 CAGE-prox-enabled activation of KRAS and FTO.
a, Experimental validation of all the eight recommended anchor sites of KRAS by ERK phosphorylation upon photo-activation. n = 2. b, Fluorescence imaging of the subcellular localization of wild-type KRAS and KRAS-Y32-ONBY before and after photo-activation, which confirmed correct membrane location for these variants inside cells. Scale bars for all frames, 10 μm. n = 2. c, Structural view of the eight computationally recommended anchor sites of KRAS by CAGE-prox. The five positive anchor sites are labelled in orange. d, Schematic of the workflow for dissecting KRAS-mediated intracellular signal transduction. Both the KRAS activation-triggered signal transduction and EGF stimulation-triggered signal transduction were compared by monitoring the downstream phosphorylation responses. e, Upon temporal activation of KRAS, the phosphorylation levels of MEK1, ERK and JNK were analysed by immunoblotting. n = 2. f, Upon temporal stimulation by EGF, the phosphorylation levels of MEK1, ERK and JNK were analysed and compared by immunoblotting. Distinct patterns of downstream kinase phosphorylation were observed with CAGE-prox-enabled temporal KRAS activation as compared to EGF-stimulated samples. n = 2. g, Structural view of the nine computationally recommended anchor sites on FTO by CAGE-prox. h, Incorporation efficiency of ONBY in each of the recommended anchor sites of FTO as measured by immunoblotting. As the incorporation efficiency at the P93 and N205 sites is extremely low, they were excluded from further experimental validations. n = 2. i, The rest of seven FTO variants were validated by the LC–MS/MS-based RNA demethylation assay. Four variants (Y106, Y108, D233 and R322) exhibited desired demethylation activity on m6A upon photo-activation. n = 3. j, The activities of three positive FTO variants (Y106, Y108, and D233) were measured by a substrate demethylation assay. Mean ± s.d.; n = 3; two-tailed t-test. All above-mentioned samples are biological replicates. P values are shown in the figure.
Extended Data Fig. 5 CAGE-prox-enabled activation of NLuc, an enzyme with undefined active sites.
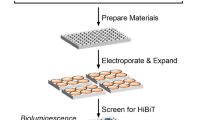
a, Structures of NLuc and its substrate. b, Energy distribution of NLuc complex models with the substrate docked. As NLuc does not have a defined active site, protein–ligand docking was performed to generate a total of 10,000 molecular docking models of the enzyme–substrate complexes. The computed parameters were shown in a scatter plot. x axis represents the interface area of each docking model by calculating the difference in solvent accessible surface areas between the apo state and the holo state. y axis represents the total energy of the docking complex calculated by Rosetta. The five docking models with either the best total energy or the largest difference in solvent-accessible surface area (∆SASA, plots labelled in orange) were selected for CAGE-prox calculation as potential anchor residues. c–g, Structural view of the five selected docking model with the substrate shown in purple sticks and the recommended CAGE-prox anchor residues in green sticks. h, Structural view of all the 15 recommended anchor sites (in green sticks) shown on the apo structure of NLuc. PDB code 5B0U. i, Experimental validation of all the 15 recommended CAGE-prox variants of NLuc. Three variants (marked in orange labels) showed strong luciferase activity (x axis) and high activation ratio (y axis) after photo-decaging. j, Validation of the recommended anchor sites on NLuc. Three out of fifteen recommended sites (S29, Y94 and Y114) were efficiently blocked by ONBY incorporation and rapidly rescued after photo-decaging. n = 2. k, The luciferase activity of three positive NLuc variants before and after photo-decaging. Mean ± s.d.; n = 3; two-tailed t-test. All above-mentioned samples are biological replicates. P values are shown in the figure.
