Abstract
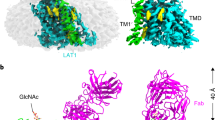
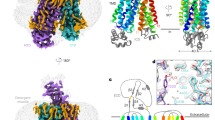
The L-type amino acid transporter 1 (LAT1; also known as SLC7A5) catalyses the cross-membrane flux of large neutral amino acids in a sodium- and pH-independent manner1,2,3. LAT1, an antiporter of the amino acid–polyamine–organocation superfamily, also catalyses the permeation of thyroid hormones, pharmaceutical drugs, and hormone precursors such as l-3,4-dihydroxyphenylalanine across membranes2,3,4,5,6. Overexpression of LAT1 has been observed in a wide range of tumour cells, and it is thus a potential target for anti-cancer drugs7,8,9,10,11. LAT1 forms a heteromeric amino acid transporter complex with 4F2 cell-surface antigen heavy chain (4F2hc; also known as SLC3A2)—a type II membrane glycoprotein that is essential for the stability of LAT1 and for its localization to the plasma membrane8,9. Despite extensive cell-based characterization of the LAT1–4F2hc complex and structural determination of its homologues in bacteria, the interactions between LAT1 and 4F2hc and the working mechanism of the complex remain largely unknown12,13,14,15,16,17,18,19. Here we report the cryo-electron microscopy structures of human LAT1–4F2hc alone and in complex with the inhibitor 2-amino-2-norbornanecarboxylic acid at resolutions of 3.3 Å and 3.5 Å, respectively. LAT1 exhibits an inward open conformation. Besides a disulfide bond association, LAT1 also interacts extensively with 4F2hc on the extracellular side, within the membrane, and on the intracellular side. Biochemical analysis reveals that 4F2hc is essential for the transport activity of the complex. Together, our characterizations shed light on the architecture of the LAT1–4F2hc complex, and provide insights into its function and the mechanisms through which it might be associated with disease.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The atomic coordinates of the BCH-bound LAT1–4F2hc complex and the JPH203-incubated LAT1–4F2hc complex have been deposited in the PDB (http://www.rcsb.org) under the accession codes 6IRT and 6IRS, respectively. The overall electron microscopy map and the BCH-focused refined electron microscopy map of the BCH-bound LAT1–4F2hc complex, the electron microscopy map of the JPH203-incubated LAT1–4F2hc complex, and the electron microscopy map of the apo wild-type LAT1–4F2hc complex have been deposited in the Electron Microscopy Data Bank (https://www.ebi.ac.uk/pdbe/emdb/) under the accession codes EMD-9722, EMD-0679, EMD-9721, and EMD-0678, respectively. Source Data for Figs. 1b–e, 3d, 4c, f and Extended Data Figs. 1a, d, 8f, 9b are available online. All other data are available from the corresponding author upon reasonable request.
References
Kanai, Y. et al. Expression cloning and characterization of a transporter for large neutral amino acids activated by the heavy chain of 4F2 antigen (CD98). J. Biol. Chem. 273, 23629–23632 (1998).
del Amo, E. M., Urtti, A. & Yliperttula, M. Pharmacokinetic role of L-type amino acid transporters LAT1 and LAT2. Eur. J. Pharm. Sci. 35, 161–174 (2008).
Fotiadis, D., Kanai, Y. & Palacín, M. The SLC3 and SLC7 families of amino acid transporters. Mol. Aspects Med. 34, 139–158 (2013).
Kageyama, T. et al. The 4F2hc/LAT1 complex transports l-DOPA across the blood–brain barrier. Brain Res. 879, 115–121 (2000).
Friesema, E. C. et al. Thyroid hormone transport by the heterodimeric human system L amino acid transporter. Endocrinology 142, 4339–4348 (2001).
Tărlungeanu, D. C. et al. Impaired amino acid transport at the blood brain barrier is a cause of autism spectrum disorder. Cell 167, 1481–1494.e18 (2016).
Kim, D. K. et al. Characterization of the system L amino acid transporter in T24 human bladder carcinoma cells. Biochim. Biophys. Acta 1565, 112–122 (2002).
Yanagida, O. et al. Human L-type amino acid transporter 1 (LAT1): characterization of function and expression in tumor cell lines. Biochim. Biophys. Acta 1514, 291–302 (2001).
Verrey, F. et al. CATs and HATs: the SLC7 family of amino acid transporters. Pflugers Arch. Eur. J. Physiol. 447, 532–542 (2004).
Scalise, M., Galluccio, M., Console, L., Pochini, L. & Indiveri, C. The human SLC7A5 (LAT1): the intriguing histidine/large neutral amino acid transporter and its relevance to human health. Front. Chem. 6, 243 (2018).
Kandasamy, P., Gyimesi, G., Kanai, Y. & Hediger, M. A. Amino acid transporters revisited: new views in health and disease. Trends Biochem. Sci. 43, 752–789 (2018).
Fort, J. et al. The structure of human 4F2hc ectodomain provides a model for homodimerization and electrostatic interaction with plasma membrane. J. Biol. Chem. 282, 31444–31452 (2007).
Meury, M. et al. Detergent-induced stabilization and improved 3D map of the human heteromeric amino acid transporter 4F2hc-LAT2. PLoS ONE 9, e109882 (2014).
Rosell, A. et al. Structural bases for the interaction and stabilization of the human amino acid transporter LAT2 with its ancillary protein 4F2hc. Proc. Natl Acad. Sci. USA 111, 2966–2971 (2014).
Jungnickel, K. E. J., Parker, J. L. & Newstead, S. Structural basis for amino acid transport by the CAT family of SLC7 transporters. Nat. Commun. 9, 550 (2018).
Gao, X. et al. Structure and mechanism of an amino acid antiporter. Science 324, 1565–1568 (2009).
Gao, X. et al. Mechanism of substrate recognition and transport by an amino acid antiporter. Nature 463, 828–832 (2010).
Fang, Y. et al. Structure of a prokaryotic virtual proton pump at 3.2 Å resolution. Nature 460, 1040–1043 (2009).
Ma, D. et al. Structure and mechanism of a glutamate-GABA antiporter. Nature 483, 632–636 (2012).
Garcia, M. L., Viitanen, P., Foster, D. L. & Kaback, H. R. Mechanism of lactose translocation in proteoliposomes reconstituted with lac carrier protein purified from Escherichia coli. 1. Effect of pH and imposed membrane potential on efflux, exchange, and counterflow. Biochemistry 22, 2524–2531 (1983).
Yamashita, A., Singh, S. K., Kawate, T., Jin, Y. & Gouaux, E. Crystal structure of a bacterial homologue of Na+/Cl−-dependent neurotransmitter transporters. Nature 437, 215–223 (2005).
Napolitano, L. et al. Novel insights into the transport mechanism of the human amino acid transporter LAT1 (SLC7A5). Probing critical residues for substrate translocation. Biochim. Biophys. Acta Gen. Subj. 1861, 727–736 (2017).
Dickens, D. et al. Modulation of LAT1 (SLC7A5) transporter activity and stability by membrane cholesterol. Sci. Rep. 7, 43580 (2017).
Gong, D. et al. Structure of the human plasma membrane Ca2+-ATPase 1 in complex with its obligatory subunit neuroplastin. Nat. Commun. 9, 3623 (2018).
Lei, J. & Frank, J. Automated acquisition of cryo-electron micrographs for single particle reconstruction on an FEI Tecnai electron microscope. J. Struct. Biol. 150, 69–80 (2005).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Grant, T. & Grigorieff, N. Measuring the optimal exposure for single particle cryo-EM using a 2.6 Å reconstruction of rotavirus VP6. eLife 4, e06980 (2015).
Zhang, K. Gctf: Real-time CTF determination and correction. J. Struct. Biol. 193, 1–12 (2016).
Scheres, S. H. Semi-automated selection of cryo-EM particles in RELION-1.3. J. Struct. Biol. 189, 114–122 (2015).
Kimanius, D., Forsberg, B. O., Scheres, S. H. & Lindahl, E. Accelerated cryo-EM structure determination with parallelisation using GPUs in RELION-2. eLife 5, e18722 (2016).
Scheres, S. H. A Bayesian view on cryo-EM structure determination. J. Mol. Biol. 415, 406–418 (2012).
Scheres, S. H. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Zhou, N., Wang, H. & Wang, J. EMBuilder: a template matching-based automatic model-building program for high-resolution cryo-electron microscopy maps. Sci. Rep. 7, 2664 (2017).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D 66, 12–21 (2010).