Extended Data Fig. 6 Orthogonal activation of the inhibitor-resistant MEK1 variants.
a, Structural view of the nine recommended anchor sites of MEK1 by CAGE-prox (PDB code 1S9I). The three experimentally validated positive variants are labelled in orange. b, Experimental validation of the nine recommended MEK1 variants by monitoring downstream ERK phosphorylation. Three positive variants (I216, D217 and N221) were identified as efficient anchor sites. n = 2. c, The screening strategy for identifying the inhibitor-resistance MEK1 mutant. The three identified CAGE-prox-activatable MEK1 variants were inspected which yielded four enzyme variant–inhibitor ‘resistant pairs’. d, Dose-dependent inhibition of the MEK1(N221)-ONBY variant showed that this variant is not resistant to inhibitor 5, with observable inhibition starting only at the 5× IC50 concentration. n = 2. e, Inhibitors screening of the MEK1(I216Y) variant identified two ‘resistant’ pairs. n = 2. f, g, Dose-dependent inhibition of the MEK1(I216Y) variant by inhibitor 2 (f) and inhibitor 6 (g). n = 3. h, Inhibitor screening on the MEK1(D217Y) variant yielded no ‘resistant’ pairs. n = 2. i, Dose-dependent inhibition the MEK1(N221Y) variant by inhibitor 2. n = 2. j, IC50 evaluation of all the four MEK1 mutant–inhibitor pairs in comparison with that of the wild-type MEK1–inhibitor pair. Mean ± s.d.; n = 3. k, The predicted binding affinities among the three mutants and eight inhibitors. The scores represent the relative binding energy of each inhibitor–mutant complex: the higher the score, the weaker the binding. A total of five inhibitor–mutant pairs (score > 0.5, coloured in red) were calculated and predicted as ‘resistant pairs’. The four resistant pairs that were validated experimentally were shown in light blue background with the numbers underlined. l, The expression level of ONBY-incorporated MEK1 (MEK1(N221)-ONBY) and the corresponding endogenous MEK1 protein. n = 2. All above-mentioned samples are biological replicates.
Extended Data Fig. 7 Cell morphological changes and activity measurement of caspase family members after photo-activation of caspase-3.
a, Apoptosis of HEK293T cells after photo–induced caspase-3 activation. Scale bars for all frames, 10 μm. n = 2. b, Apoptosis of HeLa cells after caspase-3 activation. Scale bars of all frames, 5 μm. In both cells, morphological changes upon apoptosis were observed as early as 30 min after photo-activation of caspase-3. n = 2. c, Experimental validation revealed that incorporation of ONBY at M61 blocked caspase-3 activity, which can be efficiently rescued after photo-decaging. Mean ± s.d.; n = 3; two-tailed t-test. d, Schematic of the experimental design showing that the activities of nine caspase family members were measured in HEK293T cells with either overexpression or temporal photo-activation of caspase-3. e, Normalized activity of each caspase from control cells (red), cells with transient overexpression of wild-type (WT) caspase-3 (blue), cells with overexpression of caspase-3-M61-ONBY before (green) and after (orange) photo-activation. Overexpression of the wild-type caspase-3 resulted in markedly increased activities for most of the other caspases, whereas direct photo-activation of the caspase-3 variant enabled by CAGE-prox only activated this specific enzyme without much influence on other caspases. It allows the temporal profiling of proteolytic substrates immediately after caspase-3 activation. Error bars represent mean ± s.d.; n = 3; two-tailed t-test. f, Replotting of d with activities of all caspase members shown in the same scale. Error bars represent mean ± s.d.; n = 3. All above-mentioned samples are biological replicates. P values are shown in the figure.
Extended Data Fig. 8 Analysis and verification of the identified proteolytic substrates upon caspase-3 activation.