Amunts, A. et al. Structure of the yeast mitochondrial large ribosomal subunit. Science 343, 1485–1489 (2014).
Acknowledgements
We thank the Tsinghua University branch of China National Center for Protein Sciences (Beijing) for cryo-EM and computational facility support, and X. Li for technical support in electron microscopy data acquisition. This work was funded by the National Key Basic Research (973) Program (2015CB910101) and the National Key R&D Program (2016YFA0500402) from the Ministry of Science and Technology of China, and by the National Natural Science Foundation of China (projects 31621092, 31630017, and 81861138009).
Reviewer information
Nature thanks David Dickens, Simon Newstead and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Author information
Authors and Affiliations
Contributions
Q.Z. and R.Y. conceived the project. R.Y. and X.Z. prepared the samples. Q.Z., R.Y., X.Z., and J.L. collected the electron microscopy data. Q.Z. processed the electron microscopy data and built the atomic model. Q.Z., R.Y., and X.Z. designed, analysed, and performed the biochemical experiments. Q.Z., R.Y., and X.Z. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Biochemical characterization of the LAT1–4F2hc complex.
a, IC50 measurement of eight amino acid substrates (Phe, Trp, Leu, Tyr, l-DOPA, His, Met, and Ile) and two inhibitors (JPH203and BCH) for the inhibition of Leu uptake into the LAT1–4F2hc proteoliposomes. Owing to the low solubility of JPH203, its inhibitory effect could not be measured above 2 µM. b, The IC50 values for the listed inhibitors. NA, not available. c, SEC purification. Left, LAT1 alone was eluted ~1.5 ml after the LAT1–4F2hc complex in the presence of 0.01% LMNG and 0.001% cholesteryl hemisuccinate (CHS). Right, the peak fractions were subjected to SDS–PAGE and visualized by Coomassie blue staining. The experiment was repeated twice independently with similar results. d, The LAT1(A36E) variant has similar transport activity to the wild type in the presence of 4F2hc. The measured Km and Vmax values for the LAT1(A36E)–4F2hc complex were approximately 162.3 ± 23.6 µM and 82.5 ± 5.4 nmol mg−1 min−1, respectively. Data in a and f are mean ± s.d. of three technical independent experiments.
Extended Data Fig. 2 Cryo-EM analysis of the LAT1–4F2hc complex.
a, Representative cryo-EM micrograph and 2D class averages. The raw images are similar for all three samples; HAT + BCH is shown. The scale bar for the 2D class averages (right images) is 10 nm. b, Euler angle distribution of the 3D reconstruction for LAT1–4F2hc in the presence of BCH. c, Local-resolution maps for HAT + BCH. d, Fourier shell correlation (FSC) curves of the model refined against the overall 3.5 Å map for HAT + BCH (black); against the first half map versus the same map (red); and against the first half map versus the second half map (green). The small difference between the red and green curves indicates that the refinement of the atomic coordinates was not affected by overfitting. e–g and h–j are the same as b–d, but for HAT + JPH203 and apo-HAT, respectively. For apo-HAT, only the FSC curve of the atomic model versus the electron microscopy map was calculated. k, Gold standard FSC curves for the 3D refinement of HAT + BCH (blue), BCH-focused 3D refinement of HAT + BCH (red), HAT + JPH203 (green), and apo-HAT (cyan). l, Overall structural comparison of HAT + BCH (cyan), HAT + JPH203 (yellow), and apo-HAT (pink).
Extended Data Fig. 3 Flow chart for cryo-EM data processing.
For details, see ‘Data processing’ in the Methods section. K, number of classifications.
Extended Data Fig. 4 Electron microscopy maps of representative segments.
a, The density for the transmembrane helix of 4F2hc. b, The densities of the 12 transmembrane segments of LAT1 in the HAT + BCH complex. c, The BCH density in the electron microscopy map (marked with an asterisk) is observed in the electron microscopy map of HAT + BCH (cyan mesh), but is absent in apo-HAT (pink mesh). All of the maps are contoured at 6σ.
Extended Data Fig. 5 Comparison of the substrate-binding sites of LAT1, GkApcT, and AdiC.