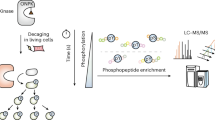
a, Schematic of the workflow for CAGE-prox-enabled temporal profiling of the proteolytic substrates immediately after caspase-3 activation. b, Venn diagram showing a summary of numbers of the proteolytic substrates identified from three independent temporal profiling experiments. A total of 544 proteins was commonly identified from all the three experiments. c–e, Sequence analysis of the cleavage sites in the identified proteolytic substrates. The logos were generated using Icelogo (https://iomics.ugent.be/icelogoserver/), with all the identified cleavage sequences aligned (the cleavage site is between the P1 and P1′ position). The sequence logos of cleavage sites at all 20 amino acids (526), Asp only (79) and Glu only (29) are shown in c, d and e, respectively. Although Asp is the most abundant cleavage site, the background of other non-caspase sites is relatively high in our results, owing to the lack of N terminus enrichment. The typical DEVD/E motifs were not observed directly, but the amino acid pattern at the P1′ position is similar to that found in previous studies41. f, Venn diagram comparing the substrates identified from this work with the proteolytic sites recorded in DegraBase (https://wellslab.ucsf.edu/degrabase/). A total of 773 overlapping substrates was found, including 544 substrates that were identified with Asp-containing peptides. g, The expression level of ONBY-incorporated caspase-3 (M61–ONBY) and the corresponding endogenous caspase-3. n = 2. h, Verification of the in vitro cleavage assay using actin, caspase-3 and PARP as positive controls. n = 2. i, Verification of the cleaved and secreted HMGB1 in the culture medium. HMGB1–Flag and wild-type caspase-3 were co-transfected into HEK293T cells, and the cell culture medium were concentrated before immunoblotting analysis. n = 2. j, Alanine screening mapped the caspase-3’s cleavage site on HMGB1 as D139/D140. n = 2. k–v, Verification of the newly identified proteolytic substrates of caspase-3. Recombinant caspase-3 protein was added into cell lysate of HEK293T followed by immunoblotting analysis. n = 2. w, Comparison of the cleavage kinetics of ATP6V1A versus ATP6V1B by caspase-3. After the recombinant wild-type caspase-3 was added into the cell lysate, the mixture was incubated at 37 °C for 0, 0.5, 1, 2, 3, 4 and 6 h, followed by immunoblotting analysis. n = 2. All above-mentioned samples are biological replicates.
Extended Data Fig. 9 CAGE-prox-enabled protein activation as a pro-drug therapy in living mice.
a, The traditional pro-drug strategy on small-molecule drugs can be extended to proteins by adopting the CAGE-prox-enabled in situ activation of therapeutic proteins. b, Schematic of the CAGE-prox-activated LF as a potential pro-drug therapy. MEK1-dependent A375 human melanoma cells were planted into mice as a xenograft model; activated LF will cause the death of MAPK-dependent tumour cells by rapid cleavage on MEK kinases. c, Experimental validation of the CAGE-prox-activatable LF variants, with the activity evaluated by cleavage efficiency on MEK3 in HEK293T cells. Incorporation of ONBY at Y659 or Y728 blocked LF’s activity, which can be efficiently rescued by photo-decaging. n = 2. d, e, Delivery and activation of LF(Y659)-ONBY was demonstrated in HeLa cells and HEK293T cells. LF(Y659)-ONBY was expressed and purified in Esherichia coli and delivered into the target cell lines by PA. n = 2. f, The growth curves of HEK293T or HeLa cells treated with LF(Y659)-ONBY/PA. Activated LF had negligible influence on the proliferation of MEK-independent HEK293T and HeLa cells. The red arrows represent the time point at which the medium was exchanged and LF–PA treatments were performed. Mean ± s.d.; n = 3. g, Photo-activation of the CAGE-prox variant of FLuc in BALB/c nude mice and optimization of the UV irradiation protocol. Increased FLuc activity were observed after UV irradiation (bottom image). n = 2. h, The caged LF-ONBY was safer than the wild-type LF (LF-WT) as determined by a dose escalation protocol. Intraperitoneal injection of wild-type LF into healthy mice every 2 days for a 2-week period caused about 50% of animal death. By contrast, injection of LF(Y659)-ONBY with a fourfold-higher dosage during the same period had negligible adverse effects on mice (n = 4). i, Photo-activation of the LF-ONBY variant has negligible influence on mouse body weight in a two-week period treatment. Mean ± s.d.; n = 8. j, Photo-activation of the LF-ONBY variant markedly reduced the tumour weight in the xenograft model. Mean ± s.d.; n = 8; two-tailed t-test. k, Evaluation of UV light penetration in vitro. UV light was irradiated through the skin and re-activation of FLuc was used to evaluate the penetration ability. Mean ± s.d.; n = 3. l, Evaluation of UV light penetration in vivo. A caged fluorescent dye was intra-tumourally injected into mice followed by UV irradiation for 5 min. The tumour was then excised and the depth of light penetration was examined. n = 2. m, n, Bio-distribution of the injected caged LF in mice. The Cy5-labelled LF protein was injected into mice, followed by imaging the whole mouse body. Mean ± s.d.; n = 3. o, The immunogenicity of LF–PA can be reduced by fusing cell-surface-targeting elements. EGF, epidermal growth factor that can target its cell-surface receptor EGFR; ZHer, an affibody that can target its receptor HER2. Mean ± s.d.; n = 3. p, The immunogenicity of LF–PA can be reduced by the addition of immunosuppressors (pentostatin combined with cyclophosphamide). Mean ± s.d.; n = 3. All above-mentioned samples are biological replicates. P values are shown in the figure.