The BCH-binding site in LAT1 is conserved in GkApcT and AdiC. Similar to the coordination of Arg by GkApcT and AdiC, the carboxyl and amino groups of BCH mainly form hydrogen bonds with atoms from TM1 and TM6, respectively. LAT1, GkApcT (PDB 6F34), and AdiC (PDB 3L1L) are coloured cyan, yellow, and green, respectively.
Extended Data Fig. 6 Sequence alignment of LAT1 homologues.
The sequences were aligned using CLUSTAL 2.1 (https://www.genome.jp/tools-bin/clustalw) with manual adjustments according to the 3D structures. The five aligned sequences are LAT1 and LAT2 from Homo sapiens, ApcT from Geobacillus kaustophilus, and AdiC and GadC from E. coli. The conserved residues that are involved in hydrophobic interactions with 4F2hc and the critical residues that are involved in the transport path are outlined with a black box. Amino acids that are identical or conserved in at least four sequences are coloured red or yellow. The secondary structural elements of LAT1 are indicated above the sequence alignment. Ala36 is marked with a solid black circle. Residues that are involved in 4F2hc interaction; that line the transport path; and that are related to autism spectrum disorders are indicated with solid magenta circles; solid blue circles; and solid blue triangles, respectively.
Extended Data Fig. 7 Sequence alignment of 4F2hc homologues.
a, Residues mediating the interactions with LAT1 on the extracellular interface are conserved in 4F2hc homologues. b, Alignment of the N-terminal segment and transmembrane helix. The residues involved in the interaction with LAT1 are outlined with a black box. The sequences of 4F2hc in eight vertebrates are aligned using CLUSTAL 2.1. The sequences, from top to bottom, are from Homo sapiens, Mus musculus, Rattus norvegicus, Oryctolagus cuniculus, Felis catus, Pelodiscus sinensis, Xenopus laevis, and Danio rerio, respectively. The residues that are invariant and highly conserved are coloured red and yellow, respectively. Phe190, Trp191, Trp194, Met197, Leu198, Ile205, Lys533, and Arg535—which are mapped to the interface of the two proteins—are indicated by solid magenta circles. Arg183 in the lipid-binding pocket is indicated by a solid blue circle.
Extended Data Fig. 8 Lipid molecules bound to LAT1.
a, Densities for the bound lipids. The densities, shown in blue mesh, are contoured at 5σ. b, Multiple hydrophobic amino acid residues (spheres) coordinate the two lipids. c, The hydrophobic tails of the lipids are bound to the hydrophobic surface of the LAT1–4F2hc complex. d, The hydrophilic heads of the two lipids are both surrounded by positively charged surface patches. e, A cholesterol-like density is located in a cleft surrounded by TM3, TM9, TM10, and TM12 of LAT1. The density, shown in blue mesh, is contoured at 7σ. f, The addition of 10% (wt/wt) cholesterol during proteoliposome preparation significantly enhanced the transport activity of the LAT1–4F2hc complex. EPL, E. coli polar lipid extract. Data are mean ± s.d. of three independent experiments. Each dot represents a technical replicate.
Extended Data Fig. 9 The LAT1 mutations A246V and P375L, which are associated with autism spectrum disorders, retain residual transport activity in vitro.
a, The mutants containing A246V or P375L displayed similar behaviour in solution to the wild-type complex when subjected to SEC. The experiment was repeated three times independently with similar results. b, The LAT1(A246V) and LAT1(P375L) mutants retained less than 5% transport activity compared with the wild type. The results of the counterflow assays are shown as mean ± s.d. of three independent experiments. Each dot represents a technical replicate. c, Ala246 of LAT1 is located at the beginning of TM6 and is involved in the interaction between TM1b and TM6a. d, Phe375 of LAT1 is located on the intracellular tip of TM9, and is surrounded by hydrophobic residues and the bound lipids. Mutation of Phe375 may affect the conformation of helix H4, which is involved in 4F2hc interaction.
Supplementary information
Supplementary Figures
This file contains uncropped blot scans.
Rights and permissions
About this article
Cite this article
Yan, R., Zhao, X., Lei, J. et al. Structure of the human LAT1–4F2hc heteromeric amino acid transporter complex. Nature 568, 127–130 (2019). https://doi.org/10.1038/s41586-019-1011-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-019-1011-z
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.