Extended Data Fig. 10 A retrospective analysis of ‘failed’ CAGE-prox predictions and other potential expanded applications of CAGE-prox.
a, The CAGE-prox predicted mutations that have been experimentally evaluated as failed can be classified into two categories: ‘leaky’ (fail to block the protein activity with the inserted caged ONBY) and ‘dead’ (fail to restore the protein activity after photo-decaging). We calculated the frequency fold change (FC) of each native amino acid in each category. The fold change was defined as the frequency of a residue in a certain category/frequency in all predicted residues. x axis represents the fold change (expressed in log2) and y axis represents the log(P value). As shown in the plot, ONBY insertions at native Gly and Ser positions are more likely to result in leaky mutants (left) whereas insertion at native Arg positions is more likely to result in dead mutants (right). As expected, a native Tyr position is less likely to fail. A total of 56 anchor residues from 7 different proteins was analysed using hypergeometric test. b, CAGE-prox-enabled control of auto-phosphorylation of Src kinase. HEK293T cells were co-transfected with Src-TAG mutants and the \({{\rm{ONBY\mbox{--}RS-tRNA}}}_{{\rm{CUA}}}^{\mathrm{pyl}}\) pair, and cultured for 24 h in the presence of ONBY. After UV-triggered decaging of the Src-OBNY variants, cells were cultured at 37 °C for another 3 h before the auto-phosphorylation level of each Src mutant was detected by immunoblotting. n = 2. c, Cell-specific targeting of a POI by CAGE-prox by adding the cancer-cell-targeting ligand to PA and the N-terminal domain of LF (LFn) to a POI, respectively. All above-mentioned samples are biological replicates.
Supplementary information
Supplementary information
This file includes the Supplementary Results for some CAGE-Prox, Supplementary Methods, 8 Supplementary Tables and Supplementary Notes about the sequence of each protein used in this study.
Supplementary Data 1
The activity measurement results of all the 60 FLuc variants before and after UV activation. The activity of each variant was adjusted by the corresponding expression level.
Supplementary Data 2
The identified proteolytic substrates immediately after caspase-3 activation. The referenced substrates of caspase-3 and apoptosis are combined from previous studies and listed in the data file.
Supplementary Video 1
The apoptotic phenotype of HEK293T cells after caspase-3 activation. Frames were captured at 5 min intervals and the start of UV irradiation was set as the time zero.
Supplementary Video 2
The apoptotic phenotype of HeLa cells after caspase-3 activation. Frames were captured at 2 min intervals and the start of UV irradiation was set as the time zero.
Source data
Rights and permissions
About this article
Cite this article
Wang, J., Liu, Y., Liu, Y. et al. Time-resolved protein activation by proximal decaging in living systems. Nature 569, 509–513 (2019). https://doi.org/10.1038/s41586-019-1188-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-1188-1
This article is cited by
-
Genetically encoded bioorthogonal tryptophan decaging in living cells
Nature Chemistry (2024)
-
DeKinomics pulse-chases kinase functions in living cells
Nature Chemical Biology (2024)
-
Lighting up kinase contacts in situ
Nature Chemical Biology (2024)
-
Mechanism-based traps enable protease and hydrolase substrate discovery
Nature (2022)
-
Directed-evolution of translation system for efficient unnatural amino acids incorporation and generalizable synthetic auxotroph construction
Nature Communications (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.